 Back
BackChromosome Mapping in Eukaryotes: Linkage, Recombination, and Genetic Mapping
Study Guide - Smart Notes

Chromosome Mapping in Eukaryotes
Introduction to Linkage and Chromosome Mapping
Chromosome mapping is a fundamental technique in genetics used to determine the relative positions of genes on a chromosome. Because there are more genes than chromosomes, some genes are inherited together, a phenomenon known as linkage. Linked genes do not assort independently, and their inheritance patterns can be used to estimate their physical proximity on a chromosome.
Linkage and Crossing Over
Genes located on the same chromosome are said to be linked. In genetic crosses, linked genes may demonstrate complete linkage (no crossing over) or incomplete linkage (crossing over occurs). Crossing over between two linked genes occurs between nonsister chromatids during meiosis, producing both parental and recombinant gametes. This process increases genetic variability.
Complete Linkage: Only parental (noncrossover) gametes are produced.
Crossing Over: Both parental and recombinant (crossover) gametes are produced.
Significance: Crossing over is a major source of genetic diversity.
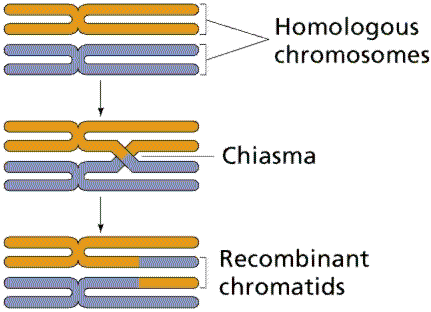
Chiasmata and Genetic Exchange
During meiosis, homologous chromosomes pair and form chiasmata, which are points of genetic exchange. The likelihood of crossing over depends on the distance between genes (interlocus distance):
The closer two genes are, the less likely crossing over will occur between them.
The farther apart two genes are, the more likely crossing over will occur.
Genetic Crosses and Linkage
Genetic crosses can reveal linkage relationships. For example, in a cross between AAbb and aaBB, if A and B are closely linked and no crossing over occurs, only parental types are produced. In contrast, unlinked genes follow Mendelian ratios (9:3:3:1 in the F2 generation).
Parental Types: Progeny with the same phenotype as one of the parents.
Recombinants: Progeny with phenotypes different from either parent.
Chromosome Mapping and Recombination Frequency
The frequency of genetic exchange (recombination) between two genes is an estimate of their relative distance on a chromosome. Recombination frequencies are additive and can be used to predict gene order.
Map Unit (mu): Defined as 1% recombination between two genes.
Centimorgan (cM): Another term for map unit; represents relative, not absolute, distance.
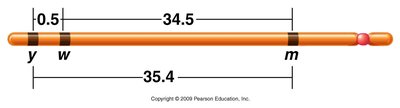
Example: Given recombination frequencies between three genes (yellow, white, miniature):
yellow-white: 0.5%
white-miniature: 34.5%
yellow-miniature: 35.4%
Single Crossover Events
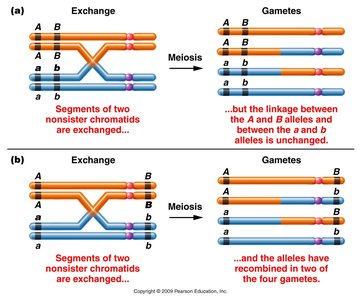
A single crossover (SCO) alters linkage between two genes only if the crossover occurs between those genes. If the exchange does not separate the alleles, only parental gametes are formed. When a single crossover occurs between two nonsister chromatids, the other two chromatids remain unchanged.
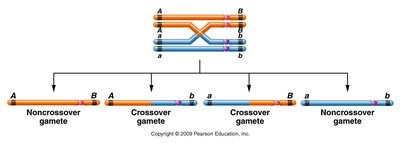
Phenotypic Ratios in Testcrosses
Testcrosses involving two genes (a and b) can yield different phenotypic ratios depending on their linkage:
Separate autosomes: Independent assortment, Mendelian ratios.
Linked, far apart: Crossover always occurs, producing recombinant types.
Linked, close together: Crossover rarely occurs, mostly parental types.
Linked, 10 mu apart: Intermediate ratio of parental and recombinant types.
Why Didn’t Gregor Mendel Find Linkage?
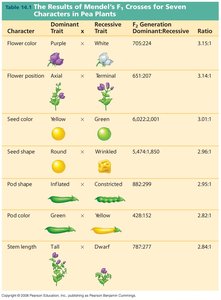
Mendel studied seven genes in pea plants, and the pea has seven chromosomes. The genes Mendel chose were either on different chromosomes or so distantly located on the same chromosome that linkage was not detected. This allowed Mendel to observe independent assortment and develop his laws.
Key Terms and Concepts
Linkage: The tendency of genes located on the same chromosome to be inherited together.
Recombination: The process by which genetic material is exchanged between chromatids during meiosis.
Chiasma: The physical site of crossing over between homologous chromosomes.
Map Unit (mu) / Centimorgan (cM): A unit of measure for genetic distance, defined as 1% recombination frequency.
Parental and Recombinant Types: Parental types resemble the original parents; recombinant types result from crossing over.
Formulas and Equations
Recombination Frequency:
Map Distance:
Summary Table: Linkage and Recombination
Condition | Phenotypic Ratio | Explanation |
|---|---|---|
Unlinked genes | 9:3:3:1 | Independent assortment |
Complete linkage | 1:2:1 | No crossing over, only parental types |
Linked, far apart | ~1:1:1:1 | Crossover always occurs, recombinant types produced |
Linked, close together | Mostly parental | Crossover rarely occurs |
Example: In a testcross between a heterozygote and a double homozygote mutant, the phenotypic ratios depend on the linkage and distance between the genes.
Additional info: The notes above expand on the original content by providing definitions, formulas, and a summary table for clarity and completeness.
