 Back
BackEnzymes: The Catalysts of Life – Structure, Function, and Regulation
Study Guide - Smart Notes

Enzymes: The Catalysts of Life
Introduction to Enzymes
Enzymes are organic catalysts that accelerate nearly all cellular reactions by lowering the activation energy required for those reactions. The presence of the appropriate enzyme determines whether a reaction can and will proceed under cellular conditions.
Activation Energy (EA): The minimum energy required for reactants to reach the transition state and form products.
Metastable State: Reactants are thermodynamically unstable but do not react due to insufficient activation energy.
Transition State: An intermediate state with higher free energy than reactants or products.
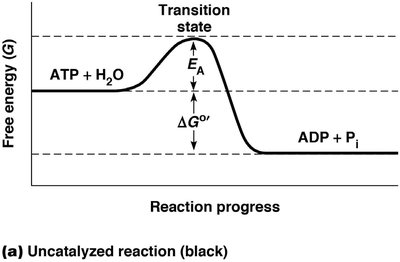
Figure: Free energy diagram showing the activation energy barrier (EA) and the change in free energy (ΔG°').
Overcoming the Activation Energy Barrier
Cells use catalysts to overcome high activation energy barriers, as increasing temperature is not feasible in isothermal biological systems. Catalysts, including enzymes, lower the EA by providing a surface for reactants to interact more efficiently.
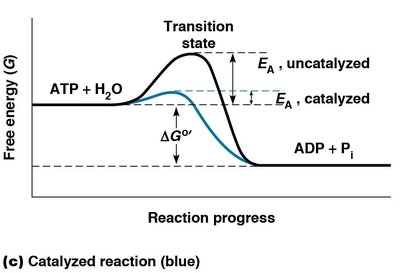
Figure: Comparison of catalyzed (blue) and uncatalyzed (black) reactions, showing reduced activation energy in the presence of an enzyme.
Properties of Enzymes as Biological Catalysts
Increase reaction rates by lowering EA.
Form transient, reversible complexes with substrates.
Alter the rate at which equilibrium is achieved, but not the equilibrium position itself.
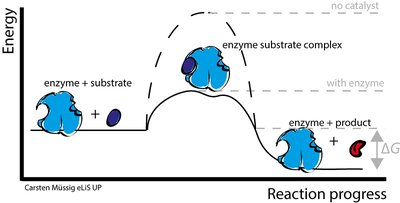
Figure: Enzyme-substrate complex formation and reduction of activation energy.
Structure and Function of Enzymes
Enzyme Composition
Most enzymes are proteins, but some RNA molecules (ribozymes) also have catalytic activity.
Figure: Enzyme action during DNA replication, illustrating the role of protein enzymes.
The Active Site
The active site is a specific cluster of amino acids within the enzyme where substrate binding and catalysis occur. The three-dimensional folding of the protein creates a groove or pocket that accommodates the substrate with high specificity.
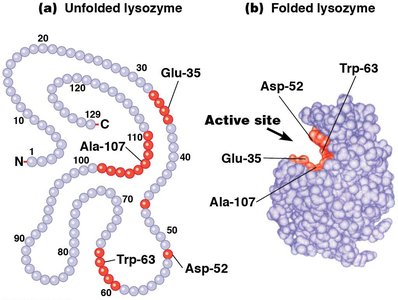
Figure: Schematic representation of an enzyme's active site and substrate binding.
Amino Acids in the Active Site
Only a subset of the 20 amino acids participate directly in catalysis, often serving as proton donors or acceptors.
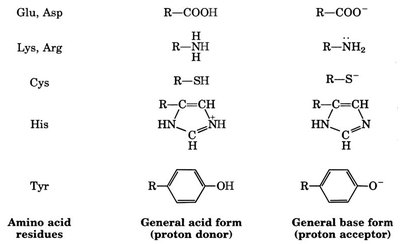
Figure: Amino acid residues involved in acid/base catalysis at the active site.
Cofactors and Coenzymes
Some enzymes require nonprotein components called cofactors (prosthetic groups) for activity. These may be metal ions or small organic molecules (coenzymes, often vitamin derivatives).
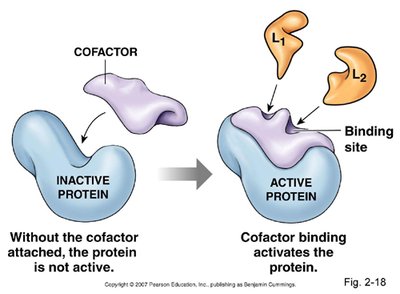
Figure: Cofactor binding is necessary for some enzymes to become active.
Enzyme Specificity and Classification
Substrate and Group Specificity
Enzymes exhibit high substrate specificity due to the shape and chemistry of the active site.
Some enzymes show group specificity, acting on a group of related substrates (e.g., carboxypeptidase A).
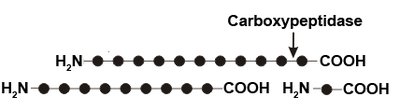
Figure: Carboxypeptidase hydrolyzes the carboxyl terminal peptide bond in polypeptides.
Enzyme Nomenclature and Classes
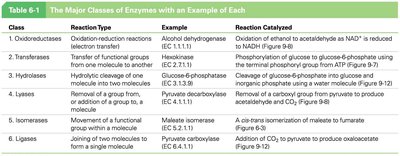
Enzymes are named based on their substrate or function and are classified into six major classes:
Class | Reaction Type | Example | Reaction Catalyzed |
|---|---|---|---|
Oxidoreductases | Oxidation-reduction | Alcohol dehydrogenase | Oxidation of ethanol to acetaldehyde |
Transferases | Transfer of functional groups | Hexokinase | Phosphorylation of glucose |
Hydrolases | Hydrolytic cleavage | Glucose-6-phosphatase | Cleavage of glucose-6-phosphate |
Lysases | Removal/addition of groups | Pyruvate decarboxylase | Removal of CO2 from pyruvate |
Isomerases | Isomerization | Maleate isomerase | Conversion of maleate to fumarate |
Ligases | Joining of molecules | Pyruvate carboxylase | Addition of CO2 to pyruvate |

Enzyme Activity: Environmental Sensitivity
Temperature Sensitivity
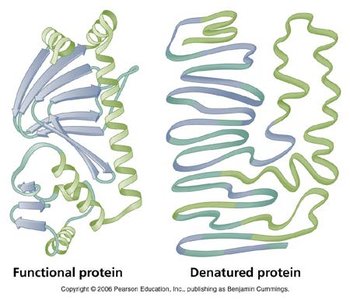
Enzyme activity increases with temperature up to an optimal point, beyond which denaturation causes loss of activity. The optimal temperature varies among enzymes and organisms.
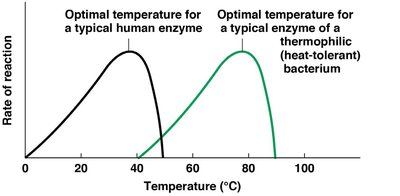
pH Sensitivity
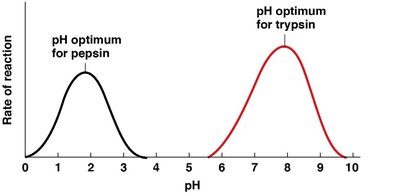
Most enzymes function within a narrow pH range, as changes in pH can disrupt ionic and hydrogen bonds essential for maintaining the active site's structure.
Other Factors
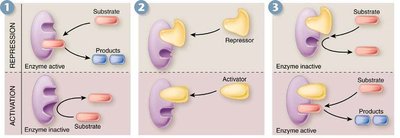
Inhibitors and Activators: Molecules that decrease or increase enzyme activity.
Ionic Strength: Affects hydrogen bonding and ionic interactions, influencing enzyme conformation.
Mechanisms of Enzyme Action
Substrate Binding and Conformational Change
Substrates bind to the enzyme's active site via noncovalent interactions, causing a conformational change that optimally orients the substrate for catalysis. This binding is reversible.
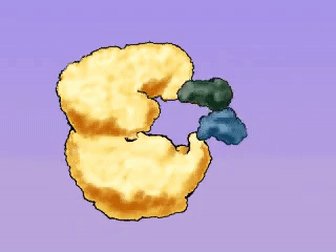
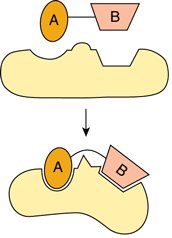
Figure: The lock-and-key model (rigid fit) and the induced-fit model (conformational change upon substrate binding).
Substrate Activation
Bond Distortion: Makes substrate bonds more susceptible to attack.
Proton Transfer: Increases substrate reactivity.
Electron Transfer: Forms temporary covalent bonds between enzyme and substrate.
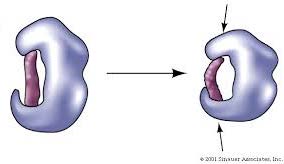
Catalytic Cycle
The enzyme catalytic cycle involves substrate binding, conformational change, conversion to product, product release, and return to the original enzyme conformation.
Ribozymes: Catalytic RNA Molecules
Discovery and Examples
Some RNA molecules, called ribozymes, possess catalytic activity. Examples include self-splicing RNA in Tetrahymena, ribonuclease P (RNase P), and ribosomal RNA (rRNA).
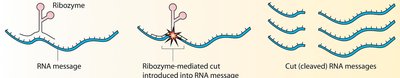
Figure: Ribozymes catalyze the cleavage of RNA messages.
Regulation of Enzyme Activity
Substrate-Level and Feedback Regulation
Substrate-level regulation: Increased substrate concentration increases reaction rate.
Feedback inhibition: Accumulation of end product inhibits an earlier step in the pathway.
Allosteric and Covalent Regulation
Allosteric regulation: Non-covalent binding of effectors at sites other than the active site, causing conformational changes that alter activity.
Covalent modification: Addition or removal of chemical groups (e.g., phosphorylation) to regulate enzyme activity.
Enzyme Inhibition
Irreversible inhibitors: Covalently bind and permanently inactivate enzymes (e.g., heavy metals, nerve gases).
Reversible inhibitors: Bind non-covalently and can dissociate. Includes competitive (bind active site) and noncompetitive (bind elsewhere) inhibitors.
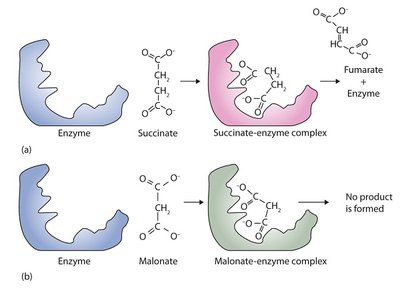
Figure: Competitive inhibition—malonate inhibits succinate dehydrogenase by occupying the active site.
Enzyme Kinetics
Michaelis–Menten Kinetics
The rate of enzymatic reactions depends on substrate concentration and can be described by the Michaelis–Menten equation:
Vmax: Maximum reaction velocity at saturating substrate concentration.
Km (Michaelis constant): Substrate concentration at which the reaction rate is half of Vmax; indicates enzyme affinity for substrate.
Where v is the initial velocity, [S] is substrate concentration, Vmax is maximum velocity, and Km is the Michaelis constant.
Lineweaver–Burk (Double-Reciprocal) Plot
The Lineweaver–Burk equation is a linear transformation of the Michaelis–Menten equation, useful for determining Km and Vmax:
This plot allows for easier determination of kinetic parameters from experimental data.
Turnover Number (kcat)
The turnover number is the number of substrate molecules converted to product per enzyme molecule per unit time at Vmax.
Summary Table: Enzyme Regulation Mechanisms
Activation | Inhibition |
|---|---|
Substrate | Product |
Covalent (phosphorylation, methylation, acetylation) | Covalent (phosphorylation, methylation, acetylation) |
Proteolytic cleavage (zymogens) | Irreversible denaturation (acids, bases, high temp, heavy metals) |
Non-covalent (allosteric activators) | Non-covalent (competitive, noncompetitive, allosteric inhibitors) |
Additional info: These notes provide a comprehensive overview of enzyme structure, function, regulation, and kinetics, suitable for college-level cell biology students.
