 Back
BackEukaryotic Gene Regulation: Mechanisms and Levels of Control (10.1)
Study Guide - Smart Notes

Eukaryotic Gene Regulation
Overview
Eukaryotic gene regulation encompasses a series of complex mechanisms that control the expression of genes at multiple levels, ensuring precise cellular function, differentiation, and response to environmental cues. Unlike prokaryotes, eukaryotes utilize chromatin structure, transcription factors, RNA processing, and post-translational modifications to regulate gene expression.
Purpose: Maintain cellular identity, respond to stimuli, conserve energy, and enable differentiation.
Levels: Chromatin structure, transcriptional, post-transcriptional, translational, and post-translational regulation.
Levels of Genetic Regulation
Chromatin Structure and Epigenetics
Chromatin structure determines the accessibility of DNA to transcription machinery. Epigenetic modifications such as DNA methylation and histone modification regulate gene expression without altering the DNA sequence.
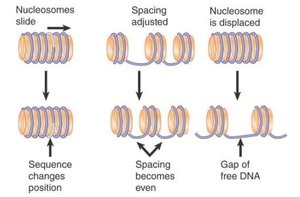
Chromatin Remodelling: Protein complexes (e.g., SWI/SNF) shift nucleosomes to expose or hide gene promoters.
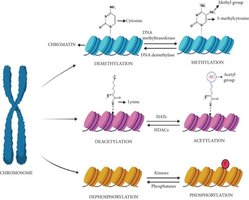
Histone Modifications: Acetylation loosens chromatin (activates transcription), deacetylation compacts chromatin (represses transcription), methylation can activate or repress depending on context.
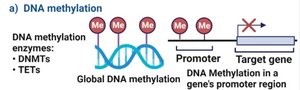
DNA Methylation: Addition of methyl groups to cytosine in CpG islands silences genes; patterns are heritable and contribute to epigenetic regulation.
Transcriptional Control
Transcriptional regulation is a major control point, involving promoters, enhancers, silencers, and transcription factors. These elements determine whether and how strongly a gene is transcribed.
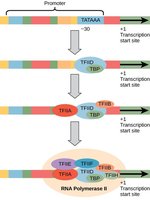
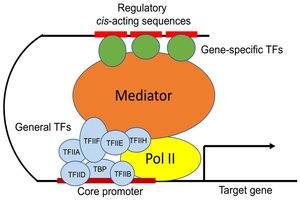
Promoters: Sites where RNA polymerase II and general transcription factors assemble.
Enhancers/Silencers: Regulatory DNA elements that can act over long distances, often cell-type specific.
Transcription Factors: Proteins that bind DNA to recruit or block transcriptional machinery, integrating signals from hormones, growth factors, or stress.
Post-Transcriptional Regulation
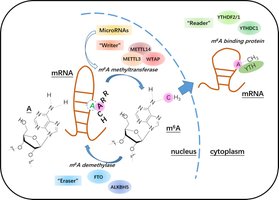
Post-transcriptional regulation involves modifications to pre-mRNA, affecting mRNA stability, transport, and translation. Alternative splicing, capping, polyadenylation, and RNA editing are key steps.
Alternative Splicing: Generates multiple mRNA isoforms from a single gene, increasing protein diversity.
5′ Capping and 3′ Polyadenylation: Protect mRNA from degradation and enhance translation.
RNA Editing: Alters nucleotide sequence of mRNA, affecting protein function.
mRNA Stability: Regulated by poly(A) tail length and RNA-binding proteins.
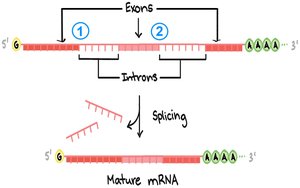
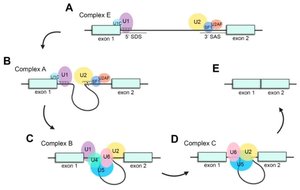
RNA Splicing
RNA splicing removes introns from pre-mRNA and joins exons to produce mature mRNA. The spliceosome, a complex of snRNPs and proteins, catalyzes this process.
Consensus Sequences: 5′ splice site (GU), 3′ splice site (AG), branch point (A).
Spliceosome: U1, U2, U4/U6, U5 snRNPs recognize splice sites and catalyze splicing.
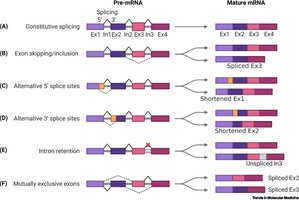
Alternative Splicing: Exon skipping, mutually exclusive exons, alternative splice sites, intron retention.
RNA Interference (RNAi)
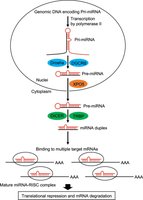
RNA interference is a gene-silencing mechanism involving small RNAs (siRNAs and miRNAs) that guide effector complexes to complementary mRNA targets, resulting in post-transcriptional repression.
siRNAs: Derived from double-stranded RNA, perfectly complementary to target mRNA, cause degradation.
miRNAs: Derived from endogenous transcripts, partially complementary, inhibit translation.
Mechanisms: mRNA cleavage, translational inhibition, transcriptional silencing, slicer-independent degradation.
Translational Control
Translational regulation determines how efficiently mRNA is translated into protein. This includes control of ribosome binding, initiation factors, and response to cellular signals.
Initiation Factors: Regulate ribosome recruitment and translation initiation.
Stress Response: Global down-regulation of translation during stress, selective translation of specific mRNAs.
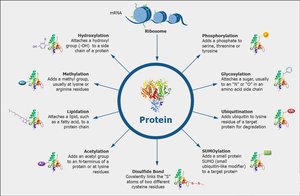
Post-Translational Modifications (PTMs)
PTMs are chemical modifications to proteins after translation, affecting their function, stability, localization, and interactions. They are crucial for modulating regulatory proteins involved in gene expression.
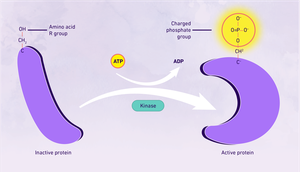
Phosphorylation: Addition of phosphate groups by kinases, alters protein activity and interactions.
Acetylation: Addition of acetyl groups to lysine residues, relaxes chromatin and facilitates transcription.
Methylation: Addition of methyl groups to histones or DNA, can activate or repress transcription.
Ubiquitination: Addition of ubiquitin, targets proteins for degradation.
Sumoylation: Addition of SUMO proteins, regulates protein localization and stability.
Glycosylation: Addition of sugar moieties, affects protein folding and cell-cell interactions.
Proteolytic Cleavage: Activation/inactivation of proteins by peptide bond cleavage.
ADP-Ribosylation: Attachment of ADP-ribose units, involved in stress responses and DNA repair.
Lipidation: Attachment of lipid groups, targets proteins to membranes.
Eukaryotic vs. Prokaryotic Regulation
Comparison Table
The mechanisms of gene regulation differ significantly between prokaryotes and eukaryotes, reflecting their structural and functional complexity.
Feature | Prokaryotes | Eukaryotes |
|---|---|---|
Gene Organization | Operons, single mRNA for multiple genes | Each gene has its own promoter, transcribed separately |
Chromatin Structure | No chromatin, DNA accessible | DNA wrapped around histones, chromatin must be modified |
Transcription & Translation | Simultaneous in cytoplasm | Transcription in nucleus, translation in cytoplasm |
Regulatory Complexity | Simple, mainly transcriptional | Complex, includes chromatin remodelling, splicing, enhancers, RNAi |
Summary
Eukaryotic gene regulation is a multi-layered process involving chromatin structure, transcription factors, RNA processing, translational control, and post-translational modifications. These mechanisms ensure precise control of gene expression, enabling cellular differentiation, adaptation, and maintenance of function. Understanding these processes is fundamental to cell biology, genetics, and biomedical research.
