 Back
BackChromosome Structure, Chromatin Organization, and Chromosomal Mutations
Study Guide - Smart Notes

Chromatin and Chromosome Structure
Chromatin Organization in Eukaryotes
Chromatin is the complex of DNA and proteins found in the nucleus of eukaryotic cells. It plays a critical role in packaging the long DNA molecules into a compact, organized structure, allowing for efficient regulation of gene expression and DNA replication.
Chromatin is dispersed throughout the nucleus during interphase and condenses into visible chromosomes during cell division.
It consists of DNA and various proteins, primarily histones, which facilitate DNA packaging.
Chromosomes occupy distinct regions called chromosome territories within the nucleus.
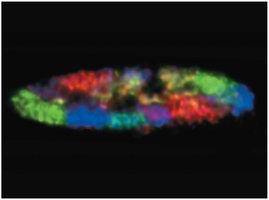
Histones and Nucleosomes
Histones are positively charged proteins that associate with DNA to form nucleosomes, the fundamental units of chromatin structure.
There are five main types of histones: H1, H2A, H2B, H3, and H4.
A nucleosome consists of DNA wrapped around a core of eight histone proteins (two each of H2A, H2B, H3, and H4).
Histone H1 binds to the linker DNA between nucleosomes, helping to further compact the chromatin.
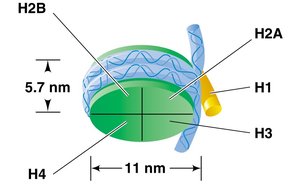
Nucleosome Arrangement and Higher-Order Structure
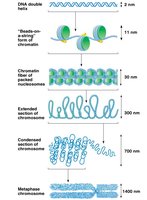
Electron microscopy reveals that chromatin fibers are composed of a linear array of nucleosomes, resembling "beads on a string." These structures are further condensed to form higher-order chromatin fibers and ultimately chromosomes.
The "beads-on-a-string" form is about 11 nm in diameter.
Nucleosomes are packed into a 30-nm chromatin fiber, which is then organized into looped domains anchored to a protein scaffold.
Further compaction leads to the highly condensed metaphase chromosome seen during cell division.

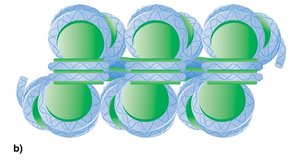
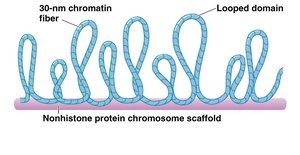
Chromatin Remodeling and Histone Modifications
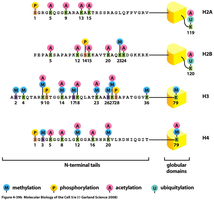
Chromatin structure is dynamic and can be remodeled to allow access to DNA for replication and gene expression. This remodeling is regulated by chemical modifications of histone proteins.
Acetylation: Addition of acetyl groups to lysine residues by histone acetyltransferase (HAT) neutralizes positive charges, loosening DNA-histone interactions and promoting gene expression.
Methylation: Addition of methyl groups to arginine or lysine residues by methyltransferases can either activate or repress gene expression, depending on the context.
Phosphorylation: Addition of phosphate groups to serine or histidine residues by kinases can alter chromatin structure and function.
Euchromatin and Heterochromatin
Chromatin exists in two main forms, which differ in their degree of compaction and transcriptional activity:
Euchromatin: Less condensed, transcriptionally active, and appears unstained during interphase.
Heterochromatin: Highly condensed, transcriptionally inactive, and appears stained during interphase. Includes regions such as telomeres and centromeres.
Chromosomal Mutations and Variations
Aneuploidy and Euploidy
Chromosomal mutations can involve changes in chromosome number or structure. These variations can have significant effects on phenotype and viability.
Aneuploidy: The number of chromosomes is not an exact multiple of the haploid set (e.g., monosomy, trisomy).
Euploidy: Complete sets of chromosomes are present (e.g., diploid, triploid, tetraploid).
Polyploidy: More than two sets of chromosomes (e.g., triploid 3n, tetraploid 4n).
Term | Explanation |
|---|---|
Aneuploidy | 2n ± x chromosomes |
Monosomy | 2n – 1 |
Trisomy | 2n + 1 |
Euploidy | Multiples of n |
Polyploidy | 3n, 4n, 5n, ... |

Monosomy and Trisomy
Monosomy and trisomy are specific types of aneuploidy that involve the loss or gain of a single chromosome, respectively.
Monosomy (2n – 1): Loss of one chromosome; often lethal due to unmasking of recessive lethal alleles and haploinsufficiency.
Trisomy (2n + 1): Gain of one chromosome; can be viable but often results in altered phenotype or developmental disorders.
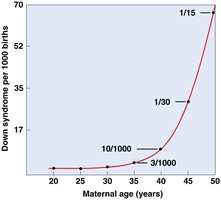
Examples include Down syndrome (trisomy 21), Patau syndrome (trisomy 13), and Edwards syndrome (trisomy 18).

Origin of Aneuploidy: Nondisjunction
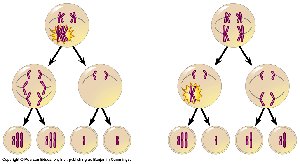
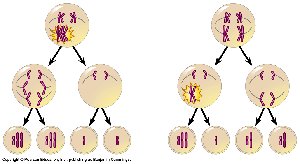
Aneuploidy often arises from nondisjunction, the failure of homologous chromosomes or sister chromatids to separate properly during meiosis.
Nondisjunction in meiosis I or II can result in gametes with abnormal chromosome numbers (n + 1 or n – 1).
Most cases of Down syndrome are due to nondisjunction during maternal meiosis.
Polyploidy: Autopolyploidy and Allopolyploidy
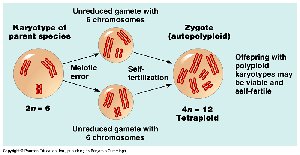
Polyploidy refers to the presence of more than two complete sets of chromosomes. It is common in plants and can arise through different mechanisms.
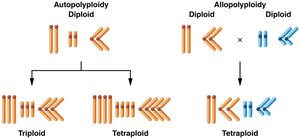
Autopolyploidy: Multiple chromosome sets from the same species, often due to meiotic errors.
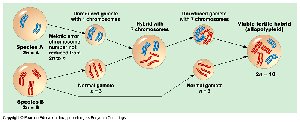
Allopolyploidy: Chromosome sets from different species, usually following hybridization and chromosome doubling.
Structural Chromosomal Mutations
Structural changes in chromosomes include deletions, duplications, inversions, and translocations. These mutations can disrupt gene function and regulation.
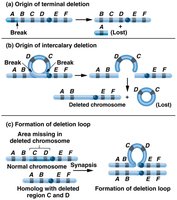
Deletions: Loss of a chromosome segment; can cause genetic diseases (e.g., Cri-du-chat syndrome).
Duplications: Repeated segments; can lead to gene redundancy or novel functions.
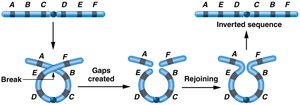
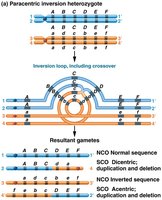
Inversions: Segment is reversed within the chromosome; can affect gene order and recombination.
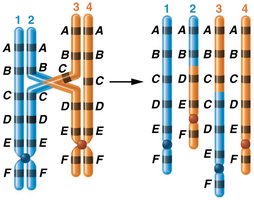
Translocations: Segment from one chromosome is transferred to another; can be reciprocal or Robertsonian.
Gene Redundancy, Amplification, and Copy Number Variants (CNVs)
Gene duplication can result in multiple copies of genes, which may provide evolutionary advantages or contribute to genetic disorders.
Gene redundancy: Multiple copies of genes (e.g., rRNA genes) ensure sufficient gene product.
Gene amplification: Selective replication of certain gene regions increases gene dosage (e.g., rRNA in oocytes).
Copy number variants (CNVs): Large-scale variations in the number of copies of DNA segments; can affect gene expression and phenotype.
Clinical Examples of Chromosomal Mutations
Down syndrome (Trisomy 21): Caused by an extra copy of chromosome 21; associated with intellectual disability and characteristic physical features.
Klinefelter syndrome (XXY): Males with an extra X chromosome; symptoms include tall stature, reduced fertility, and some feminized traits.
Turner syndrome (XO): Females with only one X chromosome; symptoms include short stature, infertility, and specific physical features.
Cri-du-chat syndrome: Deletion on the short arm of chromosome 5; causes intellectual disability and distinctive cry in infants.
Fragile X syndrome: Expansion of CGG repeats in the FMR-1 gene on the X chromosome; most common inherited cause of intellectual disability.
Conclusion: The Importance of Chromosome Structure and Number
The integrity of chromosome structure and number is essential for normal development and function. Mutations affecting chromosomes can have profound effects on phenotype, viability, and evolution.
