 Back
BackMolecular Biology Techniques: DNA Hybridization, Fingerprinting, PCR, and Sequencing
Study Guide - Smart Notes

Principles of DNA Analysis and Manipulation
DNA Denaturation and Renaturation
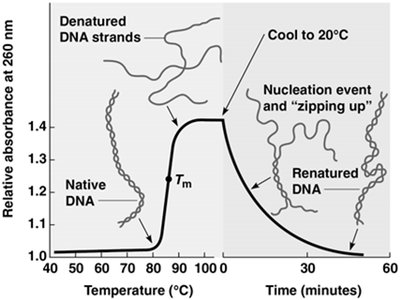
DNA consists of two complementary strands held together by noncovalent hydrogen bonds. These bonds can be disrupted by increasing temperature or altering pH, leading to strand separation (denaturation). The process is reversible; lowering the temperature allows the strands to re-anneal (renaturation), a principle underlying many molecular biology techniques.
Denaturation: The separation of double-stranded DNA into single strands by breaking hydrogen bonds, typically by heating.
Renaturation (Reannealing): The process by which complementary single-stranded DNA molecules reassociate to form double-stranded DNA when conditions return to normal.
UV Absorbance: Single-stranded DNA absorbs more UV light at 260 nm than double-stranded DNA, allowing monitoring of denaturation/renaturation.
Melting Temperature (Tm): The temperature at which half of the DNA molecules are denatured. Tm increases with higher GC content due to stronger hydrogen bonding.
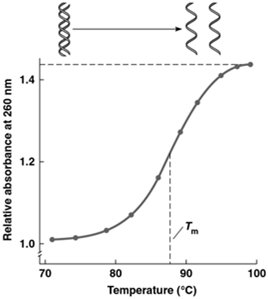
Example: DNA with 60% GC content will have a higher Tm than DNA with 40% GC content.
Membrane Transition Temperature (Comparison)
While Tm is used for DNA melting, it is also used in membrane biology to describe the temperature at which a membrane transitions from a gel-like to a fluid state. This is determined by the lipid composition of the membrane.
Unsaturated fatty acids lower Tm and increase membrane fluidity.
Saturated fatty acids raise Tm and decrease membrane fluidity.
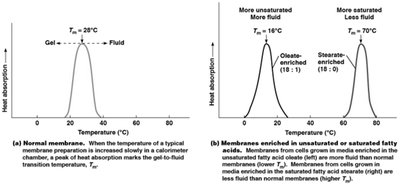
Additional info: This comparison helps clarify the use of Tm in both nucleic acid and membrane studies.
DNA Hybridization and Nucleic Acid Detection
Hybridization is the process by which single-stranded nucleic acids (DNA or RNA) form double-stranded molecules by base pairing with complementary sequences. This principle is used in several detection techniques:
FISH (Fluorescent In Situ Hybridization): Uses fluorescently labeled probes to detect specific DNA or RNA sequences in cells or tissues.
Southern Blotting: Uses radioactively or chemically labeled probes to detect specific DNA fragments separated by gel electrophoresis.
Restriction Enzymes and DNA Mapping
Restriction Enzymes and DNA Cleavage
Restriction enzymes (restriction endonucleases) are proteins that recognize specific DNA sequences (restriction sites) and cleave the DNA at or near these sites. The resulting fragments are called restriction fragments.
Blunt Ends: Produced when enzymes cut both DNA strands at the same position.
Sticky Ends: Produced when enzymes cut in a staggered manner, leaving overhanging single-stranded ends.
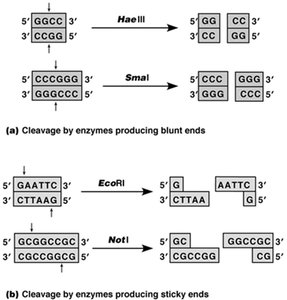
Example: EcoRI produces sticky ends, while HaeIII produces blunt ends.
Restriction Mapping
Restriction mapping involves cutting DNA with one or more restriction enzymes and analyzing the resulting fragment sizes by gel electrophoresis. This allows determination of the relative positions of restriction sites on a DNA molecule.
Single and double digests with different enzymes help construct a map of restriction sites.
Electrophoresis separates fragments by size, allowing comparison of patterns.
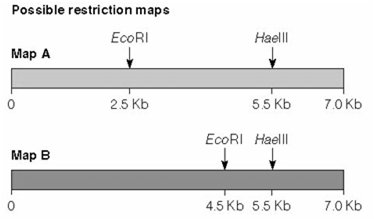
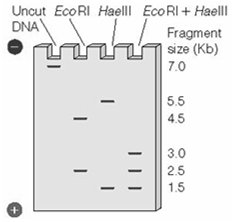
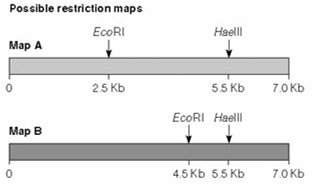
Example: Double digestion with EcoRI and HaeIII can distinguish between two similar DNA samples based on fragment patterns.
DNA Fingerprinting
Principle and Application
DNA fingerprinting uses restriction digestion and Southern blotting to generate unique patterns of DNA fragments for individual identification. Differences in DNA sequence among individuals result in different fragment lengths when digested with restriction enzymes.
Probes specific to certain DNA regions reduce the complexity of the pattern.
Used in forensic science, paternity testing, and genetic studies.
Example: DNA from a crime scene can be matched to a suspect or victim by comparing fingerprint patterns.
Polymerase Chain Reaction (PCR)
Principle and Steps
PCR is a technique for amplifying specific DNA sequences in vitro. It requires knowledge of the target sequence to design primers that flank the region of interest.
Denaturation: DNA is heated to separate strands.
Annealing: Primers bind to complementary sequences on the single-stranded DNA.
Extension: DNA polymerase synthesizes new DNA from the primers.
Repeated cycles (typically 30) result in exponential amplification of the target sequence.
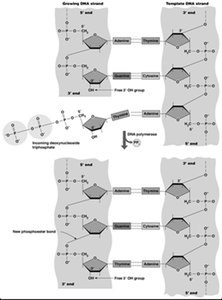
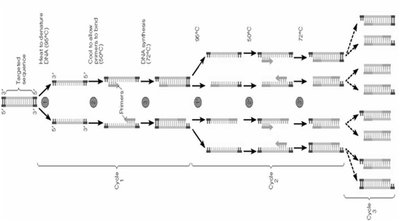
Example: PCR can amplify a single DNA molecule to billions of copies in a few hours.
DNA Sequencing
Sanger (Dideoxy) Sequencing
The Sanger method determines the order of nucleotides in DNA by incorporating chain-terminating dideoxynucleotides (ddNTPs) during DNA synthesis. Each ddNTP is labeled with a different fluorescent dye, allowing detection of the terminal base of each fragment.
DNA synthesis is performed in the presence of normal dNTPs and labeled ddNTPs.
Incorporation of a ddNTP terminates synthesis, producing fragments of varying lengths.
Fragments are separated by capillary electrophoresis, and the sequence is read by detecting the fluorescent labels.
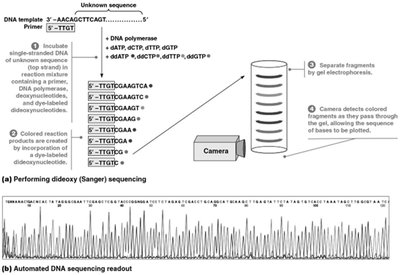
Example: Automated sequencers can rapidly determine the sequence of large DNA molecules.
Pyrosequencing and Next-Generation Sequencing (NGS)
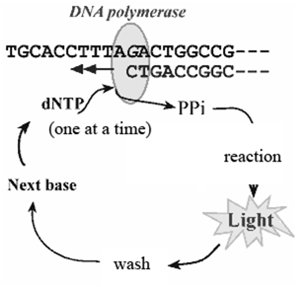
Pyrosequencing is a sequencing-by-synthesis method where the incorporation of a nucleotide releases pyrophosphate, which triggers a light-producing reaction. NGS technologies allow millions of sequencing reactions to occur in parallel, greatly increasing throughput.
Each nucleotide addition is detected by a flash of light.
Massive parallel processing enables rapid sequencing of entire genomes.
Bioinformatics and DNA Databases
Sequence Analysis Tools
Bioinformatics uses computational tools to analyze DNA and protein sequences. The National Center for Biotechnology Information (NCBI) provides databases and tools such as BLAST (Basic Local Alignment Search Tool) for sequence comparison.
BLASTn: Compares a nucleotide sequence to nucleotide databases.
BLASTp: Compares a protein sequence to protein databases.
Example: BLAST can identify homologous genes in different organisms, aiding in gene discovery and evolutionary studies.
