 Back
BackDNA Replication and Repair: Mechanisms and Molecular Players
Study Guide - Smart Notes

DNA Replication: Principles and Models
Base Pairing and the Template Mechanism
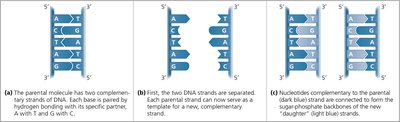
DNA replication is the process by which a cell copies its DNA, ensuring genetic information is faithfully transmitted to daughter cells. The double helix structure of DNA, with specific base pairing (A with T, G with C), allows each strand to serve as a template for the synthesis of a new complementary strand.
Base Pairing: Each parental DNA strand dictates the sequence of the new strand by complementary base pairing.
Template Mechanism: The two DNA strands separate, and new nucleotides are added according to base-pairing rules, resulting in two identical DNA molecules.
Semiconservative Replication: Each daughter DNA molecule consists of one parental and one newly synthesized strand.
Example: If the parental strand sequence is 5'-ATGC-3', the new strand will be 3'-TACG-5'.
Alternative Models of DNA Replication
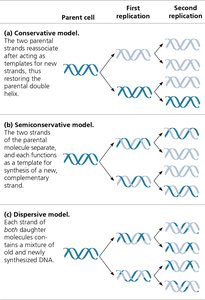
Three models were proposed to explain how DNA replicates:
Conservative Model: The parental double helix remains intact, and an entirely new double helix is synthesized.
Semiconservative Model: Each daughter molecule has one parental and one new strand (supported by experimental evidence).
Dispersive Model: Each strand of both daughter molecules contains a mixture of old and new DNA.
Experimental Evidence: Meselson-Stahl Experiment
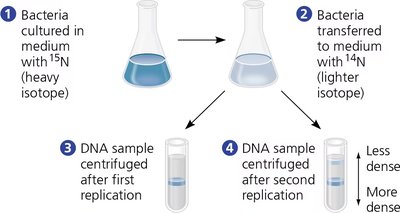
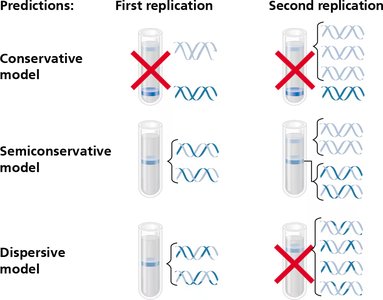
The Meselson-Stahl experiment provided strong evidence for the semiconservative model. They used isotopic labeling of DNA and density gradient centrifugation to distinguish old and new DNA strands.
Method: E. coli were grown in heavy nitrogen (15N), then transferred to light nitrogen (14N). DNA samples were collected after each replication round and centrifuged.
Results: After one replication, DNA was of intermediate density (hybrid), ruling out the conservative model. After two replications, both light and hybrid DNA were present, ruling out the dispersive model.
Mechanisms of DNA Replication
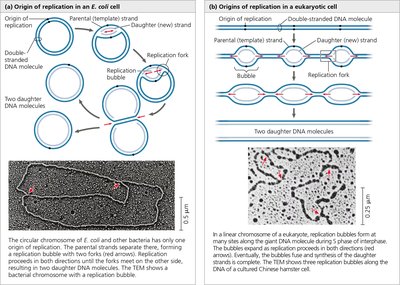
Origins of Replication and Replication Bubbles
Replication begins at specific DNA sequences called origins of replication. In bacteria, there is typically a single origin, while eukaryotic chromosomes have multiple origins, allowing for faster replication of large genomes.
Replication Bubble: The DNA strands separate at the origin, forming a bubble with two replication forks moving in opposite directions.
Bidirectional Replication: DNA synthesis proceeds in both directions from each origin.
Proteins Involved in DNA Replication
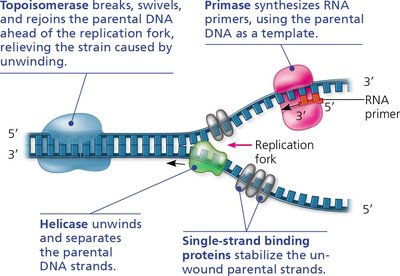
Multiple enzymes and proteins coordinate the unwinding and synthesis of new DNA strands at the replication fork:
Helicase: Unwinds and separates the parental DNA strands.
Single-Strand Binding Proteins: Stabilize unwound DNA, preventing re-annealing.
Topoisomerase: Relieves strain ahead of the replication fork by breaking, swiveling, and rejoining DNA strands.
Primase: Synthesizes short RNA primers needed to start DNA synthesis.
DNA Polymerase III: Extends the new DNA strand from the primer by adding nucleotides in the 5'→3' direction.
DNA Polymerase I: Replaces RNA primers with DNA nucleotides.
DNA Ligase: Joins Okazaki fragments on the lagging strand, forming a continuous DNA strand.
Synthesis of New DNA Strands
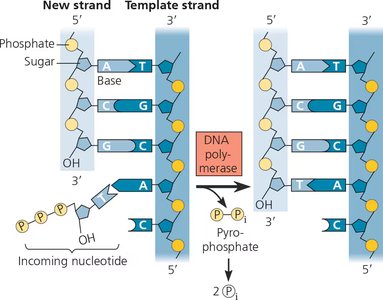
DNA polymerases add nucleotides to the 3' end of a growing DNA strand. Each incoming nucleotide is a deoxynucleoside triphosphate (dNTP), and the energy for polymerization comes from the hydrolysis of pyrophosphate.
Directionality: DNA synthesis always proceeds in the 5'→3' direction.
Energy Source: Hydrolysis of the two terminal phosphates (pyrophosphate) provides energy for the reaction.
Equation:
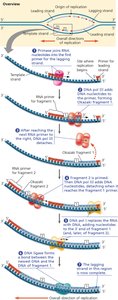
Leading and Lagging Strand Synthesis
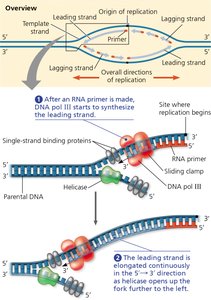
Because DNA polymerases can only add nucleotides to the 3' end, the two new strands are synthesized differently:
Leading Strand: Synthesized continuously toward the replication fork.
Lagging Strand: Synthesized discontinuously away from the fork as short Okazaki fragments, each requiring a new primer.
Okazaki Fragments: Short DNA segments on the lagging strand, later joined by DNA ligase.
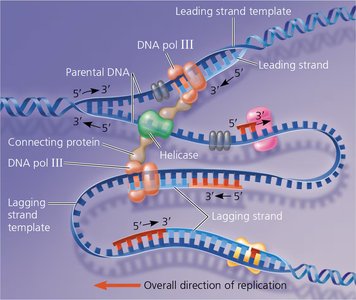
Coordination of Replication Machinery
The proteins involved in DNA replication form a large complex, sometimes called the "replisome" or "DNA replication machine." This complex coordinates the synthesis of both leading and lagging strands, possibly by anchoring to the nuclear matrix in eukaryotes and "reeling in" the parental DNA.
Proofreading and Repair of DNA
Proofreading by DNA Polymerases
DNA polymerases possess proofreading activity, correcting errors during DNA synthesis. If an incorrect nucleotide is incorporated, the enzyme removes it and replaces it with the correct one, greatly increasing fidelity.
Error Rate: Initial error rate is 1 in 105 nucleotides; after proofreading and repair, the error rate drops to 1 in 1010.
DNA Repair Mechanisms
Cells have multiple repair systems to correct DNA damage and replication errors:
Mismatch Repair: Enzymes remove and replace incorrectly paired nucleotides missed by DNA polymerase.
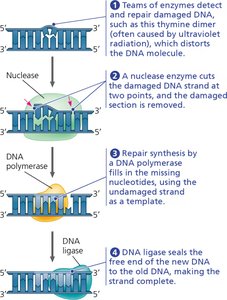
Nucleotide Excision Repair: Damaged DNA segments are excised by nucleases and replaced using the undamaged strand as a template. This is crucial for repairing lesions such as thymine dimers caused by UV light.
Telomeres and the End-Replication Problem
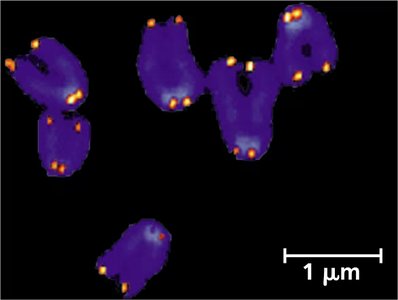
Telomeres: Protecting Chromosome Ends
Linear eukaryotic chromosomes face the end-replication problem, where the 5' ends cannot be fully replicated, leading to progressive shortening. Telomeres, repetitive noncoding sequences at chromosome ends, act as buffers to protect genes from erosion.
Telomerase: An enzyme that extends telomeres in germ cells, preventing loss of essential genetic information.
Significance: Telomere shortening is associated with aging; telomerase activity is often upregulated in cancer cells.
Summary Table: Key Proteins in DNA Replication (E. coli)
Protein/Enzyme | Function |
|---|---|
Helicase | Unwinds parental double helix at replication forks |
Single-strand binding protein | Stabilizes single-stranded DNA |
Topoisomerase | Relieves overwinding strain ahead of replication fork |
Primase | Synthesizes RNA primers |
DNA polymerase III | Main enzyme for DNA synthesis |
DNA polymerase I | Removes RNA primers and replaces with DNA |
DNA ligase | Joins Okazaki fragments |
Evolutionary Significance of DNA Replication Fidelity
High fidelity in DNA replication and repair is essential for organismal survival and the accurate transmission of genetic information. However, rare mutations that escape repair are the source of genetic variation, driving evolution and the emergence of new species.
Additional info: The notes above integrate foundational concepts from the molecular basis of inheritance, including the experimental validation of the semiconservative model, the molecular machinery of replication, and the importance of DNA repair and telomere maintenance in genome stability and evolution.
