 Back
BackDNA Replication and Repair: Mechanisms, Errors, and Cellular Strategies
Study Guide - Smart Notes

DNA Replication: Mechanisms and Requirements
Semi-Conservative Nature of DNA Replication
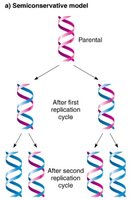
DNA replication is described as semi-conservative because each daughter DNA molecule consists of one parental (original) strand and one newly synthesized strand. This mechanism ensures the faithful transmission of genetic information across generations.
Retention of original strands: Each parental strand serves as a template for a new complementary strand.
Experimental evidence: The Meselson-Stahl experiment demonstrated that after replication, DNA molecules contain one heavy (parental) and one light (new) strand.
Genetic stability: This process preserves genetic information and minimizes errors due to complementary base pairing (A-T, G-C).
Requirements for DNA Replication
DNA replication requires several key components to ensure accuracy and efficiency:
Single-stranded DNA template: Provides the sequence to be copied.
Raw materials (dNTPs): Deoxynucleoside triphosphates serve as building blocks and energy sources.
Enzymes and proteins: Unwind DNA, synthesize primers, assemble nucleotides, and seal nicks.
Prokaryotic DNA Replication
Mode of Replication
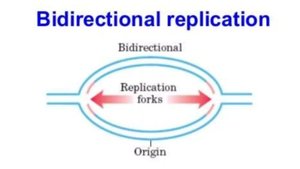
Bidirectional (Theta) Replication: Replication begins at a single origin (OriC) and proceeds in both directions, forming two replication forks.
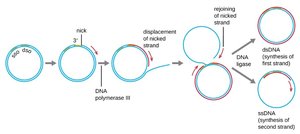
Rolling-Circle Replication: Used by plasmids and some viruses, involving strand displacement and synthesis of a new strand.
Initiation
Origin of replication (OriC): Contains AT-rich sequences recognized by DnaA protein, which bends and unwinds DNA.
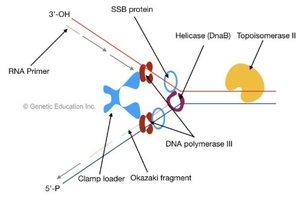
Helicase (DnaB): Unwinds DNA further, recruited by DnaC.
Single-stranded binding proteins (SSBs): Stabilize unwound DNA & prevents re-annealing or secondary structure formation.
DNA gyrase (topoisomerase II): Relieves tortional strain & supercoiling ahead of the fork caused by unwinding.
Elongation
Primase: Synthesizes short RNA primers which provide a 3' -OH group for DNA polymerase to extend.
DNA polymerase III: Main enzyme for DNA elongation, highly processive due to the β-clamp that secures DNA polymerase to the template strand. adds dNTPs to the 3'-OH of the RNA primer
Leading strand: Synthesized continuously toward the fork. (5'-3')
Lagging strand: Synthesized discontinuously as Okazaki fragments, each initiated by a new primer. (5'-3')
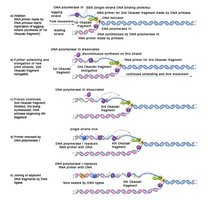
DNA polymerase I: Removes RNA primers from okazaki fragments and replaces them with DNA.
DNA ligase: Joins Okazaki fragments by forming phosphodiester bonds
the replisome is a protein complex at the replication fork that coordinates helicase, primase, DNAA polymerase and other factors
Termination
Ter sites bound by Tus proteins: Terminate replication by blocking helicase activity.
Topoisomerase IV: separates interlinked daughter chromosomes.
Proofreading by DNA polymerase III and mismatch repair : Ensure high fidelity (error rate ~1 in 10⁹ bp).
Eukaryotic DNA Replication
Mode of Replication
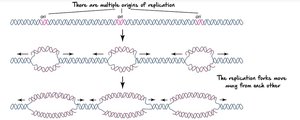
Bidirectional replication from multiple origins: Ensures timely duplication of large, linear chromosomes.
Initiation
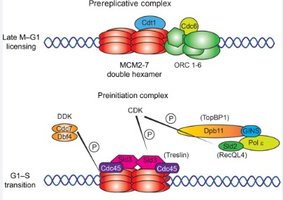
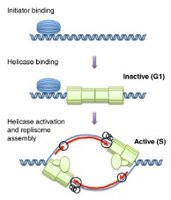
Replication licensing: Ensures replication only occurs once per cell cycle. Key factors include Cdc6, Cdt1, and the MCM2-7 helicase complex loaded onto origins during G1 phase of cell cycle .
Activation: Cyclin-dependent kinases (CDKs) phosphorylate licensing factors and helicase to initiate replication.
Unwinding
helicase (MCM complex): separates two strands creating a replication bubble
Replication protein A (RPA): binds to single stranded DNA to stabilize it and prevent re-annealing or formation of secondary structures
Topoisomerase: relieves supercoiling and tortional strain caused by unwinding ahead of the replication fork.
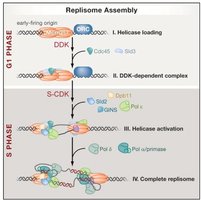
Elongation
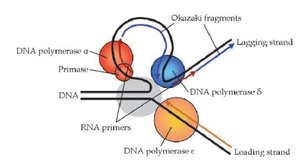
DNA polymerase α-primase: Synthesizes RNA primer and short DNA stretch. This primer proves a 3'-OH group for further DNA synthesis.
DNA polymerase ε: Synthesizes leading strand continuously towards the replication fork (5'-3').
DNA polymerase δ: Synthesizes lagging strand (Okazaki fragments) away from replication fork (5'-3').
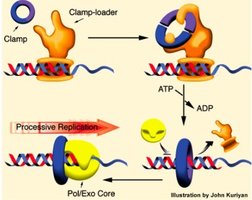
PCNA (sliding clamp): Ensures processivity of polymerases.
RFC (clamp loader): Loads PCNA onto DNA Ensuring processivity of polymerases..
Histone chaperones (CAF-1, ASF1): Assemble nucleosomes on new DNA.
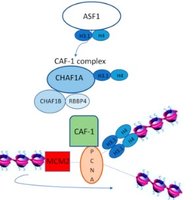
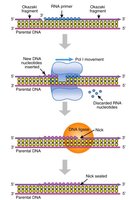
Primer Removal and Gap Filling
RNase H and FEN1: Remove RNA primers from Okazaki fragments.
DNA polymerase δ: Extends DNA to fill gaps left by primer removal
DNA ligase I: Joins Okazaki fragments by forming phosphodiester bonds.
Termination
Telomeres: Repetitive sequences at chromosome ends that prevent loss of genetic information, maintained by telomerase in germ, stem, and cancer cells.
Proofreading and mismatch repair: DNA polymerases ε and δ have 3'→5' exonuclease activity to remove mismatched nucleotides; mismatch repair corrects errors missed by polymerase proofreading post-replication.
Comparison Table: Prokaryotic vs. Eukaryotic DNA Replication
Feature | Prokaryotic Cells | Eukaryotic Cells |
|---|---|---|
Origins of replication | Single origin | Multiple origins |
Chromosome structure | Circular | Linear |
Enzymes | DNA polymerase III | DNA polymerases α, δ, ε |
Telomeres/telomerase | Absent | Present |
Nucleosome assembly | Not required | Essential |
Location | Cytoplasm | Nucleus |
DNA Replication Errors and Mutations
Sources of Replication Errors
Mismatched base pairing: Incorrect nucleotide incorporation.
Slippage in repetitive sequences: Causes insertions or deletions (indels).
Chemical modifications: Oxidative stress, alkylation, deamination.
External mutagens: UV radiation, chemicals, viruses.
Types of Base Mutations
Point mutations: Single nucleotide changes.
Transition mutations: Purine ↔ Purine or Pyrimidine ↔ Pyrimidine (A↔G, C↔T).
Transversion mutations: Purine ↔ Pyrimidine (A↔C, G↔T). likely to cause structural changes in DNA and functional changes in the resulting protein
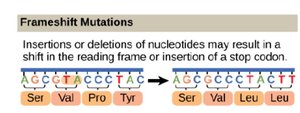
Insertions/Deletions (Indels): Addition or removal of nucleotides, may cause frameshifts resulting in different amino acid sequences
Expanding nucleotide repeats: Increase in repeat number (e.g., CAG in Huntington's disease). disrupts normal gene function.
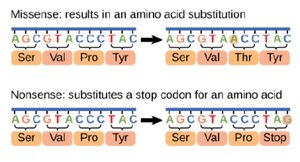
Missense mutation: Base substitution changes amino acid. can alter protein function
Nonsense mutation: Base substitution creates a stop codon leading to a non-functional protein.
Silent mutation: Codon change does not alter amino acid.
Neutral mutation: Amino acid change does not affect protein function.
Loss-of-function mutation: Reduces/eliminates protein activity. can result in nonsense mutations, frameshifts or missense mutations that impair protein structure
Gain-of-function mutation: Protein acquires new/enhanced function. often caused my missense mutations or gene duplications

Summary Table: Types of Mutations
Type of Mutation | Description | Example/Effect |
|---|---|---|
Transition | Purine ↔ Purine or Pyrimidine ↔ Pyrimidine | A ↔ G or C ↔ T; less disruptive |
Transversion | Purine ↔ Pyrimidine | A ↔ C or G ↔ T; more disruptive |
Insertion/Deletion | Addition/removal of nucleotides | Frameshifts, altered protein coding |
Expanding repeats | Increased repeat number | Huntington's disease |
Missense | Changes one amino acid | Sickle cell anemia (Glu→Val) |
Nonsense | Creates stop codon | Truncated protein |
Silent | Codon change, same amino acid | No effect |
Neutral | Amino acid change, no effect | Similar amino acid substitution |
Loss-of-function | Reduces/eliminates activity | Tumor suppressor mutations |
Gain-of-function | New/enhanced function | Oncogene activation |
DNA Repair Mechanisms
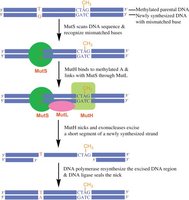
Mismatch Repair (MMR)
Corrects errors that escape proofreading, such as mismatched bases and small indels.
Detection: MutS (prokaryotes) or MSH (eukaryotes) recognize mismatches.
Strand discrimination: Newly synthesized strand identified by lack of methylation (prokaryotes) or presence of nicks (eukaryotes).
Excision and resynthesis: Exonuclease removes error; DNA polymerase and ligase fill and seal the gap.
Direct Repair
Reverses specific DNA damage without removing bases.
Photoreactivation: Photolyase repairs UV-induced pyrimidine dimers using light energy.
Alkyl transferase: Removes alkyl groups from guanine (e.g., O6-methylguanine).
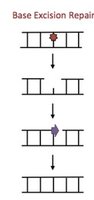
Base Excision Repair (BER)
Repairs small, non-helix-distorting lesions (e.g., deaminated, oxidized, or alkylated bases).
DNA glycosylase: Removes damaged base, creating an abasic site.
AP endonuclease: Cleaves backbone at abasic site.
DNA polymerase and ligase: Fill and seal the gap.
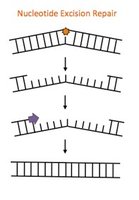
Nucleotide Excision Repair (NER)
Removes bulky, helix-distorting lesions (e.g., thymine dimers, chemical adducts).
Damage recognition: GG-NER (global genome) or TC-NER (transcription-coupled).
Strand incision: Endonucleases cut on both sides of lesion.
Excision and resynthesis: DNA polymerase fills gap; ligase seals nick.
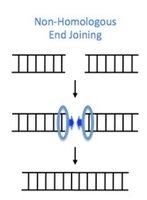
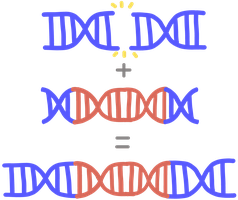
Repair of Double-Strand Breaks (DSBs)
Homology-Directed Repair (HDR): Uses homologous sequence as template for accurate repair. High fidelity.
Non-Homologous End Joining (NHEJ): Directly joins DNA ends, error-prone due to possible nucleotide loss.
Additional info:
Replication factories in eukaryotes are discrete nuclear sites where replication proteins are concentrated.
Proper chromatin assembly after replication is essential for gene regulation and genome stability.
