 Back
BackDNA Structure and Replication: Foundations of Genetic Material
Study Guide - Smart Notes

DNA Structure and Replication
Discovery of DNA as Genetic Material
The identification of DNA as the hereditary material was a pivotal moment in biology. Early experiments distinguished DNA from proteins as the carrier of genetic information.
Griffith-Avery Experiment: Demonstrated transformation in bacteria, suggesting a 'transforming principle' (later identified as DNA).
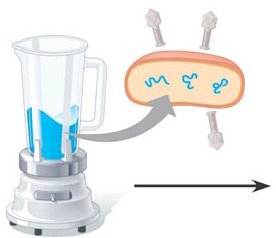
Hershey-Chase Experiment: Used bacteriophages to show that DNA, not protein, enters bacterial cells and directs viral replication.
Watson and Crick: Deduced the double helix structure of DNA, integrating chemical and physical data.
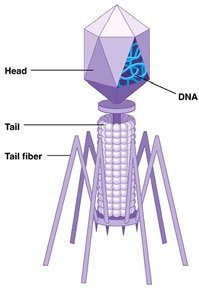
Example: The Hershey-Chase experiment used radioactive isotopes to label DNA and protein, confirming DNA as the genetic material.
DNA Structure
DNA is a double-stranded helical molecule composed of nucleotides. Its structure is essential for its function as genetic material.
Nucleotide: Consists of a phosphate group, deoxyribose sugar, and a nitrogenous base (A, T, G, C).
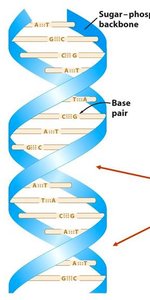
Double Helix: Two antiparallel strands held together by hydrogen bonds between complementary bases.
Base Pairing: Adenine pairs with Thymine (A-T), and Guanine pairs with Cytosine (G-C).
Major and Minor Grooves: The helical structure creates grooves that are important for protein binding.
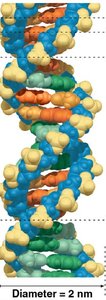
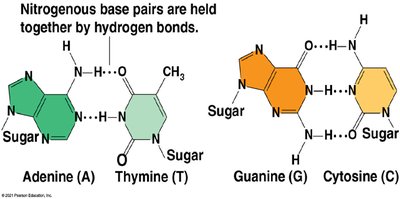
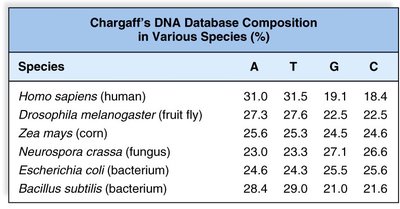
Example: Chargaff's rules state that the amount of A equals T, and G equals C in DNA from any species.
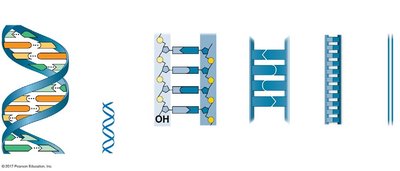
Key Features Defining DNA Structure
Double-stranded helix
Antiparallel orientation (5' to 3' and 3' to 5')
Right-handed helix
Major and minor grooves
DNA Packaging in Chromosomes
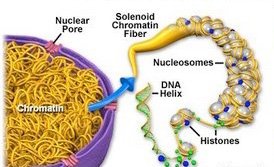
To fit within the nucleus, DNA is highly compacted with proteins into chromatin and chromosomes.
Nucleosome: Basic unit of DNA packaging, consisting of DNA wrapped around histone proteins.
Chromatin: The complex of DNA and proteins that forms chromosomes.
Levels of Compaction: DNA wraps around histones, forming nucleosomes ('beads on a string'), which further coil and fold into higher-order structures.

DNA Replication
Overview of DNA Replication
DNA replication is the process by which a cell duplicates its DNA before cell division. It ensures genetic continuity between generations of cells.
Semiconservative Model: Each new DNA molecule consists of one parental and one newly synthesized strand.
Origin of Replication (ORI): Specific sequence where replication begins.
Replication Fork: The Y-shaped region where the DNA is split into two strands for copying.

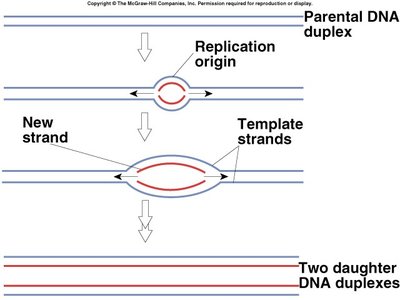
Steps of DNA Replication
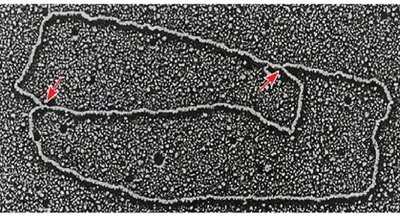
Unwinding the DNA: Helicase separates the two DNA strands at the ORI, creating a replication bubble.
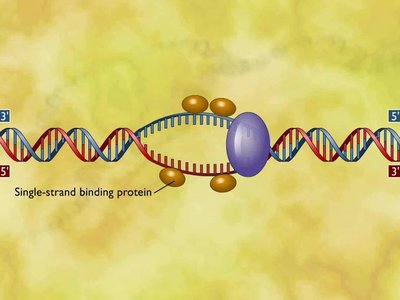
Stabilization: Single-stranded binding proteins (SSBP) prevent re-annealing of the strands.
Relieving Tension: Topoisomerase prevents supercoiling ahead of the replication fork.
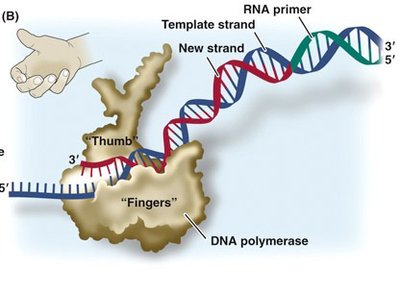
Primer Synthesis: Primase synthesizes a short RNA primer to provide a starting point for DNA polymerase.
DNA Synthesis: DNA polymerase adds nucleotides to the 3' end of the primer, synthesizing the new strand in a 5' to 3' direction.
Leading and Lagging Strands: The leading strand is synthesized continuously, while the lagging strand is synthesized in short Okazaki fragments.
Primer Removal and Ligation: DNA polymerase I removes RNA primers and replaces them with DNA; DNA ligase seals the gaps between fragments.

Leading vs. Lagging Strand Synthesis
Leading Strand: Synthesized continuously toward the replication fork.
Lagging Strand: Synthesized discontinuously away from the fork in Okazaki fragments.
Okazaki Fragments: Short DNA segments on the lagging strand, later joined by DNA ligase.
Limitations and Solutions in DNA Replication
DNA Polymerase Limitations: Cannot initiate synthesis without a primer; can only add nucleotides to the 3' end.
End Replication Problem: Linear chromosomes shorten with each replication due to incomplete synthesis of 5' ends.
Telomeres: Repetitive DNA sequences at chromosome ends that protect coding regions.
Telomerase: Enzyme that extends telomeres in germ cells and some stem cells.
Proofreading and DNA Repair
High fidelity in DNA replication is achieved through proofreading and repair mechanisms.
Proofreading: DNA polymerase checks and corrects errors during synthesis.
Mismatch Repair: Enzymes correct errors missed by DNA polymerase.
Nucleotide Excision Repair: Removes and replaces damaged DNA segments.
Key Terms and Enzymes
Helicase: Unwinds the DNA double helix.
Single-strand binding protein (SSBP): Stabilizes unwound DNA.
Primase: Synthesizes RNA primers.
DNA Polymerase: Synthesizes new DNA strands.
Ligase: Joins Okazaki fragments.
Topoisomerase: Relieves supercoiling.
Telomerase: Extends telomeres.
Summary Table: Leading vs. Lagging Strand
Feature | Leading Strand | Lagging Strand |
|---|---|---|
Synthesis | Continuous | Discontinuous (Okazaki fragments) |
Direction | Toward replication fork | Away from replication fork |
Primer Requirement | One primer | Multiple primers |
DNA Ligase Role | Minimal | Joins fragments |
Additional info:
Mutations arising from replication errors are a source of genetic variation, which is essential for evolution by natural selection.
Shortening of telomeres is associated with cellular aging and cancer biology.
