 Back
BackGene Expression: From Gene to Protein (Transcription, Translation, and Mutations)
Study Guide - Smart Notes

Gene Expression: From Gene to Protein
Overview of Gene Expression
Gene expression is the process by which information from DNA is used to synthesize functional gene products, primarily proteins. This process involves two main stages: transcription and translation. Proteins serve as the link between genotype and phenotype, dictating the traits of an organism.
Transcription: The synthesis of RNA from a DNA template.
Translation: The synthesis of a polypeptide (protein) from an mRNA template.
Central Dogma: The flow of genetic information: DNA → RNA → Protein.
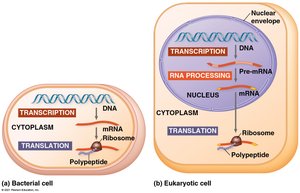
Differences Between Prokaryotic and Eukaryotic Gene Expression
Gene expression differs between prokaryotic and eukaryotic cells, primarily due to cellular structure.
Prokaryotes: Transcription and translation occur simultaneously in the cytoplasm.
Eukaryotes: Transcription occurs in the nucleus, followed by RNA processing. Translation occurs in the cytoplasm.
RNA Processing: Eukaryotic cells modify RNA transcripts (pre-mRNA) before translation.

Transcription: DNA-Directed Synthesis of RNA
Basic Principles of Transcription
Transcription is the first stage of gene expression, where RNA polymerase synthesizes RNA from a DNA template. The RNA produced is complementary to the template strand of DNA.
RNA Polymerase: Enzyme that catalyzes RNA synthesis; does not require a primer.
Promoter: DNA sequence where RNA polymerase binds to initiate transcription.
Transcription Unit: The stretch of DNA that is transcribed into RNA.
Stages of Transcription
Transcription occurs in three stages:
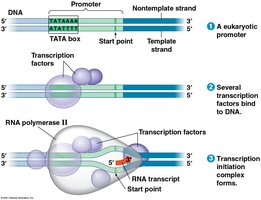
Initiation: RNA polymerase binds to the promoter, often with the help of transcription factors (in eukaryotes, the TATA box is crucial).
Elongation: RNA polymerase moves along the DNA, adding nucleotides to the 3' end of the growing RNA strand.
Termination: Transcription ends when RNA polymerase reaches a terminator sequence (bacteria) or a polyadenylation signal (eukaryotes).
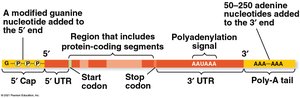
RNA Processing in Eukaryotes
Modification of mRNA Ends
Before mRNA leaves the nucleus, it undergoes several modifications:
5' Cap: Addition of a modified guanine nucleotide to the 5' end.
Poly-A Tail: Addition of 50–250 adenine nucleotides to the 3' end.
These modifications facilitate export, protect mRNA, and help ribosome binding.
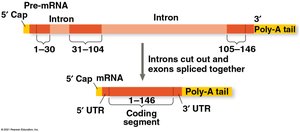
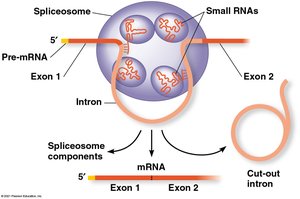
RNA Splicing: Removal of Introns
Most eukaryotic genes contain noncoding regions called introns that are removed during RNA splicing. The remaining coding regions, exons, are joined together.
Spliceosome: A complex of proteins and small RNAs that catalyzes the removal of introns.
Alternative Splicing: Allows a single gene to code for multiple polypeptides by varying which exons are included.
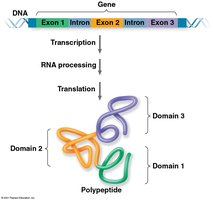
Protein Structure and Exon Shuffling
Proteins often have modular domains, and different exons may code for different domains. Exon shuffling can lead to new protein functions.
Domains: Discrete functional regions of a protein.
Exon Shuffling: Mixing and matching exons between genes during evolution.
Translation: RNA-Directed Synthesis of Polypeptides
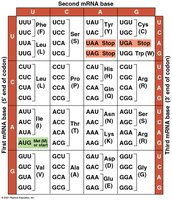
Genetic Code and Codons
The genetic code is a set of rules by which information encoded in mRNA is translated into proteins. Each codon (three-nucleotide sequence) specifies an amino acid.
64 Codons: 61 code for amino acids, 3 are stop signals.
Redundancy: Multiple codons can code for the same amino acid.
Reading Frame: Codons must be read in the correct grouping.
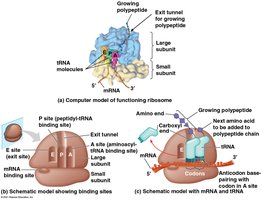
Translation Process
Translation occurs in three stages: initiation, elongation, and termination. Ribosomes facilitate the decoding of mRNA into a polypeptide chain.
Initiation: Small ribosomal subunit binds mRNA and initiator tRNA (methionine), then large subunit joins.
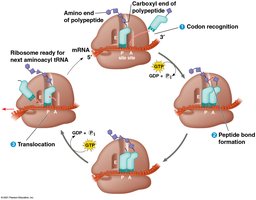
Elongation: Amino acids are added one by one to the growing chain, involving codon recognition, peptide bond formation, and translocation.
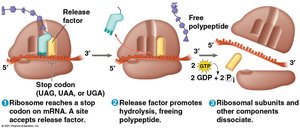
Termination: Occurs when a stop codon is reached; a release factor triggers the release of the polypeptide.
Transfer RNA (tRNA) Structure and Function
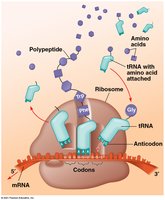
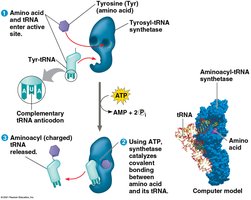
tRNA molecules carry amino acids to the ribosome and match them to the mRNA codons via their anticodon.
Anticodon: Three-nucleotide sequence on tRNA complementary to mRNA codon.
Aminoacyl-tRNA Synthetase: Enzyme that attaches the correct amino acid to tRNA.
Wobble: Flexible pairing at the third base of a codon allows some tRNAs to bind to multiple codons.
Post-Translational Modifications and Protein Targeting
Protein Folding and Modifications
After translation, polypeptides fold into their functional three-dimensional shapes. Additional modifications may be required for activity.
Folding: Spontaneous or assisted by chaperone proteins.
Modifications: Glycosylation (adding sugars), methylation (adding methyl groups), and others.
Targeting Proteins to Specific Locations
Proteins are directed to their proper cellular locations by signal peptides. Ribosomes can be free in the cytosol or bound to the endoplasmic reticulum.
Signal Peptide: Short sequence that directs the polypeptide to the ER or other organelles.
SRP (Signal Recognition Particle): Binds signal peptide and directs ribosome to ER membrane.
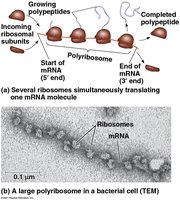
Polyribosomes
Multiple ribosomes can translate a single mRNA simultaneously, forming a polyribosome, which increases the efficiency of protein synthesis.
Mutations and Their Effects on Protein Structure and Function
Types of Mutations
Mutations are changes in the genetic material that can affect protein structure and function. They can be spontaneous or induced by mutagens.
Point Mutations: Changes in a single nucleotide pair.
Substitutions: Replace one nucleotide pair with another; can be silent, missense, or nonsense.
Insertions/Deletions: Addition or loss of nucleotide pairs; often cause frameshift mutations.
Mutagens: Physical or chemical agents that cause mutations; most carcinogens are mutagens.
Effects of Mutations
Mutations can lead to abnormal proteins, loss of function, or new traits. The impact depends on the type and location of the mutation.
Silent Mutation: No effect on protein sequence.
Missense Mutation: Changes one amino acid in the protein.
Nonsense Mutation: Creates a premature stop codon.
Frameshift Mutation: Alters the reading frame, often resulting in a nonfunctional protein.
Summary Table: Types of Mutations
Mutation Type | Description | Effect on Protein |
|---|---|---|
Silent | Substitution, no change in amino acid | No effect |
Missense | Substitution, changes amino acid | Possible loss or change of function |
Nonsense | Substitution, creates stop codon | Truncated, usually nonfunctional protein |
Frameshift | Insertion/deletion, alters reading frame | Usually nonfunctional protein |
Additional info:
Gene expression is regulated at multiple levels, including transcription, RNA processing, translation, and post-translational modification.
Alternative splicing increases protein diversity in eukaryotes.
The genetic code is nearly universal, allowing genes from one species to be expressed in another.
