 Back
BackCell Signaling and Signal Transduction Mechanisms
Study Guide - Smart Notes

Signal Transduction Mechanisms: Cell Communication
Introduction to Cell Signaling
Cell signaling is the process by which cells detect and respond to external stimuli, allowing them to coordinate activities and adapt to their environment. This process involves the transmission of signals from the cell surface to the interior, resulting in specific cellular responses.
Signaling molecules (ligands) can be secreted or membrane-bound and include hormones, growth factors, neurotransmitters, and pheromones.
Some receptors are activated by changes in metabolite levels (e.g., oxygen, nutrients) or physical stimuli (e.g., light, touch, heat).
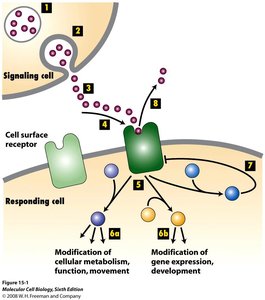
Types of Cell Signaling
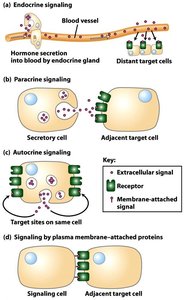
Cells communicate using different signaling mechanisms based on the distance between the signaling and target cells.
Endocrine signaling: Hormones are released into the bloodstream and act on distant target cells.
Paracrine signaling: Signals act on nearby cells.
Autocrine signaling: Cells respond to signals they themselves produce.
Juxtacrine signaling: Direct contact between signaling and target cells via membrane-attached proteins.
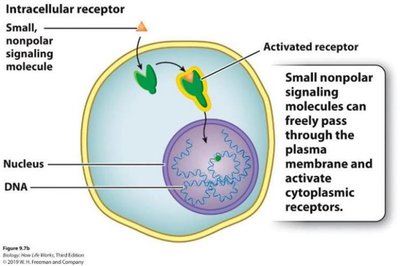
Cell Surface and Intracellular Receptors
Receptors are proteins that bind signaling molecules and initiate cellular responses. They can be located on the cell surface or inside the cell.
Cell surface receptors: Bind hydrophilic or large ligands that cannot cross the plasma membrane.
Intracellular receptors: Bind small, nonpolar ligands that diffuse through the membrane (e.g., steroid hormones).
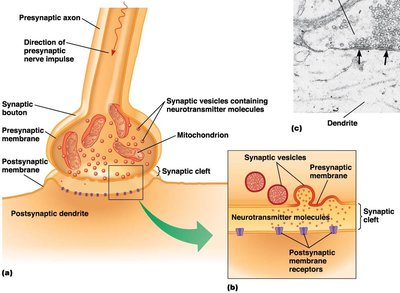
Synaptic Signaling and Neurotransmitters
Neurons communicate via chemical synapses, where neurotransmitters are released from the presynaptic neuron and bind to receptors on the postsynaptic cell.
Neurotransmitters are stored in synaptic vesicles and released upon arrival of an action potential.
They diffuse across the synaptic cleft and bind to specific receptors, triggering a response in the postsynaptic cell.
Specificity and Sensitivity of Receptor-Ligand Interactions
Receptors exhibit specificity for their ligands, and the sensitivity of a cell to a signal depends on the number and affinity of its receptors.
Binding specificity: Determined by molecular complementarity between ligand and receptor.
Dissociation constant (Kd): The concentration of free ligand at which half the receptors are occupied. High affinity receptors have low Kd values.
Different cell types may have different responses to the same ligand due to distinct receptor types or downstream signaling pathways.
Cell Type | Ligand | Receptor | Response |
|---|---|---|---|
Skeletal Muscle | Acetylcholine | Ion Channel | Contraction |
Heart Muscle | Acetylcholine | GPCR | Slows contraction rate |
Pancreatic Cells | Acetylcholine | GPCR | Exocytosis of digestive enzymes |
Agonists and Antagonists
Synthetic ligands can modulate receptor activity:
Agonists: Activate the receptor, mimicking the natural ligand.
Antagonists: Bind the receptor without activating it, blocking the natural ligand's effect.
Highly Conserved Components of Cell Signaling
Several intracellular proteins and small molecules are common to many signaling pathways:
Protein kinases: Enzymes that add phosphate groups to proteins, often activating or deactivating them.
Protein phosphatases: Remove phosphate groups from proteins.
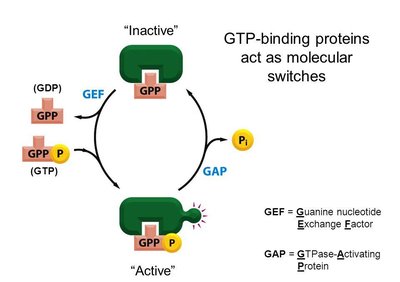
GTP-binding proteins (G-proteins): Act as molecular switches, cycling between active (GTP-bound) and inactive (GDP-bound) states.
Second messengers: Small molecules like cAMP and Ca2+ that amplify and propagate signals within the cell.
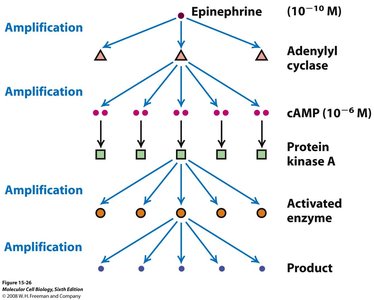
Second Messengers and Signal Amplification
Second messengers rapidly diffuse within the cell, amplifying the signal initiated by ligand-receptor binding.
cAMP: Activates protein kinase A, leading to phosphorylation of target proteins.
Ca2+: Triggers processes such as muscle contraction, secretion, and neurotransmitter release.
Signal amplification allows a small number of activated receptors to produce a large cellular response.
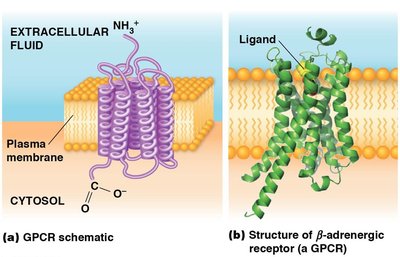
G Protein Coupled Receptors (GPCRs)
Structure and Function of GPCRs
GPCRs are a large family of cell surface receptors that respond to a variety of external signals. They are characterized by seven transmembrane domains and interact with trimeric G proteins to transmit signals inside the cell.
Ligand binding activates the receptor, which then activates the associated G protein by promoting the exchange of GDP for GTP on the alpha subunit.
The activated G protein can then regulate downstream effectors such as enzymes or ion channels.
GPCR Signaling Pathways
Different GPCRs activate distinct intracellular signaling cascades:
GPCR—Gαs/Gαi—Adenylyl cyclase—cAMP—PKA
GPCR—Gαq—Phospholipase C—IP3/DAG—PKC
GPCR—Gαt—cGMP phosphodiesterase—cGMP
GPCR—Gαi—Ion channels
Example: Epinephrine binds to β-adrenergic receptors (a type of GPCR) to mediate the fight-or-flight response, increasing glucose and fatty acid availability for energy.
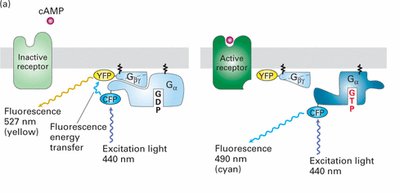
Experimental Analysis of G Protein Activation
Fluorescence Resonance Energy Transfer (FRET) can be used to study the activation of G proteins in living cells. When the G protein subunits are together, energy transfer occurs between fluorescent tags; upon activation and dissociation, the energy transfer is lost, indicating G protein activation.
Summary Table: Major Components of Signal Transduction
Component | Function | Example |
|---|---|---|
Ligand | Initiates signaling by binding receptor | Epinephrine, Acetylcholine |
Receptor | Detects ligand and transduces signal | GPCR, Tyrosine kinase receptor |
G Protein | Molecular switch, relays signal | Trimeric G protein |
Second Messenger | Amplifies and propagates signal | cAMP, Ca2+ |
Effector Protein | Executes cellular response | Protein kinase A, Ion channel |
Additional info: The notes above integrate foundational concepts from cell biology, including receptor-ligand interactions, second messenger systems, and the molecular mechanisms underlying GPCR signaling, as well as experimental approaches to studying these pathways.
