 Back
BackEnzymes: The Catalysts of Life – Structure, Function, and Regulation
Study Guide - Smart Notes

Enzymes: The Catalysts of Life
Introduction to Enzymes
Enzymes are biological catalysts that dramatically increase the rate of chemical reactions in cells by lowering the activation energy required for those reactions. They are essential for both spontaneous and non-spontaneous reactions, ensuring that cellular processes occur efficiently and under precise regulation.
Enzyme: A protein (or RNA molecule) that catalyzes chemical reactions in biological systems.
Substrate: The specific reactant molecule upon which an enzyme acts.
Product: The molecule(s) produced from the enzymatic reaction.
Enzymes do not alter the overall energy balance (ΔG) of a reaction; they only affect the rate at which equilibrium is reached.
Activation Energy and the Metastable State
Activation Energy (EA) and Reaction Rates
For a chemical reaction to occur, reactant molecules must overcome an energy barrier known as the activation energy (EA). This barrier ensures that only molecules with sufficient energy can react, preventing uncontrolled reactions in the cell.
Activation Energy (EA): The minimum energy required for reactants to reach the transition state and form products.
Most cellular reactions have high EA, resulting in a metastable state where reactants are thermodynamically unstable but kinetically stable.
Enzymes lower EA, allowing reactions to proceed rapidly at physiological temperatures.
Biological Importance of Activation Energy
Regulation of Metabolism: High EA allows cells to control when and where reactions occur via enzyme presence and activity.
Prevention of Extraneous Reactions: High EA prevents random, destructive reactions in the cell.
Energetic Efficiency: Cells release energy gradually, harvesting it efficiently through controlled enzymatic steps.
Enzymes as Biological Catalysts
Properties of Catalysts
Increase reaction rates by lowering EA.
Form transient, reversible complexes with substrates.
Do not alter the equilibrium position of the reaction.
Most enzymes are proteins, though some RNA molecules (ribozymes) also possess catalytic activity.
The Active Site
The active site is a specific region of the enzyme where substrate binding and catalysis occur. It is formed by the three-dimensional folding of the protein and contains key amino acid residues that interact with the substrate.
Common active site amino acids: cysteine, histidine, serine, aspartate, glutamate, lysine.
These residues participate in substrate binding and catalysis, often acting as proton donors/acceptors.
Prosthetic Groups
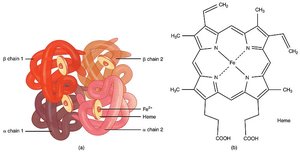
Many enzymes require nonprotein cofactors called prosthetic groups, which are often metal ions or coenzymes derived from vitamins. These groups are essential for enzyme function and are usually located near the active site.
Example: Heme group in hemoglobin and catalase.
Enzyme Specificity
Enzymes exhibit high substrate specificity due to the precise fit between the active site and the substrate. Some enzymes are highly specific for a single substrate, while others show group specificity, acting on a family of related molecules.
Enzyme Classification
Enzymes are classified into six major classes based on their function:
Oxidoreductases – catalyze oxidation-reduction reactions
Transferases – transfer functional groups
Hydrolases – catalyze hydrolysis reactions
Lyases – add or remove groups to form double bonds
Isomerases – catalyze isomerization changes
Ligases – join two molecules with covalent bonds
Factors Affecting Enzyme Activity
Temperature
Enzyme activity generally increases with temperature due to increased molecular motion, but excessive heat leads to denaturation and loss of function. Each enzyme has an optimal temperature for activity.
Human enzymes: optimal at 37°C; denature above 50–55°C.
Thermophilic enzymes: function at high temperatures (e.g., DNA polymerase from hot springs).
Psychrophilic enzymes: function at low temperatures (e.g., enzymes from Listeria bacteria).
pH
Most enzymes are active within a narrow pH range, typically 3–4 units. Changes in pH can alter the charge of amino acids at the active site, disrupting enzyme structure and function.
Ionic Strength and Other Factors
Enzyme activity is also sensitive to ionic strength, inhibitors, and activators, which can affect the enzyme's conformation and interactions.
Mechanisms of Enzyme Action
Substrate Binding and the Induced-Fit Model
Substrate binding occurs at the active site, often involving hydrogen and ionic bonds. The induced-fit model describes how substrate binding induces a conformational change in the enzyme, optimizing the fit and positioning catalytic residues for the reaction.
Substrate Activation Mechanisms
Bond distortion: Weakens substrate bonds, making them more susceptible to catalysis.
Proton transfer: Increases substrate reactivity.
Electron transfer: Forms temporary covalent bonds between enzyme and substrate.
Ribozymes: Catalytic RNA Molecules
Discovery and Function
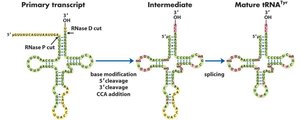
Ribozymes are RNA molecules with catalytic activity, supporting the RNA World Hypothesis. Examples include self-splicing RNA in Tetrahymena thermophila and the RNA component of ribonuclease P, which processes tRNA precursors.
Enzyme Kinetics
Michaelis–Menten Kinetics
Enzyme kinetics studies the rates of enzyme-catalyzed reactions. The Michaelis–Menten equation describes how reaction velocity (v0) depends on substrate concentration ([S]):
Vmax: Maximum reaction velocity at saturating substrate concentration.
Km: Substrate concentration at which the reaction rate is half of Vmax; a measure of enzyme affinity for substrate.
kcat: Turnover number, the number of substrate molecules converted to product per enzyme per second at Vmax.
Enzyme Inhibition
Types of Inhibition
Irreversible inhibitors: Bind covalently, permanently inactivating the enzyme (e.g., heavy metals, nerve gas).
Reversible inhibitors: Bind noncovalently and can dissociate; include competitive and noncompetitive inhibitors.
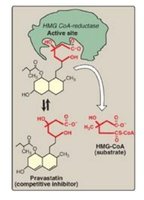
Competitive Inhibition
Competitive inhibitors resemble the substrate and bind to the active site, blocking substrate access. Increasing substrate concentration can overcome competitive inhibition.
Noncompetitive and Uncompetitive Inhibition
Noncompetitive inhibitors bind to a site other than the active site, causing a conformational change that reduces enzyme activity. Uncompetitive inhibitors bind only to the enzyme-substrate complex.
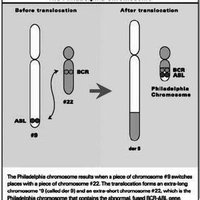
Clinical Example: Gleevec and the Philadelphia Chromosome
Chronic myeloid leukemia (CML) is driven by the BCR-ABL fusion protein, a constitutively active tyrosine kinase resulting from the Philadelphia chromosome translocation. Gleevec (imatinib) is a competitive inhibitor that targets this kinase, but resistance can develop through mutations, gene amplification, or drug efflux.
Enzyme Regulation
Substrate-Level Regulation
Enzyme activity can be regulated by substrate and product concentrations, allowing rapid adjustment to cellular needs.
Allosteric Regulation
Allosteric enzymes have regulatory sites distinct from the active site. Binding of allosteric effectors (activators or inhibitors) induces conformational changes that modulate enzyme activity. Allosteric regulation is crucial for controlling metabolic pathways and often involves feedback inhibition by pathway end products.
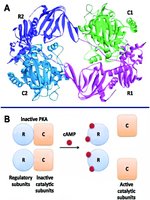
Covalent Modification
Enzyme activity can be regulated by the reversible addition or removal of chemical groups, such as phosphorylation (by kinases) and dephosphorylation (by phosphatases). This mechanism allows rapid and reversible control of enzyme function.
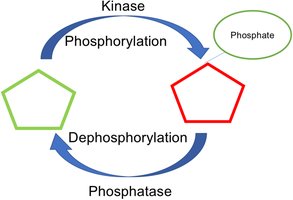
Proteolytic Cleavage
Some enzymes are synthesized as inactive precursors (zymogens) and activated by irreversible proteolytic cleavage. This is common for digestive enzymes like trypsin and chymotrypsin.
Summary Table: Types of Enzyme Inhibition
Type | Binding Site | Effect on Vmax | Effect on Km | Reversibility |
|---|---|---|---|---|
Competitive | Active site | No change | Increases | Reversible |
Noncompetitive | Allosteric site | Decreases | No change | Reversible |
Uncompetitive | Enzyme-substrate complex | Decreases | Decreases | Reversible |
Irreversible | Active or allosteric site (covalent) | Decreases | Varies | Irreversible |
Key Equations
Michaelis–Menten equation:
Turnover number:
Practice Questions
Which of the following best describes the function of an enzyme? Answer: Enzymes lower the activation energy of a reaction.
Where does a substrate bind on an enzyme? Answer: Active site
What does a low Km value indicate? Answer: The enzyme has a high affinity for its substrate.
What is an allosteric inhibitor? Answer: A molecule that binds to a site other than the active site, causing a conformational change that reduces enzyme activity.
