 Back
BackTranscription and the Genetic Code: Cell Biology Study Notes
Study Guide - Smart Notes

Transcription and the Central Dogma of Molecular Biology
Overview of the Central Dogma
The central dogma of molecular biology describes the directional flow of genetic information within a cell: from DNA to RNA to protein. This process involves two major steps: transcription (DNA to RNA) and translation (RNA to protein). While transcription involves converting one nucleic acid type to another, translation changes the "language" from nucleic acids to amino acids. Not all RNA is translated; non-coding RNAs regulate gene expression and form structural components.
Transcription: DNA is used as a template to synthesize RNA (mRNA, rRNA, tRNA).
Translation: mRNA is decoded to synthesize proteins.
Non-coding RNA: Includes regulatory and structural RNAs that do not code for proteins.


One Gene, One Polypeptide Hypothesis
Historical Experiments
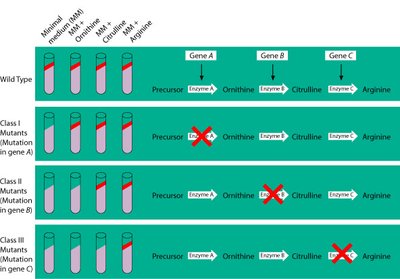
Early experiments by Beadle and Tatum using Neurospora crassa established the "one gene = one enzyme" hypothesis. Later, Pauling and Ingram showed that a single gene mutation could alter one amino acid in hemoglobin, leading to sickle cell anemia. This evolved into the "one gene = one polypeptide" concept, and further research revealed that a single gene can produce multiple polypeptides through alternative splicing.
Beadle & Tatum: Mutations in single genes disrupted specific enzymes.
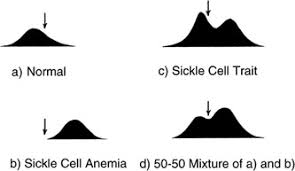
Pauling & Ingram: Electrophoresis showed altered migration of sickle cell hemoglobin due to a single amino acid change (Glu → Val).
One gene = multiple polypeptides: Alternative splicing allows one gene to produce several proteins.
Stages of Transcription
Four Main Stages
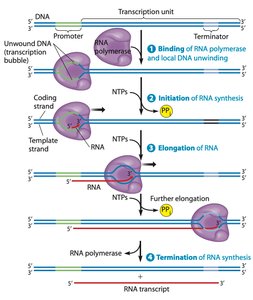
Transcription is divided into four stages: binding, initiation, elongation, and termination. Each stage is essential for accurate RNA synthesis.
Binding: RNA polymerase binds to promoter regions near the start of the gene.
Initiation: The DNA helix is unwound, and RNA synthesis begins with activated ribonucleotide triphosphates (rNTs).
Elongation: RNA polymerase moves along the DNA, adding rNTs to the growing RNA strand.
Termination: RNA polymerase detaches, and the RNA transcript is released.
Prokaryotic Transcription
RNA Polymerase and Promoters
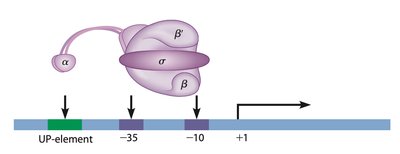
Prokaryotic transcription is simpler than eukaryotic transcription. The RNA polymerase consists of five subunits (2 α, 2 β, 1 σ). The core enzyme can synthesize RNA but cannot initiate transcription without the σ subunit. Promoter sequences, such as the -10 (TATAAT) and -35 (TTGACA) regions, are critical for RNA polymerase binding.
Core enzyme: 2 α + 2 β subunits.
σ subunit: Required for initiation.
Promoter sequences: -10 (TATAAT), -35 (TTGACA).
Initiation and Elongation
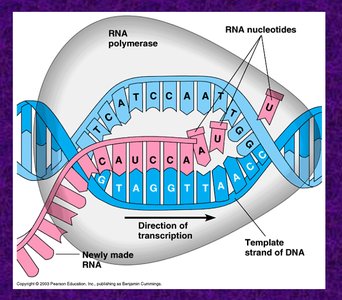
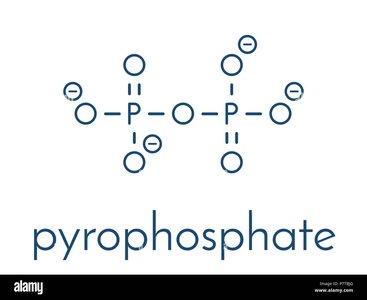
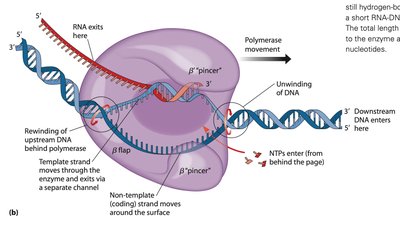
During initiation, the DNA helix is unwound, and activated rNTs are added. The σ subunit detaches after about nine rNTs are added, allowing the core enzyme to continue elongation. New rNTs are added to the 3' end of the RNA, and topoisomerase prevents DNA supercoiling.
Initiation: Helix unwinding, rNT addition, σ subunit detachment.
Elongation: Core enzyme adds rNTs, DNA is re-wound.
Termination
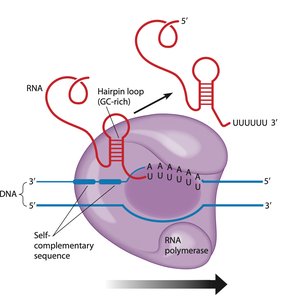
Termination occurs when RNA polymerase encounters specific signals. The Rho factor protein unwinds RNA from DNA in an ATP-dependent manner, or GC-rich sequences form a hairpin loop, causing the RNA to detach.
Rho factor: ATP-dependent unwinding.
GC-rich hairpin loop: Self-complementary sequences cause RNA release.
Eukaryotic Transcription
RNA Polymerases and Promoters
Eukaryotes have three nuclear RNA polymerases (I, II, III), each with distinct roles and promoter requirements. RNA polymerase II transcribes mRNA and requires multiple transcription factors for initiation. Promoter elements include the initiator sequence (Inr), TATA box, BRE, and DPE.
RNA polymerase I: rRNA synthesis.
RNA polymerase II: mRNA synthesis.
RNA polymerase III: tRNA and other small RNAs.
Promoter elements: Inr, TATA box, BRE, DPE.

Initiation Complex and Processing
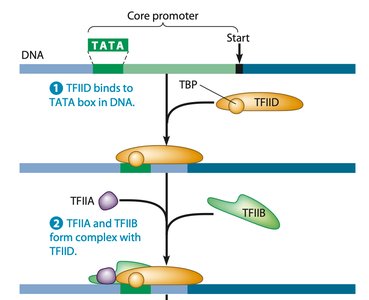
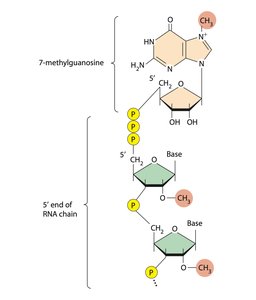
Transcription initiation in eukaryotes requires assembly of a pre-initiation complex, including RNA polymerase II and transcription factors (TFIID, TFIIB, etc.). After transcription, RNA undergoes processing: addition of a 5' cap, polyA tail, and splicing to remove introns.
Pre-initiation complex: RNA polymerase II + transcription factors.
RNA processing: 5' cap, polyA tail, splicing.
RNA Processing and Splicing
Introns, Exons, and Spliceosomes
Primary RNA transcripts contain both introns (non-coding regions) and exons (coding regions). Introns are removed by spliceosomes, large complexes of RNA and proteins. The GU-AG rule defines the splice sites. Alternative splicing allows for multiple polypeptides from a single gene.
Introns: Non-coding regions excised from RNA.
Exons: Coding regions joined to form mature RNA.
Spliceosomes: Catalyze intron removal.
Alternative splicing: Generates protein diversity.
The Genetic Code and Codons
Triplet Code and Reading Frames
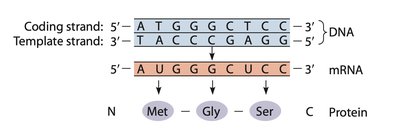
The genetic code is based on triplet codons, with three nucleotides coding for one amino acid. There are 64 possible codons, including start (AUG) and stop (UAA, UAG, UGA) signals. The code is degenerate, meaning multiple codons can specify the same amino acid. Reading frames are critical; frameshift mutations can disrupt protein synthesis.
Triplet code: Three nucleotides per codon.
Start codon: AUG (methionine).
Stop codons: UAA, UAG, UGA.
Degenerate code: Multiple codons for one amino acid.
Reading frames: Correct grouping of codons is essential.
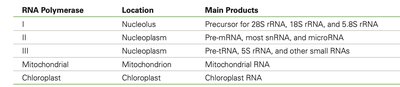
Summary Table: RNA Polymerases in Eukaryotes
This table summarizes the types, locations, and main products of eukaryotic RNA polymerases.
RNA Polymerase | Location | Main Products |
|---|---|---|
I | Nucleolus | Precursor for 28S rRNA, 18S rRNA, and 5.8S rRNA |
II | Nucleoplasm | Pre-mRNA, most snRNA, and microRNA |
III | Nucleoplasm | Pre-tRNA, 5S rRNA, and other small RNAs |
Mitochondrial | Mitochondrion | Mitochondrial RNA |
Chloroplast | Chloroplast | Chloroplast RNA |
Key Equations and Concepts
Base pairing during transcription:
Direction of RNA synthesis:
Triplet code: possible codons
Transcription Review
Binding: Promoters, start point
Initiation: RNA polymerase, "scrunching", small number of NT added
Elongation: RNA polymerase core, re-winding
Termination: Two signals
Processing: PolyA tail/capping, splicing
Additional info: These notes expand on lecture content with definitions, examples, and diagrams to provide a comprehensive overview of transcription and the genetic code for cell biology students.
