 Back
BackProtein Synthesis and Processing: Translation and Regulation
Study Guide - Smart Notes

Protein Synthesis & Processing
Overview of Translation
Translation is the process by which the genetic code carried by mRNA is decoded to produce a specific polypeptide, or protein. This process is central to gene expression and involves multiple steps and molecular components.
Gene Expression: Involves transcription (DNA to RNA), RNA processing, translation (RNA to protein), and post-translational modifications.
Functional Product: The end result is an active protein, often an enzyme, that performs cellular functions.
Key Steps: Transcription, RNA processing (splicing, capping, base modification), translation, folding, and further processing (lipid addition, glycosylation, peptide cleavage).
Structure of mRNA and Translation Initiation
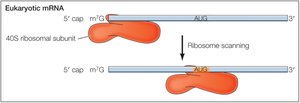
mRNA contains untranslated regions (UTRs) at both the 5' and 3' ends, which play important roles in translation regulation and ribosome binding. Translation typically begins at the AUG start codon, which is not at the very 5' end but after the 5' UTR.
5' UTR: Contains signals that direct ribosome binding and localization of the start codon.
Translation Start Site: The AUG codon marks the beginning of the protein coding sequence.
UTR: Abbreviation for untranslated region.
Components of Translation
Key Molecular Players
Translation requires several components, each with a distinct role:
mRNA: Carries the genetic code, read 5' to 3', specifying each amino acid with a three-base codon.
tRNA: Serves as an adaptor between the mRNA codon and the amino acid being incorporated. Each tRNA has a unique anticodon and an amino acid attachment site.
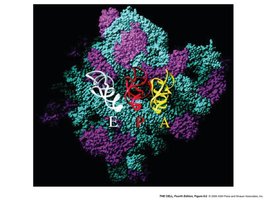
Ribosome: The site and machinery of protein synthesis, composed of large and small subunits.
Non-ribosomal Protein Factors: Facilitate initiation, elongation, and termination of translation.
Translation Factors Table
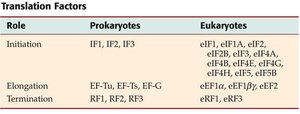
Translation factors are essential for the regulation and efficiency of protein synthesis. The table below compares the factors involved in prokaryotic and eukaryotic translation:
Role | Prokaryotes | Eukaryotes |
|---|---|---|
Initiation | IF1, IF2, IF3 | eIF1, eIF1A, eIF2, eIF2B, eIF3, eIF4A, eIF4B, eIF4E, eIF4G, eIF4H, eIF5, eIF5B |
Elongation | EF-Tu, EF-Ts, EF-G | eEF1α, eEF1βγ, eEF2 |
Termination | RF1, RF2, RF3 | eRF1, eRF3 |
tRNA Structure and Function
tRNA Structure
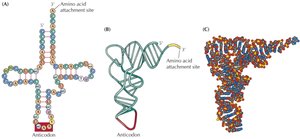
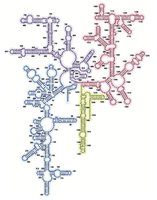
tRNAs are 70-80 nucleotides long and adopt a cloverleaf or L-shaped structure. Each tRNA has a unique anticodon and a CCA sequence at the 3' end for amino acid attachment.
Anticodon Loop: Recognizes the mRNA codon.
Adaptor Function: Links the genetic code to the amino acid sequence.
Aminoacyl tRNA Synthetases
These enzymes recognize both the amino acid and the tRNA anticodon, catalyzing the attachment of the correct amino acid to its tRNA. This process requires ATP and results in a "charged" tRNA.
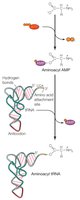
Codon Recognition and Wobble
Codon-anticodon pairing is not always strict; the third base (wobble position) allows for flexibility, enabling one tRNA to recognize multiple codons.
Wobble Base: Allows non-standard base pairing, increasing efficiency.
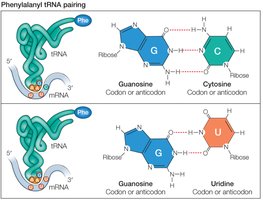
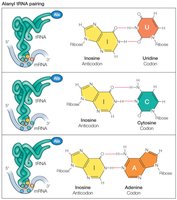
Ribosome Structure and Function
Ribosome Subunits
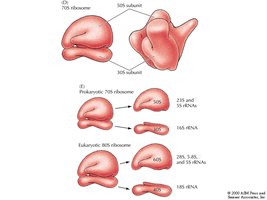
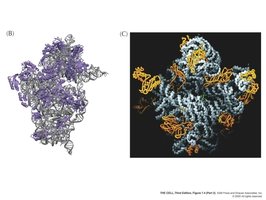
Ribosomes are composed of a large and a small subunit. Prokaryotic and eukaryotic ribosomes are similar in overall structure but differ in details, such as rRNA composition and size.
Prokaryotic Ribosome: 70S (50S + 30S)
Eukaryotic Ribosome: 80S (60S + 40S)
rRNA: Large subunit rRNA acts as a peptidyl transferase (23S in prokaryotes, 28S in eukaryotes), functioning as a ribozyme.
Process of Translation
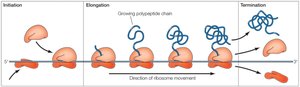
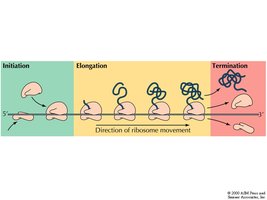
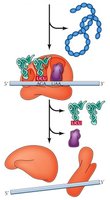
Translation Steps: Initiation, Elongation, Termination
Translation occurs in three main stages: initiation, elongation, and termination. Each stage involves specific factors and mechanisms.
Initiation: Assembly of the ribosome on the mRNA, recognition of the start codon, and binding of the initiator tRNA.
Elongation: Sequential addition of amino acids to the growing polypeptide chain.
Termination: Release of the completed polypeptide when a stop codon is encountered.
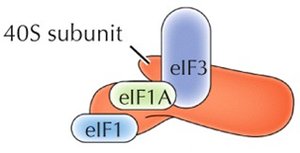
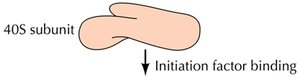
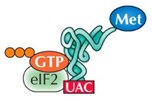
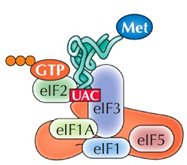
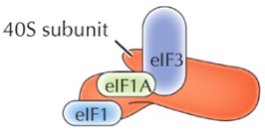
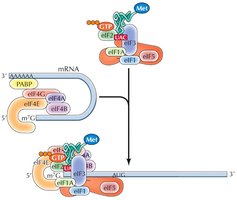
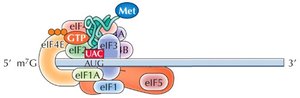
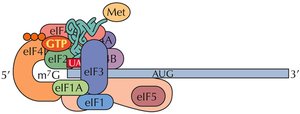
Initiation in Eukaryotes
The small ribosomal subunit (40S) binds to the mRNA's 5' cap and scans for the AUG start codon. Initiation factors (eIFs) and a "charged" initiator tRNA (tRNAimet) are required. Once the start codon is located, eIFs are released and the large subunit binds.
Elongation
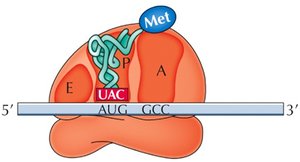
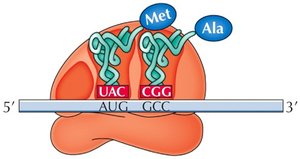
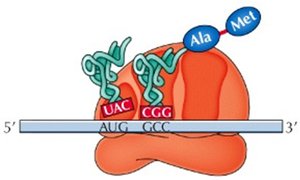
The ribosome has three sites for tRNA binding: P (peptidyl), A (aminoacyl), and E (exit). The initiator tRNA binds the P site, and subsequent tRNAs enter the A site. Peptide bonds form, and the ribosome translocates along the mRNA, requiring GTP for energy.
P Site: Holds the tRNA with the growing peptide chain.
A Site: Accepts the incoming aminoacyl-tRNA.
E Site: Exit site for uncharged tRNA.
Termination
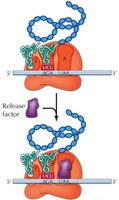
When a stop codon is reached, no tRNA matches it. Instead, release factors (RFs) bind, stimulating hydrolysis of the bond between tRNA and the polypeptide, releasing the completed protein and dissociating the ribosome.
Regulation of Translation
Translation Initiation Regulation
Translation initiation is tightly regulated by signals in the mRNA and by various mechanisms, including global and specific regulation. The efficiency of initiation can be affected by sequences surrounding the AUG codon (e.g., Kozak sequence in eukaryotes).
Polycistronic vs. Monocistronic mRNA
Prokaryotic mRNAs are often polycistronic, containing multiple translation start sites, while eukaryotic mRNAs are typically monocistronic, with a single translation start site.
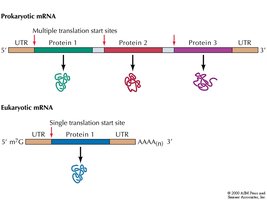
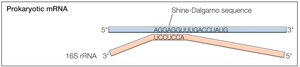
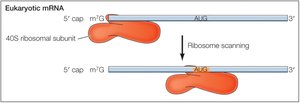
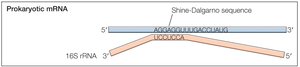
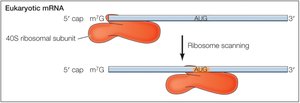
Localization of AUG: Prokaryotes vs. Eukaryotes
In prokaryotes, the Shine-Dalgarno sequence aligns the ribosome with the start codon. In eukaryotes, the small ribosomal subunit binds the 5' cap and scans for the AUG codon, with initiation efficiency influenced by surrounding sequences.
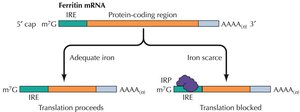
Regulation by Noncoding RNAs and Translational Repressors
Noncoding RNAs (miRNA, siRNA) can cause degradation of mRNA or inhibit translation. Translational repressor proteins, such as those regulating ferritin, can block translation in response to cellular signals (e.g., iron availability).
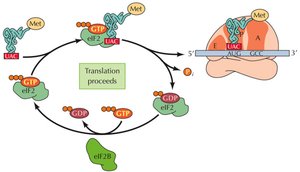
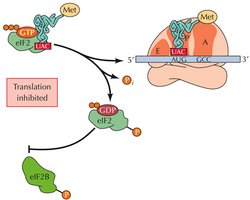
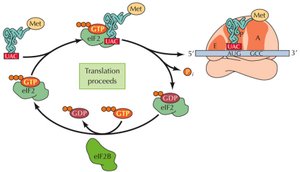
Global Regulation via Initiation Factors
Phosphorylation of initiation factors (e.g., eIF-2) can globally regulate translation, especially in response to stress or nutrient deprivation. Phosphorylation blocks GDP/GTP exchange, preventing recycling of eIF-2 and inhibiting translation.
Summary Table: Translation Steps and Regulation
Step | Key Components | Regulation |
|---|---|---|
Initiation | mRNA, ribosome, eIFs, tRNAimet | mRNA signals, eIF phosphorylation, repressor proteins |
Elongation | Ribosome, tRNAs, elongation factors | Energy (GTP), codon-anticodon pairing |
Termination | Release factors, ribosome | Stop codon recognition |
Additional info: The notes expand on brief slide points to provide full academic context, including definitions, examples, and regulatory mechanisms. All images included are directly relevant to the adjacent explanations, reinforcing key concepts in translation and its regulation.
