 Back
BackCH5.2: Chromosome Mapping in Eukaryotes: Principles and Applications
Study Guide - Smart Notes

Chromosome Mapping in Eukaryotes
Linkage and Recombination
Chromosome mapping is a fundamental technique in genetics used to determine the relative positions of genes on a chromosome. The process relies on the principles of linkage and recombination, which describe how genes located close together on the same chromosome tend to be inherited together, while recombination events can separate them.
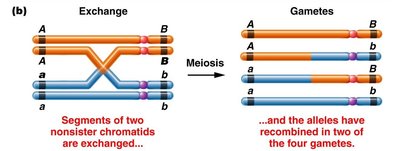
Linkage: Genes located on the same chromosome are said to be linked and do not assort independently.
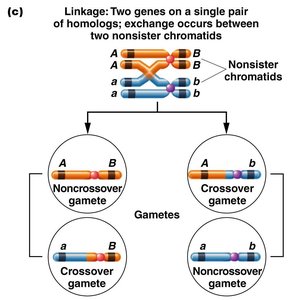
Recombination: The exchange of genetic material between homologous chromosomes during meiosis can produce new combinations of alleles, known as recombinant gametes.
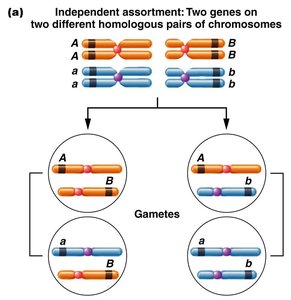
Independent Assortment: Genes on different chromosomes assort independently, producing four types of gametes in equal proportions.
Single and Double Crossovers
Crossovers are essential for mapping gene distances. A single crossover (SCO) alters linkage between two genes only if the crossover occurs between those genes. Double crossovers (DCOs) are rarer and provide information about the order of three genes.
Single Crossover (SCO): Used to determine the distance between two linked genes.
Double Crossover (DCO): Used to determine the order of three genes on a chromosome.
Map Unit (mu): One map unit is defined as 1% recombination between two genes, also called a centimorgan (cM).
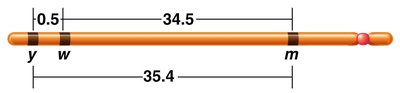
Three-Point Mapping
Three-point mapping is a technique used to determine the order and distances between three genes. It requires a parent heterozygous for all three genes, accurate determination of gamete genotypes, and a large sample size.
Criteria for Mapping: Parent must be heterozygous for all three genes, cross must allow genotype determination, and a large sample size is needed.
Gene Order: The order of genes is determined by analyzing the frequency of crossover classes.
Reciprocal Classes: F2 phenotypes complement each other and are called reciprocal classes.
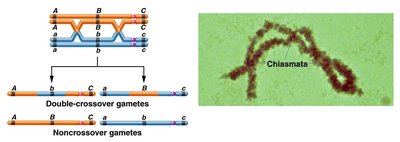
Calculating Recombination Frequencies
The expected frequency of double-crossover gametes is much lower than that of single-crossover gametes. The probability of two independent crossover events is the product of their individual probabilities (product law).
Product Law:
Example: If crossover between A and B occurs 20% of the time, and between B and C 30% of the time, the expected DCO frequency is (6%).
Gene Sequence Determination
Determining the sequence of genes requires analysis of multiple crossovers. The gene that switches position in the DCO phenotype compared to the NCO arrangement is the one in the middle.
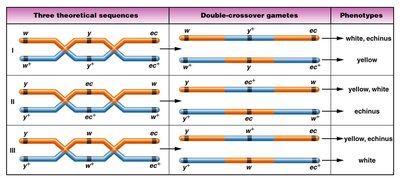
Mapping Example: Corn Genes
Consider a cross involving three linked genes (bm, v, pr) in corn. The arrangement of alleles, gene sequence, and map distances are determined by analyzing crossover classes and their frequencies.
Noncrossover Classes: Occur with the highest frequency.
Gene Sequence: Identified by the gene that shifts in DCOs.
Map Distance: Calculated by adding SCO and DCO frequencies between gene pairs.
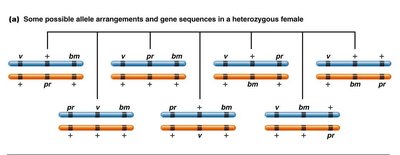
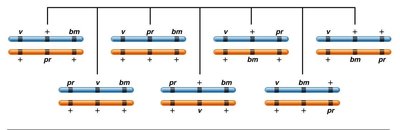
Interference and Mapping Accuracy
Interference affects the recovery of multiple exchanges. Positive interference occurs when fewer DCOs than expected are observed; negative interference occurs when more are observed. Mapping accuracy decreases as the distance between genes increases due to undetected multiple exchanges.
Interference Calculation:
Example: If expected DCO is 9.7% and observed is 7.8%, interference is positive.
Mapping Drosophila Genes
Drosophila genes have been extensively mapped, providing a reference for genetic studies. Chromosome maps show the relative positions of many genes.

Human Chromosome Mapping and Synteny Testing
Somatic cell hybridization and synteny testing are used to assign genes to specific human chromosomes. DNA markers and annotated databases now allow for precise mapping using recombination frequencies and physical distances.
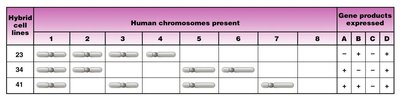
Somatic Cell Hybridization: Fusion of cells from different species, followed by chromosome loss, allows mapping of gene products to specific chromosomes.
Synteny Testing: Correlates gene product presence with chromosome presence in hybrid cell lines.
DNA Markers: Serve as landmarks for genetic and physical maps.

Summary Table: Crossover Classes and Mapping
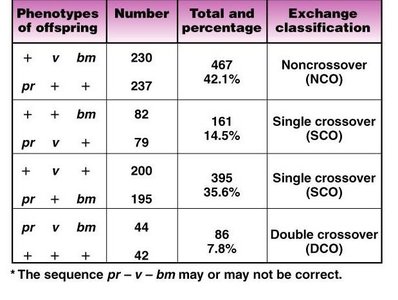
The following table summarizes the classification of crossover events, their frequencies, and the resulting phenotypes in a three-point mapping experiment.
Phenotypes of Offspring | Number | Total and Percentage | Exchange Classification |
|---|---|---|---|
+ v bm | 230 | 467 (42.1%) | Noncrossover (NCO) |
pr + + | 237 | ||
+ v + | 82 | 161 (14.5%) | Single crossover (SCO) |
pr + bm | 79 | ||
+ + bm | 200 | 395 (35.6%) | Single crossover (SCO) |
pr v + | 195 | ||
pr v bm | 44 | 86 (7.8%) | Double crossover (DCO) |
+ + + | 42 |
Key Terms and Concepts
Map Unit (mu): 1% recombination frequency.
Centimorgan (cM): Unit of genetic distance.
Interference: Reduction in observed DCOs compared to expected.
Synteny: Presence of genes on the same chromosome.
Physical Map: Actual base-pair distance between genes.
Practice Questions
Two genes that are 60 map units apart are expected to show what percent recombination? Answer: 50% (maximum recombination frequency).
The phenotypic classes of offspring representing double crossovers occur less frequently than single crossover classes.
In a three-point mapping experiment where no double crossovers are observed, complete positive interference has occurred.
Synteny testing allows researchers to assign genes to specific chromosomes.
Additional info: The notes expand on the original content by providing definitions, examples, and context for key concepts in chromosome mapping, including the use of tables and images to clarify crossover events and mapping strategies.
