 Back
BackChromosome Mapping in Eukaryotes: Linkage, Recombination, and Genetic Distance
Study Guide - Smart Notes

Chromosome Mapping in Eukaryotes
Introduction to Linkage and Chromosome Mapping
Chromosome mapping is a fundamental technique in genetics used to determine the relative positions of genes on a chromosome. Genes that are located on the same chromosome tend to be inherited together, a phenomenon known as linkage. The study of linkage and recombination frequencies allows geneticists to construct genetic maps, which are crucial for understanding inheritance patterns and gene interactions.
Linkage and Independent Assortment
Independent Assortment: Genes located on different chromosomes assort independently during meiosis, resulting in a variety of genetic combinations in gametes.
Linkage: Genes that are located close together on the same chromosome tend to be inherited together and do not assort independently. These genes are said to be linked.
Complete Linkage: In rare cases where two genes are so close together that crossing over does not occur between them, only parental (nonrecombinant) gametes are produced.
Recombination: If crossing over occurs between two linked genes, both parental and recombinant gametes are produced, increasing genetic variability.
Mechanism of Crossing Over
During meiosis, homologous chromosomes pair and may exchange segments in a process called crossing over. The physical point where this exchange occurs is known as a chiasma. Crossing over results in recombinant chromatids, which carry new combinations of alleles.
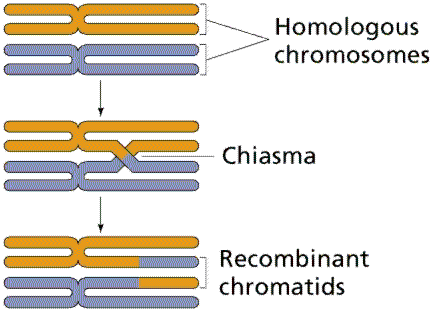
Chiasmata: The sites of crossing over between nonsister chromatids.
Genetic Variability: Crossing over is a major source of genetic diversity in sexually reproducing organisms.
Linkage and Recombination Frequency
The likelihood of crossing over between two genes depends on the distance between them on the chromosome. The closer two genes are, the less likely a crossover will occur between them; the farther apart, the more likely recombination will occur.
Parental Types: Progeny with the same phenotype as one of the parents.
Recombinants: Progeny with new combinations of traits not found in either parent.
Genetic Mapping and Map Units
Geneticists use recombination frequencies to estimate the relative distances between genes. One map unit (mu), also called a centimorgan (cM), is defined as a 1% recombination frequency between two genes.
Additivity: Recombination frequencies between linked genes are additive, allowing for the construction of genetic maps.
Relative Distance: Map units represent relative, not absolute, physical distances.
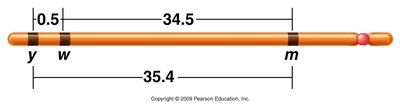
Example: If the recombination frequency between yellow and white is 0.5%, between white and miniature is 34.5%, and between yellow and miniature is 35.4%, the gene order is yellow – white – miniature.
Single Crossover Events
A single crossover (SCO) event between two genes can result in recombinant gametes if the crossover occurs between the genes. If the crossover occurs outside the region between the genes, only parental gametes are produced.
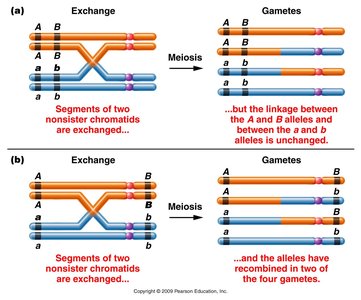
Effect of SCO: Only chromatids involved in the crossover are recombinant; the others remain unchanged.
Maximum Recombination: If a crossover occurs 100% of the time between two genes, the maximum observed recombination frequency is 50%.
Test Crosses and Phenotypic Ratios
Test crosses are used to determine the linkage relationship between genes. The phenotypic ratios observed in the progeny can indicate whether genes are unlinked, completely linked, or partially linked due to crossing over.
Unlinked Genes: Produce a 1:1:1:1 ratio in a test cross.
Completely Linked Genes: Only parental types are observed.
Partially Linked Genes: Both parental and recombinant types are observed, with the proportion of recombinants reflecting the distance between the genes.
Why Mendel Did Not Observe Linkage
Gregor Mendel did not detect linkage in his pea plant experiments because the seven genes he studied were either on different chromosomes or so far apart on the same chromosome that crossing over always occurred, resulting in independent assortment.
Summary Table: Linkage and Recombination Outcomes
Condition | Expected Phenotypic Ratio | Explanation |
|---|---|---|
Genes on separate chromosomes | 1:1:1:1 (test cross) | Independent assortment; all combinations equally likely |
Genes completely linked | 1:1 (parental types only) | No crossing over; only parental gametes formed |
Genes partially linked | Parental > Recombinant | Some crossing over; proportion of recombinants reflects distance |
Key Equations
Recombination Frequency (RF):
Map Distance (in map units or centimorgans):
Additional info: Chromosome mapping is a foundational tool in genetics, enabling the study of gene interactions, inheritance patterns, and the identification of genes associated with traits or diseases.
