 Back
BackTranslation: From mRNA to Protein – Mechanisms and Molecular Players
Study Guide - Smart Notes

Translation: The Genetic Code to Protein Synthesis
Overview of Translation
Translation is the process by which the genetic information encoded in messenger RNA (mRNA) is used to assemble a specific sequence of amino acids, forming a polypeptide chain that folds into a functional protein. This process is the final step in the central dogma of molecular biology: DNA → RNA → Protein.
Translation occurs in the cytoplasm on ribosomes.
It involves decoding the mRNA sequence into a chain of amino acids using transfer RNAs (tRNAs) and various protein factors.
The genetic code is read in triplets called codons.
The Genetic Code
Codons and the Non-Overlapping Code
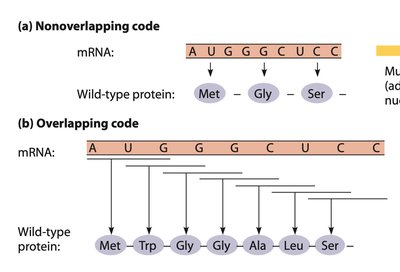
The genetic code consists of 64 possible codons, each made up of three nucleotides. Each codon specifies an amino acid or a stop signal during translation. The code is degenerate, meaning that multiple codons can code for the same amino acid, and it is non-overlapping, so each nucleotide is part of only one codon.
Codons: Triplets of nucleotides in mRNA that specify amino acids.
Non-overlapping: Each nucleotide is read only once in a single reading frame.
Degeneracy: More than one codon can specify the same amino acid.
Start and Stop Codons
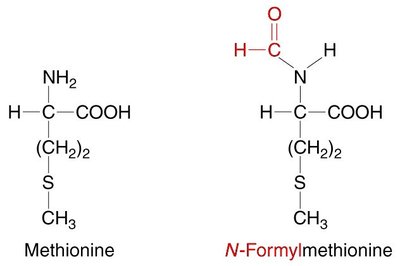
Start codon (AUG): Codes for methionine (Met) and signals the start of translation.
Stop codons (UAA, UAG, UGA): Signal the end of translation; do not code for amino acids.
Key Molecular Players in Translation
Ribosomes
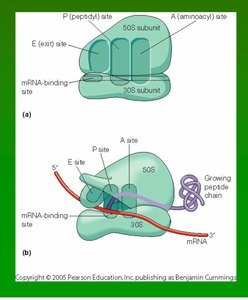
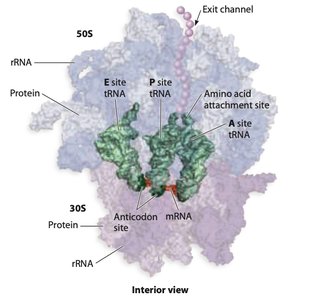
Ribosomes are large ribonucleoprotein complexes composed of ribosomal RNA (rRNA) and proteins. They consist of two subunits (large and small) and contain three main binding sites for tRNA: the A (aminoacyl), P (peptidyl), and E (exit) sites.
Bacterial ribosome (70S): Large subunit (50S) + small subunit (30S)
Eukaryotic ribosome (80S): Large subunit (60S) + small subunit (40S)
Four important sites: mRNA binding site, A site, P site, E site
Transfer RNA (tRNA)
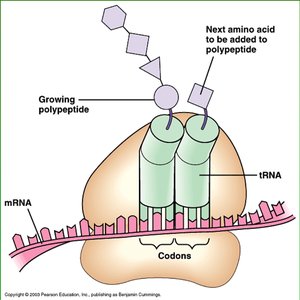
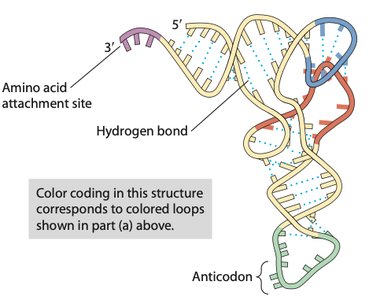
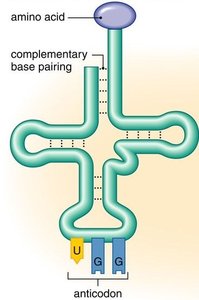
tRNAs are adaptor molecules that bring specific amino acids to the ribosome. Each tRNA has an anticodon region that base-pairs with a complementary codon on the mRNA and an acceptor stem where the corresponding amino acid is attached.
Anticodon: A sequence of three nucleotides complementary to the mRNA codon.
Amino acid attachment site: Always at the 3' end of the tRNA, ending with an adenine nucleotide.
Charged tRNA: tRNA with an amino acid attached (also called aminoacyl-tRNA).
Aminoacyl-tRNA Synthetases
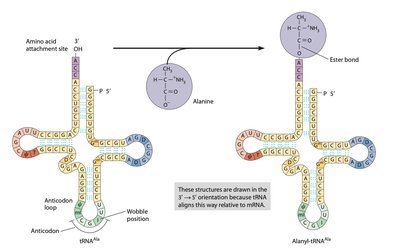
These enzymes catalyze the attachment of amino acids to their corresponding tRNAs, forming aminoacyl-tRNAs. There are usually 20 different synthetases, one for each amino acid. The reaction is highly specific and involves ATP hydrolysis.
Specificity: Each synthetase recognizes one amino acid and its compatible tRNAs.
Reaction: Amino acid + tRNA + ATP → aminoacyl-tRNA + AMP + PPi
Proofreading: Synthetases have proofreading activity to ensure correct amino acid attachment.
Codon-Anticodon Pairing and the Wobble Hypothesis
Anticodon-Codon Interaction
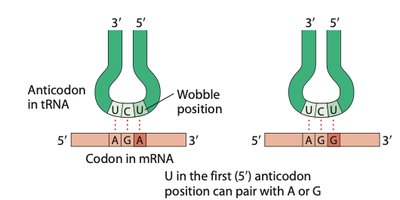
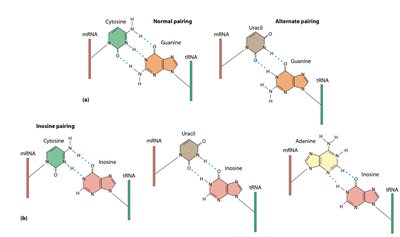
The anticodon of tRNA pairs with the codon of mRNA in an antiparallel fashion. The first base of the anticodon (5' end) pairs with the third base of the codon (3' end), which is known as the wobble position.
Wobble hypothesis: Flexibility in base pairing at the third codon position allows some tRNAs to pair with more than one codon.
Inosine (I): A modified base in tRNA that can pair with U, C, or A in the mRNA.
Steps of Translation
1. Initiation
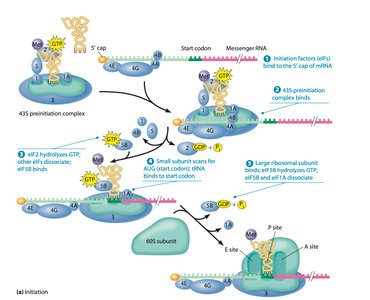
Initiation involves the assembly of the translation machinery at the start codon of the mRNA. In prokaryotes, the Shine-Dalgarno sequence helps position the ribosome, while in eukaryotes, the 5' cap structure is recognized by initiation factors.
Prokaryotes: Initiation factors (IF1, IF2, IF3), fMet-tRNA, Shine-Dalgarno sequence
Eukaryotes: eIFs, Met-tRNA, 5' cap structure
Small ribosomal subunit binds mRNA, then the initiator tRNA, then the large subunit joins
2. Elongation
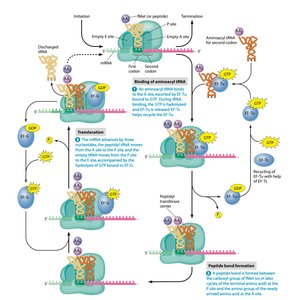
During elongation, amino acids are added one by one to the growing polypeptide chain. Each cycle involves binding of an aminoacyl-tRNA to the A site, peptide bond formation, and translocation of the ribosome along the mRNA.
Peptide bond formation: Catalyzed by the ribosome (ribozyme activity of rRNA).
Translocation: Ribosome moves three nucleotides along mRNA, shifting tRNAs between sites.
Elongation factors: EF-Tu (prokaryotes) or eEF1A (eukaryotes) help deliver aminoacyl-tRNAs.
3. Termination
Termination occurs when a stop codon enters the A site. Release factors bind to the stop codon, prompting the ribosome to release the completed polypeptide and dissociate from the mRNA.
Stop codons: UAA, UAG, UGA
Release factors: Proteins that mimic tRNA structure and trigger hydrolysis of the bond between the polypeptide and tRNA.
Energy Requirements of Translation
Translation is an energy-intensive process, requiring the hydrolysis of multiple high-energy phosphate bonds for each amino acid added to the polypeptide chain. ATP and GTP are consumed during tRNA charging, initiation, elongation, and termination.
At least 4 phosphate bonds are hydrolyzed per amino acid addition.
Additional energy is used for proofreading and assembly of translation components.
Molecular Chaperones
Molecular chaperones are proteins that assist in the proper folding of newly synthesized polypeptides and prevent misfolding or aggregation. Some chaperones act during translation, while others function after the polypeptide is released from the ribosome.
Ensure correct secondary and tertiary structure formation.
Prevent non-specific interactions between nascent polypeptides.
Summary Table: Key Steps and Components of Translation
Step | Main Components | Key Events |
|---|---|---|
Initiation | Small and large ribosomal subunits, mRNA, initiator tRNA, initiation factors | Assembly of initiation complex at start codon |
Elongation | Elongation factors, aminoacyl-tRNAs, ribosome | Sequential addition of amino acids, peptide bond formation, translocation |
Termination | Release factors, ribosome, mRNA | Recognition of stop codon, release of polypeptide, ribosome dissociation |
Key Equations
Aminoacyl-tRNA formation:
Additional info: The notes above integrate and expand upon the provided lecture outline, filling in details about the molecular mechanisms, the roles of each translation component, and the energy requirements. The images included are directly relevant to the described processes and structures, reinforcing the textual explanations.
