 Back
BackRegulation of Gene Expression: Mechanisms and Biological Significance
Study Guide - Smart Notes

Regulation of Gene Expression
Introduction to Gene Expression Regulation
Gene expression regulation is essential for cellular function, adaptation, and development. Cells control which genes are expressed, when, and to what extent, allowing them to respond to environmental changes and differentiate into specialized types.
Bacterial Gene Regulation
Transcriptional Regulation in Bacteria
Bacteria often regulate gene expression at the transcriptional level to conserve resources and respond efficiently to environmental changes. Two primary mechanisms are feedback inhibition and gene regulation at the transcriptional level.
Feedback inhibition: The end product of a metabolic pathway inhibits an enzyme involved earlier in the pathway, preventing overproduction.
Gene regulation: Cells can adjust enzyme production by regulating the transcription of genes encoding those enzymes.
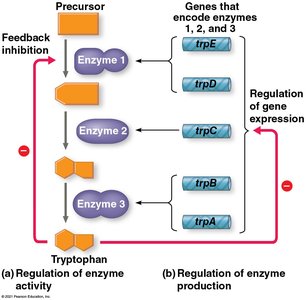
The Operon Model
An operon is a cluster of functionally related genes regulated together by a single "on-off switch" called the operator. The operon includes the operator, promoter, and the genes they control. A repressor protein can bind to the operator to block transcription, and its activity is often regulated by other molecules (corepressors or inducers).
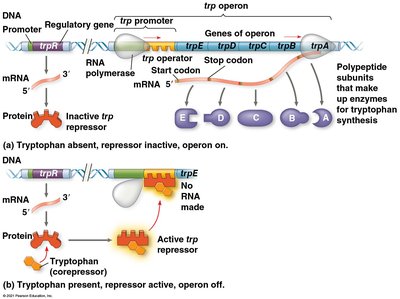
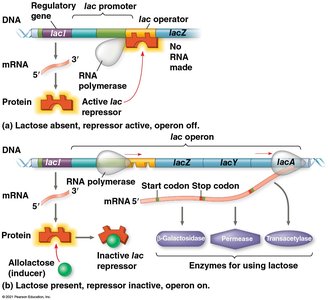
Repressible and Inducible Operons
Repressible operon: Usually on; can be turned off by a repressor (e.g., trp operon).
Inducible operon: Usually off; can be turned on by an inducer that inactivates the repressor (e.g., lac operon).
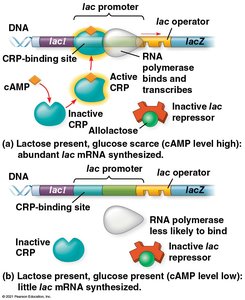
Positive Gene Regulation
Some operons are also regulated by activator proteins, such as the cyclic AMP receptor protein (CRP). When glucose is scarce, CRP is activated by cAMP and enhances transcription of the lac operon.
Eukaryotic Gene Regulation
Overview of Eukaryotic Gene Expression Control
Gene expression in eukaryotes is regulated at multiple levels, including chromatin structure, transcription, RNA processing, translation, and protein modification or degradation. This regulation is crucial for cell specialization and response to environmental signals.
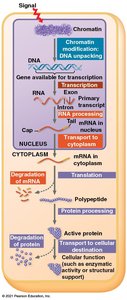
Regulation of Chromatin Structure
Chromatin structure influences gene accessibility. Genes in tightly packed heterochromatin are usually not expressed, while those in loosely packed euchromatin are more accessible. Chemical modifications of histones (e.g., acetylation) and DNA (e.g., methylation) play key roles in regulating chromatin structure and gene expression.
Histone acetylation: Addition of acetyl groups to histone tails loosens chromatin, promoting transcription.
DNA methylation: Addition of methyl groups to DNA bases reduces transcription and can cause long-term gene silencing.
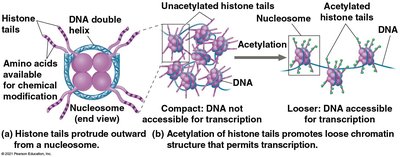
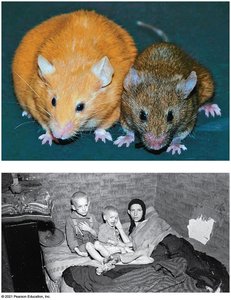
Epigenetic Inheritance
Epigenetic inheritance refers to the transmission of traits by mechanisms not involving changes in DNA sequence, such as chromatin modifications. These can explain differences in gene expression between genetically identical individuals.
Transcriptional Regulation in Eukaryotes
Transcription initiation is controlled by the interaction of transcription factors with control elements in the DNA. General transcription factors are required for all protein-coding genes, while specific transcription factors bind to enhancers or proximal control elements to regulate particular genes.
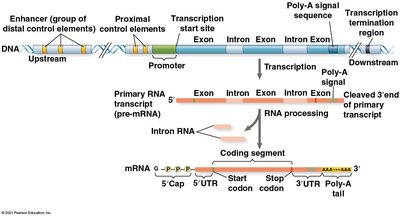
Enhancers and Activators
Enhancers are distal control elements that can greatly increase transcription of associated genes. Activator proteins bind to enhancers and facilitate the assembly of the transcription initiation complex at the promoter.
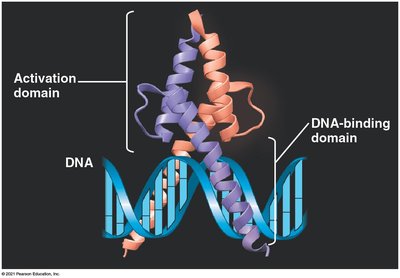
Combinatorial Control and Coordinated Gene Expression
Gene expression is often regulated by combinations of control elements and transcription factors, allowing precise spatial and temporal control. In eukaryotes, co-expressed genes may be scattered across the genome but share common control elements recognized by the same activators.
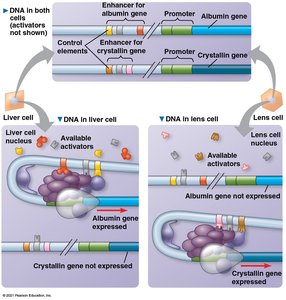
Nuclear Architecture
Chromosome conformation and the spatial organization of chromatin within the nucleus can influence gene expression. Regions of chromosomes may interact at transcription factories, specialized sites rich in transcription machinery.
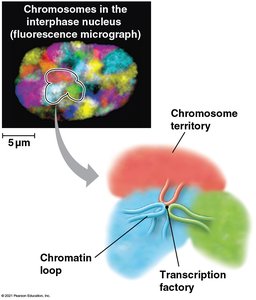
Post-Transcriptional Regulation
Gene expression can be regulated after transcription through mechanisms such as alternative RNA splicing, mRNA degradation, and translational control.
Alternative RNA splicing: Different mRNAs can be produced from the same primary transcript, increasing protein diversity.
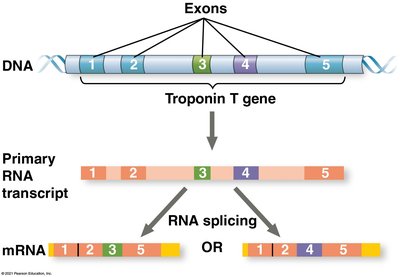
mRNA degradation: The stability of mRNA molecules affects how much protein is produced. Sequences in the 3' UTR influence mRNA lifespan.
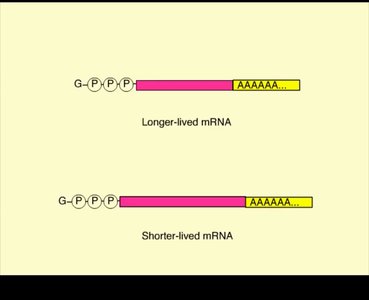
Translational regulation: Regulatory proteins can block translation initiation, and global translation can be regulated in response to cellular signals.
Protein processing and degradation: Proteins may be chemically modified, cleaved, or marked for degradation by ubiquitin and destroyed by proteasomes.
Noncoding RNAs and Gene Expression
Roles of Noncoding RNAs (ncRNAs)
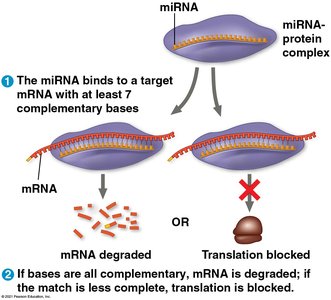
Many noncoding RNAs (ncRNAs) play crucial roles in gene regulation. These include microRNAs (miRNAs), small interfering RNAs (siRNAs), and piwi-interacting RNAs (piRNAs).
miRNAs: Bind to complementary mRNA sequences, leading to mRNA degradation or translational repression.
siRNAs: Similar to miRNAs, involved in RNA interference (RNAi) and defense against viruses.
piRNAs: Induce heterochromatin formation and silence transposons in animal germ cells.
Differential Gene Expression in Development
Cell Differentiation and Morphogenesis
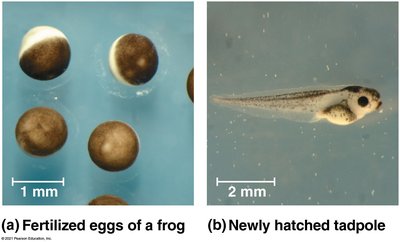
During development, differential gene expression leads to the formation of specialized cell types, tissues, and organs. This process involves cytoplasmic determinants, inductive signals, and sequential gene regulation.

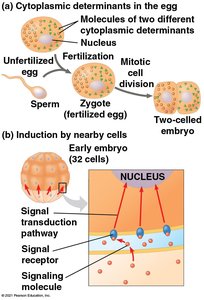
Pattern Formation and Axis Establishment
Pattern formation is the spatial organization of tissues and organs, guided by positional information. In Drosophila, maternal effect genes and morphogen gradients (e.g., bicoid protein) establish body axes and segment identity.
Gene Regulation and Cancer
Genetic Changes Leading to Cancer
Cancer results from mutations that disrupt normal gene regulation, particularly in genes controlling the cell cycle. These include:
Oncogenes: Mutated proto-oncogenes that promote uncontrolled cell division.
Tumor-suppressor genes: Normally inhibit cell division; mutations can remove these restraints.
Mutations in genes such as ras (a G protein) and p53 (a tumor suppressor) are common in cancers. Multiple mutations are usually required for cancer to develop, and both inherited and environmental factors contribute to cancer risk.
Summary Table: Key Mechanisms of Gene Regulation
Level of Regulation | Mechanism | Example |
|---|---|---|
Transcriptional | Operons, transcription factors, enhancers | trp operon, lac operon, eukaryotic enhancers |
Chromatin Structure | Histone modification, DNA methylation | Acetylation, methylation, epigenetic inheritance |
Post-Transcriptional | Alternative splicing, mRNA degradation | Troponin T gene splicing, mRNA stability |
Translational | Regulatory proteins, miRNAs | miRNA-mediated repression |
Post-Translational | Protein modification, degradation | Ubiquitin-proteasome pathway |
