 Back
BackEukaryotic Organelle Structure and Function: Isolation, Sorting, and Roles
Study Guide - Smart Notes

Eukaryotic Organelle Structure and Function
Methods for Isolation of Organelles
Isolation of cellular organelles is essential for studying their structure and function. Centrifugation is the primary method used to separate organelles based on size and density.
Differential Centrifugation: Sequential centrifugation at increasing speeds separates cell components into pellets and supernatants. Large components sediment first, followed by medium and small components.
Density Gradient Centrifugation: Organelles are separated by their density in a gradient solution (e.g., sucrose or cesium chloride), forming distinct bands that can be extracted for analysis.
Svedberg Unit (S): A measure of sedimentation rate during centrifugation, used to characterize ribosomes and other macromolecules.
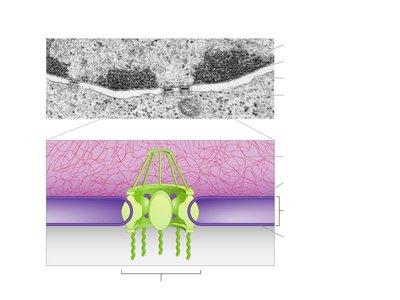
Structure of the Nuclear Envelope and Nuclear Pore Complex
The nuclear envelope is a double-membrane structure that surrounds the nucleus, containing nuclear pore complexes for regulated transport between the nucleus and cytoplasm.
Inner and Outer Membranes: The envelope consists of two lipid bilayers.
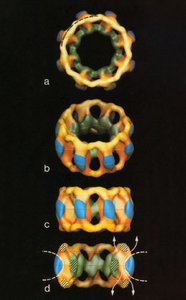
Nuclear Pore Complex: Large protein assemblies that mediate selective transport of molecules.
Symmetry and Structure: Advanced imaging reveals the three-dimensional symmetry of the nuclear pore complex.
Endoplasmic Reticulum (ER)
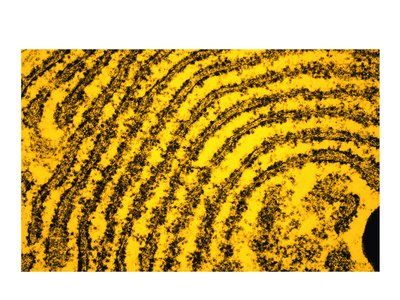
The ER is a network of membranes involved in protein and lipid synthesis, and is divided into rough and smooth regions.
Rough ER (RER): Studded with ribosomes, responsible for protein synthesis and folding.
Smooth ER (SER): Lacks ribosomes, involved in lipid synthesis, drug detoxification, and carbohydrate metabolism.
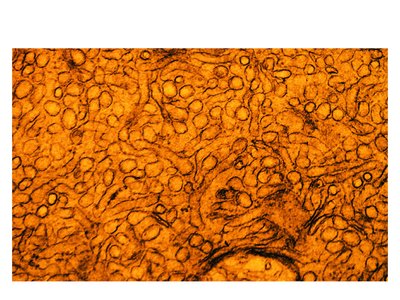
Smooth ER Functions
Hydroxylation reactions (addition of OH groups)
Drug detoxification
Glycogen catabolism (conversion of glycogen to glucose)
Membrane production
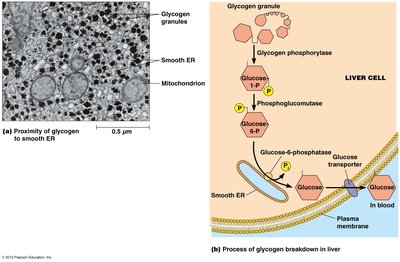
Steroid Biosynthesis in Smooth ER
Synthesis of cholesterol and steroid hormones
Enzyme HMG-CoA reductase is key in cholesterol biosynthesis
Statins target HMG-CoA reductase to lower cholesterol
Membrane Biosynthesis
ER is the primary source of membrane lipids
Fatty acids synthesized in cytoplasm and incorporated into ER membrane
Phospholipid translocators (flippases) transfer lipids across the bilayer, establishing membrane asymmetry
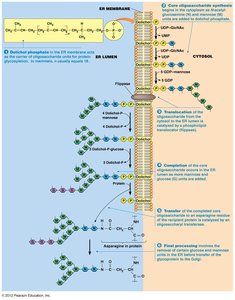
Rough ER Functions
Protein folding by chaperones
Glycosylation for post-translational modification
Insertion of membrane proteins
Sorting and export of proteins to Golgi apparatus
Golgi Apparatus
The Golgi apparatus is a stack of membrane-bound sacs responsible for modifying, sorting, and packaging proteins and lipids.
Packaging and sorting proteins for transport or export
Terminal glycosylation and other post-translational modifications (phosphorylation, sulfonation, methylation)
Protein Glycosylation and Sorting
Core glycosylation occurs in RER; terminal glycosylation in Golgi
Sugars act as signals for protein sorting
Three possibilities for terminal glycosylation: no change, editing away sugars, or addition of new sugars
Anterograde and Retrograde Transport
Anterograde: Movement of material toward the plasma membrane (exocytosis)
Retrograde: Return of vesicles from Golgi to ER, balancing lipid flow and supplying materials for new vesicles
Retention and Retrieval Tags
ER proteins contain retention tags (e.g., RXR) to prevent escape
Retrieval tags (e.g., KDEL, KKXX) ensure proteins are returned from Golgi to ER
Golgi Protein Sorting by Membrane-Spanning Domain Length
Membrane thickness increases from CGN to TGN
Proteins are sorted based on the length of their hydrophobic domains
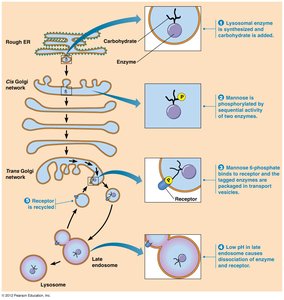
Targeting of Lysosomal Proteins
Lysosomal enzymes are tagged with mannose-6-phosphate in the Golgi, ensuring their delivery to lysosomes.
N-glycosylation and removal of glucose/mannose units
Phosphorylation of mannose residues forms mannose-6-phosphate tag
Tag binds receptor, directing enzyme to lysosome

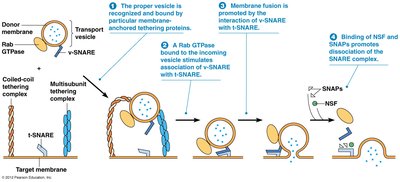
SNARE Proteins and Vesicle Fusion
SNARE proteins mediate the specificity of vesicle fusion with target membranes, ensuring correct delivery of cargo.
v-SNAREs on vesicles, t-SNAREs on target membranes
Complementary interaction allows recognition and fusion
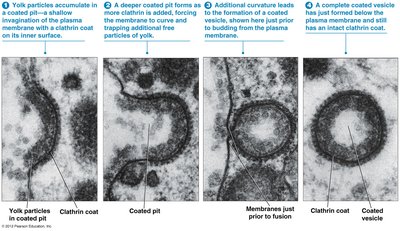
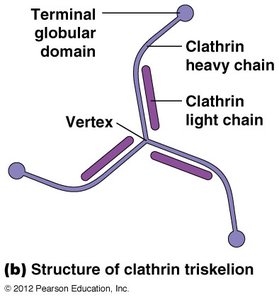
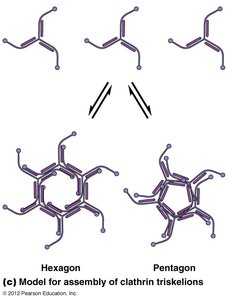
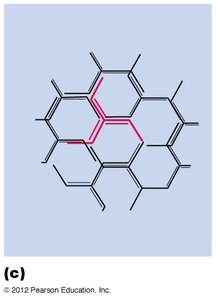
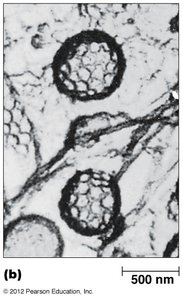
Receptor-Mediated Endocytosis and Clathrin-Coated Vesicles
Cells internalize macromolecules via receptor-mediated endocytosis, using clathrin-coated vesicles for selective sorting and transport.
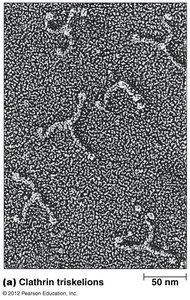
Clathrin forms a coat around vesicles, aiding in cargo selection and vesicle formation
Adaptins help clathrin coat the vesicle; dynamin closes the vesicle

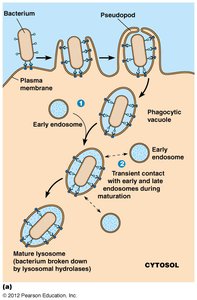
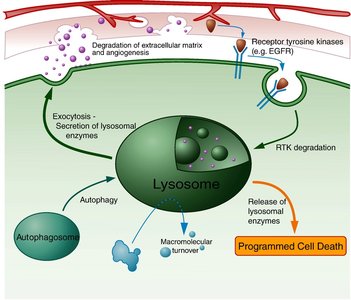
Lysosomes
Lysosomes are membrane-bound organelles containing hydrolytic enzymes for digestion and recycling of cellular components.
Digest nutrients, destroy pathogens, recycle damaged organelles
Maintain acidic pH (4-5) for enzyme activity
Protective mechanisms: pro-enzymes and pH requirements
Autophagic vs. heterophagic lysosomes: autophagy digests internal components, heterophagy digests external material
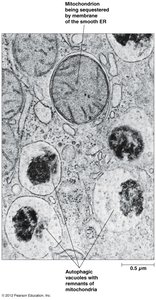

Plant Vacuole
Plant vacuoles are multifunctional organelles analogous to lysosomes, with additional roles in storage and maintaining turgor pressure.
Acidic environment for digestion
Storage of proteins, malate, anthocyanins, and waste
Regulation of cytosolic pH and turgor pressure
Peroxisomes
Peroxisomes are organelles involved in detoxification, fatty acid oxidation, and metabolism of nitrogen-containing compounds.
Detoxify oxygen radicals and harmful compounds
Oxidize fatty acids (β-oxidation)
Break down nitrogen compounds and unusual substances
Contain enzymes like superoxide dismutase and catalase for hydrogen peroxide metabolism
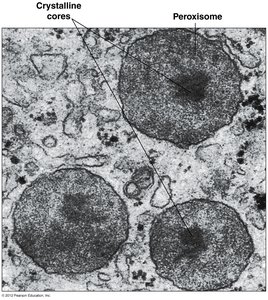
Hydrogen Peroxide Metabolism:
Superoxide dismutase generates H2O2
Catalase and peroxidase detoxify H2O2
Result: hydrogen peroxide is degraded inside the peroxisome
Fatty Acid Oxidation:
Animal cells: long-chain fatty acids oxidized in peroxisomes, shorter chains in mitochondria
Plants and yeast: complete oxidation in peroxisomes
Metabolism of Nitrogen-Containing Compounds:
Urate oxidase oxidizes urate from nucleic acid/protein catabolism
Catabolism of Unusual Substances:
D-amino acids and xenobiotics are degraded in peroxisomes
Includes breakdown of short-chain hydrocarbons
Additional info: These notes cover the structure, function, and sorting mechanisms of eukaryotic organelles, with emphasis on the ER, Golgi, lysosomes, vacuoles, and peroxisomes. The included images directly illustrate key concepts such as organelle structure, protein sorting, vesicle formation, and enzymatic functions.
