 Back
BackChromosome Mapping in Eukaryotes: Principles and Applications
Study Guide - Smart Notes

Chromosome Mapping in Eukaryotes
Single and Double Crossovers: Linkage and Recombination
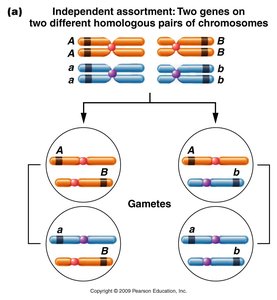
Chromosome mapping is a fundamental technique in genetics for determining the relative positions of genes on a chromosome. The process relies on the occurrence of crossovers during meiosis, which can separate linked genes and produce recombinant gametes.
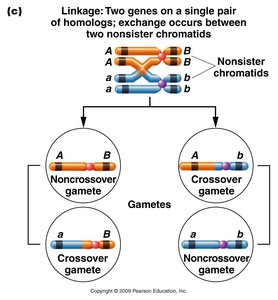
Single crossover (SCO): Alters linkage between two genes only if the crossover occurs between them, resulting in recombinant gametes.
Map unit (mu): Defined as 1 percent recombination between two genes; also called centimorgans (cM). These are relative, not absolute, distances.
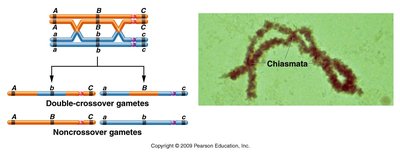
Double crossover (DCO): Involves two separate exchanges and is used to determine the order of three genes on a chromosome.
Product law: The probability of two independent crossover events occurring simultaneously is the product of their individual probabilities.
Example: If crossover between A and B occurs 20% of the time, and between B and C 30% of the time, the expected DCO frequency is .
Three-Point Mapping and Gene Order Determination
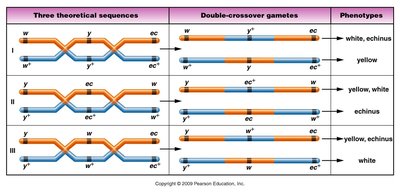
Three-point mapping is used to determine the sequence and distances between three linked genes. This method requires careful analysis of crossover classes and large sample sizes.
Criteria for mapping cross:
Parent must be heterozygous for all three genes.
Cross must allow accurate determination of gamete genotypes from offspring phenotypes.
Large sample size is needed to recover all crossover classes.
Gene order determination: The gene that switches position in the DCO phenotype compared to the noncrossover (NCO) arrangement is the one in the middle.
Map distance calculation: The distance between two genes in a three-point cross is the percentage of all detectable exchanges (SCO and DCO) between them.
Example: If SCO between v and pr is 14.5% and DCO is 7.8%, then the distance is mu.
Interference and Mapping Accuracy
Interference describes the phenomenon where one crossover event reduces the likelihood of another nearby crossover. This affects the recovery of multiple exchanges and the accuracy of mapping, especially as gene distances increase.
Positive interference: Fewer DCOs than expected occur.
Negative interference: More DCOs than expected occur.
Calculation:
Expected DCO:
Observed DCO: 7.8%
Interference is positive in this case.
Mapping accuracy: High when genes are close together; decreases as distance increases due to undetected multiple exchanges.
Generation of Human Chromosome Maps: Synteny Testing and DNA Markers
Human chromosome mapping has advanced through somatic cell hybridization and the use of DNA markers. Synteny testing allows researchers to assign genes to specific chromosomes by correlating gene expression with chromosome presence in hybrid cell lines.
Somatic cell hybridization: Fusion of two cells in culture, resulting in gradual loss of chromosomes from one species.
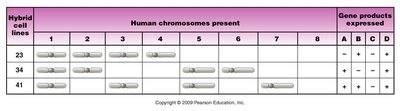
Synteny testing: Panel of cell lines, each with a few human chromosomes, is used to correlate gene products with chromosome presence.
DNA markers: Serve as landmarks for genetic mapping; recombination frequency forms genetic maps, while base-pair distance forms physical maps.
Hybrid cell lines | Human chromosomes present | Gene products expressed |
|---|---|---|
23 | 1, 2, 3, 4 | A, B |
34 | 2, 4, 5, 6 | B, C |
41 | 3, 5, 7, 8 | C, D |

Key Concepts and Review Questions
Map units (mu/cM): Relative measure of genetic distance based on recombination frequency.
Double crossovers: Occur less frequently than single crossovers.
Synteny testing: Assigns genes to specific chromosomes.
Interference: Complete positive interference means no DCOs are observed.
Recombination frequency: Two genes 60 mu apart are expected to show 50% recombination (maximum for linked genes).
Additional info:
Gene mapping is essential for understanding genetic linkage, inheritance patterns, and for applications in genomics and biotechnology.
Physical maps (base-pair distances) complement genetic maps (recombination frequencies) for comprehensive chromosome analysis.
