 Back
BackMeiosis, Sexual Reproduction, and Cell Cycle Regulation: Study Notes for Cell Biology
Study Guide - Smart Notes

Meiosis and Sexual Reproduction
The Principle of Meiosis
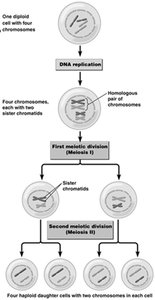
Meiosis is a specialized type of cell division that reduces the chromosome number by half, producing four haploid cells from one diploid cell. This process is essential for sexual reproduction and genetic diversity.
Reduction of Chromosome Number: Meiosis ensures gametes (egg and sperm) have half the chromosome number of somatic cells.
Diploid to Haploid: Begins with a diploid cell (2n) and ends with four haploid cells (n).
DNA Replication: One round of DNA replication precedes two successive cell divisions (Meiosis I and Meiosis II).
Meiosis I: Reduction division separates homologous chromosomes.
Meiosis II: Similar to mitosis, separates sister chromatids.
Result: Four genetically unique haploid cells.
Chromosome Structure and Karyotype
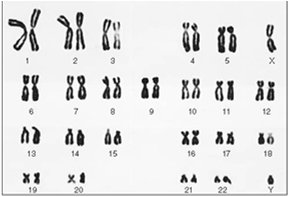
Chromosomes are thread-like structures carrying genetic information. A karyotype is the complete set of chromosomes in a cell, arranged by size and shape.
Homologous Chromosomes: Pairs of chromosomes with similar structure and gene content.
Karyotype Analysis: Used to identify chromosomal abnormalities and determine chromosome number.
Meiosis and Sexual Reproduction
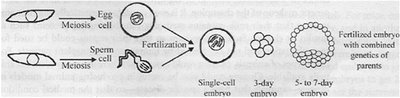
Sexual reproduction involves the fusion of two gametes (egg and sperm), each produced by meiosis, resulting in a zygote with a complete set of chromosomes.
Gametes: Haploid cells formed by meiosis.
Zygote: Diploid cell formed by fertilization.
Embryonic Development: Zygote undergoes mitotic divisions to form a multicellular embryo.
Organismal Cloning: Reproduction Without Meiosis
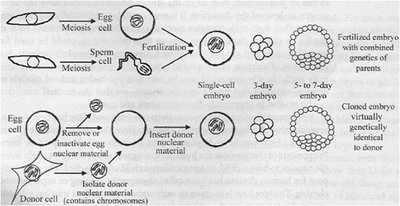
Cloning is a form of asexual reproduction where an organism is produced without meiosis or fertilization, resulting in a genetically identical copy of the donor.
Nuclear Transfer: Donor nucleus is inserted into an enucleated egg cell.
Development: The cloned embryo develops similarly to a fertilized embryo but is genetically identical to the donor.
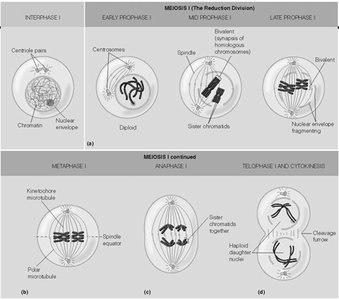
Stages of Meiosis I
Meiosis I is the first division in meiosis, reducing chromosome number and separating homologous chromosomes.
Interphase: Chromosomes replicate.
Prophase I: Homologous chromosomes pair and exchange genetic material (crossing over).
Metaphase I: Homologous pairs align at the cell equator.
Anaphase I: Homologous chromosomes separate.
Telophase I and Cytokinesis: Two haploid cells are formed.
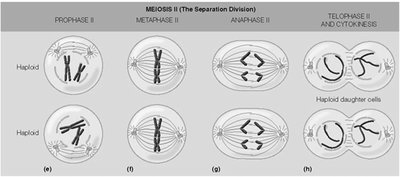
Stages of Meiosis II
Meiosis II resembles mitosis, separating sister chromatids in each haploid cell.
Prophase II: Chromosomes condense.
Metaphase II: Chromosomes align at the equator.
Anaphase II: Sister chromatids separate.
Telophase II and Cytokinesis: Four haploid daughter cells are produced.
Cellular Movement: The Sliding-Filament Model of Muscle Contraction
Sliding-Filament Model
The sliding-filament model explains how muscles contract by the sliding of actin and myosin filaments past each other.
Thick Filaments: Composed of myosin.
Thin Filaments: Composed of actin.
Contraction: Myosin heads bind to actin, pulling the filaments and shortening the sarcomere.
Comparison Table: Changes During Muscle Contraction
Item | Changed? |
|---|---|
Length of thick filaments | N |
Length of thin filaments | N |
Length of A band | N |
Overlapping of the filaments | Y |
Length of H zone | Y |
Length of I band | Y |
Length of Sarcomere | Y |
Cell Adhesion and Leukocyte Rolling
Leukocyte Rolling Arrest
Leukocyte rolling is a process by which white blood cells move along the vascular endothelium, mediated by selectins and integrins, and is crucial for immune response and inflammation.
Selectins: Mediate initial rolling of leukocytes.
Integrins: Activate firm adhesion and arrest.
ICAM: Intercellular adhesion molecule involved in firm attachment.
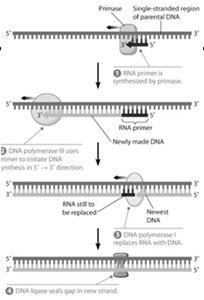
DNA Replication and Telomere Extension
Key Enzymes in DNA Replication
DNA replication is a complex process involving multiple enzymes to ensure accurate duplication of genetic material.
Helicase: Unwinds DNA helix.
Topoisomerase: Relieves supercoiling.
ssDNA-binding proteins: Stabilize single-stranded DNA.
Primase: Synthesizes RNA primers.
Polymerase: Synthesizes new DNA strands.
Ligase: Seals nicks in DNA.
Exonucleases: Remove incorrect bases.
Telomerase: Extends telomeres at chromosome ends.
Cell Cycle Regulation and DNA Damage Checkpoints
Cell Cycle Regulation: DNA Damage Checkpoint
The cell cycle is tightly regulated to ensure proper cell division. DNA damage checkpoints prevent the progression of the cell cycle in the presence of DNA damage.
p53 Protein: A transcription factor stabilized by DNA damage; activates p21 expression.
p21: Inhibits cyclin-Cdk complexes, arresting the cell cycle.
ATM: Ataxia telangiectasia mutated gene, involved in DNA damage response.
PUMA: p53 upregulated modulator of apoptosis, promotes cell death if damage is irreparable.
Growth Factor Signaling and Cell Cycle Entry
Epithelial growth factors provide positive signals for cells to pass the restriction point and enter the cell cycle, involving a cascade of signaling molecules.
MAPK Pathway: Includes MAPK, Raf, MEK, and Ras-GTP.
Rb-E2F: Rb protein regulates E2F transcription factor, controlling S-phase entry.
RTK: Receptor tyrosine kinase, initiates signaling.
Cyclin D-Cdk4/6: Promotes cell cycle progression.
EGF: Epidermal growth factor, stimulates pathway.
c-Jun: Nuclear transcription factor activated by MAPK.
Example Pathway Order: RTK → Ras-GTP → Raf → MEK → MAPK → c-Jun → Cyclin D-Cdk4/6 → Rb → E2F → S-phase entry
Additional info: The notes above expand on brief points with academic context, definitions, and examples to ensure completeness and clarity for cell biology students.
