Back
BackThe Nucleus: Structure, Function, and Genetic Information Flow
Study Guide - Smart Notes

The Flow of Genetic Information in Cells
Overview of Genetic Information Flow
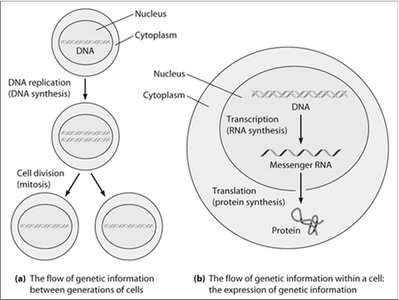
The flow of genetic information in eukaryotic cells involves the maintenance and utilization of DNA within the nucleus. Genetic information is replicated and passed to daughter cells during cell division, and is also expressed through transcription and translation to produce proteins essential for cellular function.
DNA Replication: The process by which DNA is copied before cell division, ensuring genetic continuity.
Transcription: The synthesis of RNA from a DNA template within the nucleus.
Translation: The process by which messenger RNA (mRNA) is decoded in the cytoplasm to synthesize proteins.
The Structure and Components of the Nucleus
Nuclear Envelope and Associated Structures
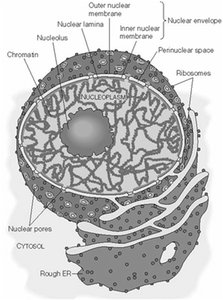
The nucleus is surrounded by a double-membrane structure called the nuclear envelope, which separates the nucleoplasm from the cytoplasm. The outer membrane is continuous with the rough endoplasmic reticulum (ER), while the inner membrane is connected to the nuclear lamina, a supportive protein network.
Nuclear Envelope: Composed of inner and outer membranes, with the outer membrane continuous with the rough ER.
Nuclear Pore Complexes (NPCs): Large protein assemblies that regulate the exchange of molecules between the nucleus and cytoplasm.
Nucleoplasm: The semi-fluid matrix inside the nucleus, containing chromatin and the nuclear matrix.
Nucleolus: A prominent subnuclear structure involved in ribosomal RNA (rRNA) synthesis and ribosome assembly.
The Nuclear Matrix and Nuclear Lamina
Structural Support and Organization
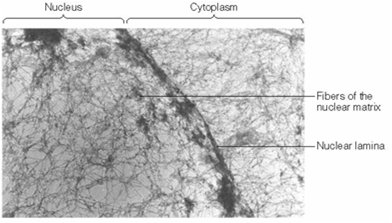
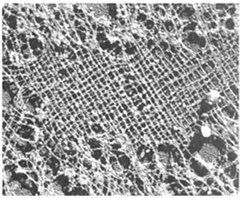
The nuclear matrix (nucleoskeleton) is a three-dimensional fibrous network that provides structural support and organizes the contents of the nucleus. The nuclear lamina, composed of intermediate filament proteins called lamins (A, B, C), forms a thin layer beneath the inner nuclear membrane and is essential for nuclear stability and chromatin organization.
Nuclear Matrix: Insoluble network remaining after chromatin extraction; organizes nuclear processes.
Nuclear Lamina: Supports the nuclear envelope and anchors chromatin; mutations in lamins can cause diseases such as progeria.

Genetic Diseases Related to Nuclear Structure
Progeria and Laminopathies
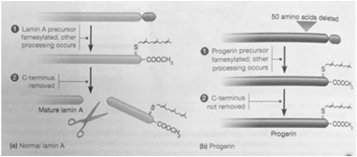
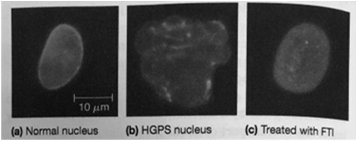
Mutations in nuclear lamina proteins can lead to diseases known as laminopathies. One example is Hutchinson-Gilford Progeria Syndrome (HGPS), a premature aging disorder caused by a mutation in lamin A, resulting in abnormal nuclear morphology and impaired cellular function.
Progeria: Characterized by rapid aging due to defective lamin A processing.
Laminopathies: A group of disorders caused by mutations in lamins or associated proteins.
The Nucleolus
Structure and Function
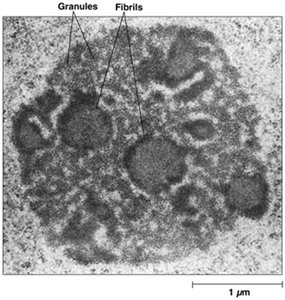
The nucleolus is a large, spherical, non-membranous organelle within the nucleus, formed by liquid-liquid phase separation. It is the site of rRNA synthesis, processing, and assembly of ribosomal subunits. Nucleoli form around nucleolar organizing regions (NORs) containing rDNA repeats.
rRNA Synthesis: 28S, 18S, and 5.8S rRNAs are transcribed and processed in the nucleolus.
Ribosome Assembly: Ribosomal proteins imported from the cytoplasm are assembled with rRNA to form ribosomal subunits.
Chromatin Organization in the Nucleus
Chromosome Territories and Chromatin Structure
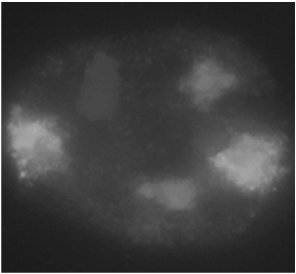
Chromatin fibers are dispersed throughout the nucleus in a nonrandom fashion, with each chromosome occupying a distinct territory. This organization is essential for gene regulation and efficient DNA replication and repair.
Chromosome Territories: Discrete regions occupied by individual chromosomes.
In Situ Hybridization: Technique used to visualize specific DNA sequences within the nucleus.
Nuclear Pores and Nucleocytoplasmic Transport
Structure and Function of Nuclear Pore Complexes (NPCs)
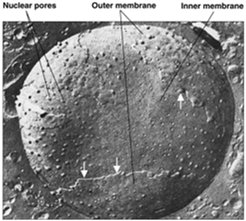
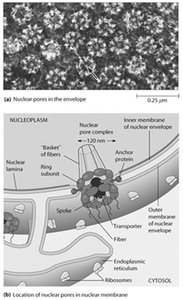
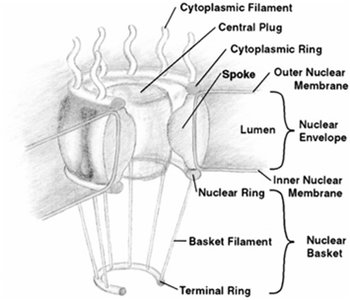
Nuclear pores are large protein complexes embedded in the nuclear envelope, facilitating selective exchange of macromolecules between the nucleus and cytoplasm. Each NPC is composed of multiple nucleoporins (Nups) arranged in an octagonal symmetry, with a central transporter and associated rings and fibers.
Number of NPCs: 3000–4000 per nucleus.
Diameter: ~120 nm.
Function: Regulate import and export of proteins, RNA, and ribonucleoprotein complexes.
Macromolecular Transport into and out of the Nucleus
Nuclear Import and Export Mechanisms
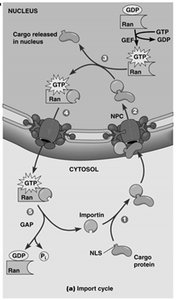
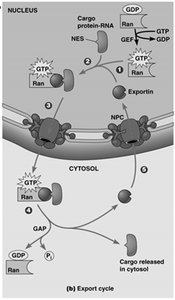
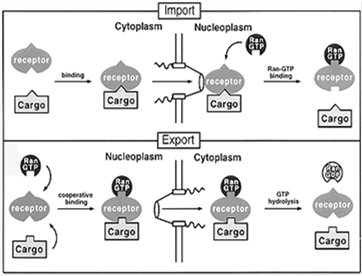
Transport through the NPC is highly regulated and involves specific signal sequences and transport receptors. Import and export are mediated by nuclear localization signals (NLS) and nuclear export signals (NES), respectively, and require energy provided by the Ran GTPase system.
Nuclear Import: Proteins with NLS are recognized by importins, which facilitate their transport into the nucleus.
Nuclear Export: Proteins and RNAs with NES are recognized by exportins, which mediate their export to the cytoplasm.
Ran GTPase: Provides directionality to transport by cycling between GTP- and GDP-bound forms, regulated by RanGEF (nuclear) and RanGAP (cytoplasmic).
DNA Structure and Organization
The Double Helix and DNA Measurement
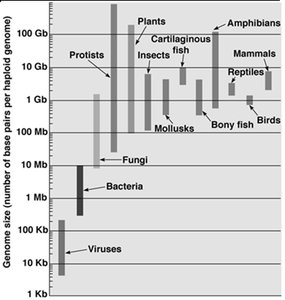
DNA is a double helix with antiparallel strands, held together by hydrogen bonds between complementary bases (A=T, G≡C). DNA length is measured in base pairs (bp), with larger units such as kilobase (Kb), megabase (Mb), and gigabase (Gb) used for longer sequences. The total DNA content of an organism is its genome, which varies in size among different organisms.
Base Pairing: Purine-pyrimidine pairing ensures uniform helix diameter.
Genome Size: Generally correlates with organismal complexity, but with exceptions.
DNA Supercoiling and Topoisomerases
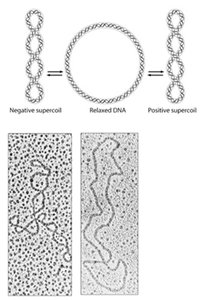
Relaxed and Supercoiled DNA Forms
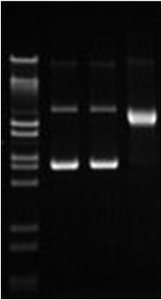
DNA can exist in relaxed or supercoiled forms. Supercoiling compacts DNA and is regulated by enzymes called topoisomerases. Positive supercoiling twists DNA in the same direction as the helix, while negative supercoiling twists it in the opposite direction. Topoisomerases introduce transient breaks to relieve or introduce supercoils, which is essential during DNA replication and transcription.
Type I Topoisomerases: Introduce single-strand breaks to relax supercoils.
Type II Topoisomerases: Introduce double-strand breaks; e.g., DNA gyrase in bacteria.