 Back
BackChromosome Mapping in Eukaryotes: Three-Point Crosses, Gene Order, and Mapping Accuracy
Study Guide - Smart Notes

Chromosome Mapping in Eukaryotes
Introduction to Chromosome Mapping
Chromosome mapping is a fundamental technique in genetics used to determine the relative positions of genes on a chromosome. By analyzing recombination frequencies between genes, geneticists can construct linkage maps that reveal gene order and distances. This process is essential for understanding inheritance patterns and for identifying genes associated with specific traits or diseases.
Single and Double Crossovers
Crossovers during meiosis are the basis for genetic recombination. A single crossover (SCO) event between two genes can separate linked alleles, resulting in recombinant gametes. The frequency of recombination is used to estimate the distance between genes, measured in map units (mu) or centimorgans (cM), where 1 mu = 1% recombination.
Single crossovers can only alter linkage between two genes if the crossover occurs between them.
Double crossovers (DCOs) involve two separate exchanges and are less frequent than SCOs. DCOs are crucial for determining the order of three genes on a chromosome.
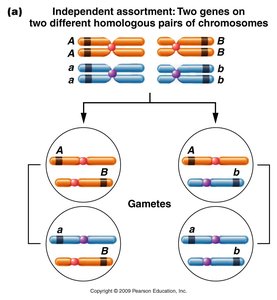
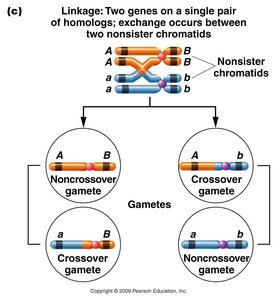
Three-Point Test Crosses
Three-point crosses are used to map the order and distances between three linked genes. This method requires:
A parent heterozygous for all three genes.
The ability to determine the genotype of gametes by observing offspring phenotypes.
A large sample size to recover all crossover classes.
In a three-point cross, the noncrossover (NCO) phenotypes are the most abundant, while double-crossover (DCO) phenotypes are the least abundant. Reciprocal classes of phenotypes (e.g., wild type vs. mutant for all three genes) help identify gene order.
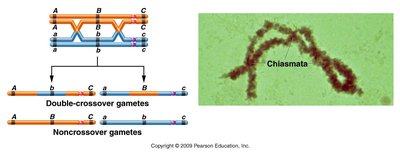
Determining Gene Order
To determine the order of three genes, compare the DCO phenotypes to the NCO phenotypes. The gene that differs between these classes is in the middle. For example, if the DCO class has only one gene switched compared to the NCO class, that gene is the central gene.
Possible gene orders must be considered and tested against observed crossover classes.
The correct order is the one where the DCO class matches the expected switch in the middle gene.
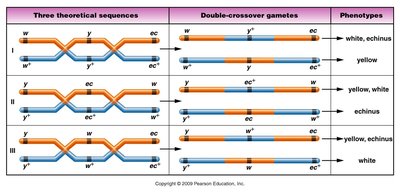
Calculating Map Distances
The distance between two genes in a three-point cross is the percentage of all detectable exchanges (SCOs and DCOs) between them. For each gene pair, add the frequencies of relevant SCO and DCO classes to obtain the map distance:
For genes v and pr:
For genes pr and bm:
Interference and Coincidence
Interference describes how one crossover event can influence the likelihood of another nearby crossover. It is calculated by comparing the observed and expected frequencies of DCOs:
Expected DCO frequency = (distance between gene 1 and 2) × (distance between gene 2 and 3)
Interference (I) =
Positive interference: fewer DCOs than expected
Negative interference: more DCOs than expected
Example: If expected DCO = 9.7% and observed DCO = 7.8%, interference is positive.
Accuracy of Mapping and Limitations
Mapping accuracy decreases as the distance between genes increases, due to undetected multiple crossovers. When genes are far apart, the maximum observable recombination frequency is 50%, which is indistinguishable from independent assortment.
Applications: Human Chromosome Mapping
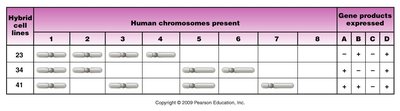
Somatic cell hybridization and synteny testing are used to assign genes to specific human chromosomes. By fusing cells from different species and analyzing which human chromosomes and gene products are retained, researchers can correlate gene expression with chromosome presence.

Modern Mapping: DNA Markers and Physical Maps
Today, chromosome mapping also uses DNA markers and annotated databases. Genetic maps are based on recombination frequencies, while physical maps use actual base-pair distances. DNA markers serve as landmarks for locating genes and constructing high-resolution maps.
Summary Table: Key Concepts in Chromosome Mapping
Concept | Definition | Example/Application |
|---|---|---|
Map Unit (mu) | 1% recombination frequency | 10 mu = 10% recombination |
Single Crossover (SCO) | Exchange between two homologous chromosomes | Used to map two genes |
Double Crossover (DCO) | Two exchanges between homologous chromosomes | Used to determine gene order |
Interference | Effect of one crossover on another nearby | Positive or negative interference |
Synteny Testing | Assigning genes to chromosomes using hybrid cells | Human gene mapping |
Additional info: The notes above expand on the original content by providing definitions, formulas, and examples for each key concept, ensuring the material is self-contained and suitable for exam preparation.
