 Back
BackCh 10: Chromosome Structure, Organization, and Transposable Elements
Study Guide - Smart Notes

Chromosome Structure and Organization
Introduction to Chromosomes
Chromosomes are the fundamental structures that contain the genetic material of an organism. They are composed of DNA and associated proteins, and their organization is crucial for gene expression, replication, and segregation during cell division. The genome refers to all the genetic material in an organism, which in bacteria is typically a single circular chromosome, while in eukaryotes it includes multiple linear chromosomes as well as organellar genomes (mitochondria and chloroplasts).
Genome: The complete set of genetic material in an organism.
Chromosome: A DNA-protein complex that carries genetic information.
Gene: A segment of DNA that encodes a functional product, usually a protein.
Organization of Bacterial Chromosomes
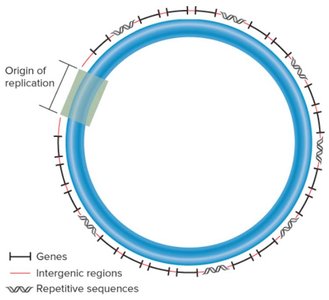
Bacterial chromosomes are typically circular DNA molecules, a few million base pairs in length, containing thousands of genes. Most bacteria have a single chromosome, but it may be present in multiple copies. The DNA is organized into loop domains and is compacted by supercoiling and DNA-binding proteins.
Intergenic regions: Noncoding DNA between genes.
Origin of replication: Specific site where DNA replication begins.
Repetitive sequences: Short DNA sequences repeated throughout the chromosome.
Compaction of Bacterial Chromosomes
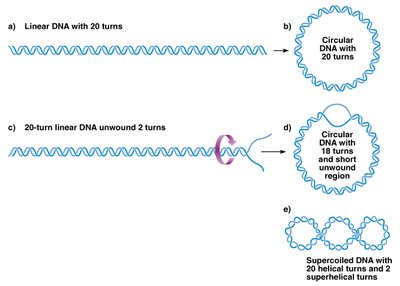
To fit within the cell, bacterial DNA is compacted by forming loop domains and by supercoiling. Nucleoid-associated proteins (NAPs) help organize and compact the DNA, while supercoiling further reduces its size. Supercoiling can be negative (underwinding) or positive (overwinding), and is regulated by enzymes called topoisomerases.
Loop domains: Structural loops of DNA anchored by proteins.
Supercoiling: Additional twisting of the DNA helix, which can be negative or positive.
Topoisomerases: Enzymes that control DNA supercoiling.
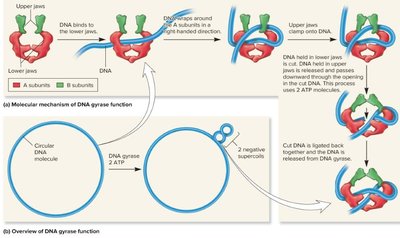
Control of Supercoiling in Bacteria
Supercoiling is regulated by two main enzymes:
DNA gyrase (topoisomerase II): Introduces negative supercoils using ATP and can relax positive supercoils.
DNA topoisomerase I: Relaxes negative supercoils.
The balance between these enzymes determines the overall supercoiling state of bacterial DNA.
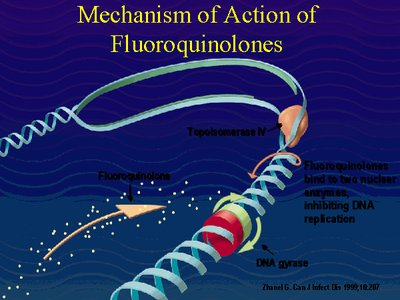
DNA Gyrase Inhibitors
DNA gyrase is essential for bacterial survival. Inhibitors of this enzyme, such as quinolones (e.g., ciprofloxacin) and coumarins, are used as antibiotics because they block DNA replication in bacteria without affecting eukaryotic topoisomerases.
Organization of Eukaryotic Chromosomes
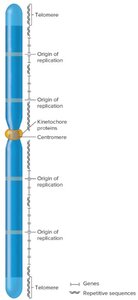
Eukaryotic chromosomes are linear and found in sets. Humans have two sets of 23 chromosomes. Eukaryotic chromosomes are much larger than bacterial chromosomes and contain more genes, as well as large amounts of noncoding and repetitive DNA. Three key DNA sequences are required for chromosome function:
Origins of replication: Multiple sites where DNA replication begins.
Centromeres: Regions important for chromosome segregation during cell division.
Telomeres: Specialized ends of chromosomes important for stability and replication.
Repetitive DNA Sequences in Eukaryotes
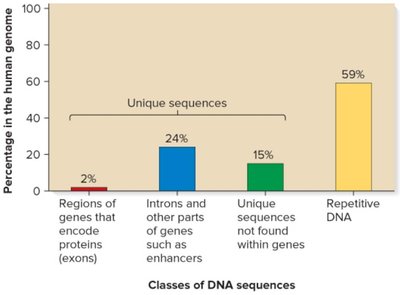
Eukaryotic genomes contain unique, moderately repetitive, and highly repetitive sequences. The amount of repetitive DNA can vary greatly between species and is not always correlated with organismal complexity.
Unique sequences: Found once or a few times in the genome (e.g., most genes).
Moderately repetitive: Found hundreds to thousands of times (e.g., rRNA genes, histone genes, transposable elements).
Highly repetitive: Found tens of thousands to millions of times (e.g., Alu elements in humans).
Transposable Elements (TEs)
Introduction to Transposable Elements
Transposable elements (TEs), or "jumping genes," are DNA sequences that can move to new locations within the genome. First discovered by Barbara McClintock in corn, TEs are now known to be widespread in bacteria, fungi, plants, and animals. TEs can cause mutations and contribute to genome evolution.
Transposition: The process by which TEs move within the genome.
Mutable site: A chromosomal location where a TE has inserted, causing genetic instability.
Types of Transposable Elements
There are two main types of transposition pathways:
Simple transposition (cut-and-paste): The TE is excised from one location and inserted into another.
Retrotransposition: The TE is transcribed into RNA, reverse transcribed into DNA, and inserted at a new site. This is common in eukaryotes.
Structure of Transposable Elements
Insertion elements: The simplest TEs, flanked by inverted repeats and often containing a gene for transposase.
Simple transposons: Carry additional genes, such as antibiotic resistance genes.
LTR retrotransposons: Contain long terminal repeats and genes for reverse transcriptase and integrase.
Non-LTR retrotransposons: Lack LTRs; may encode reverse transcriptase and endonuclease.
Autonomous vs. Nonautonomous Elements
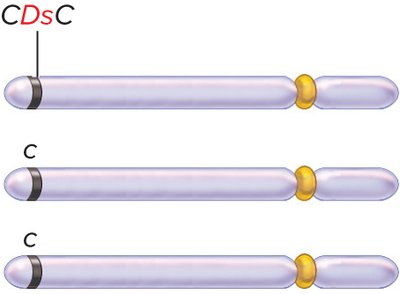
Autonomous elements: Contain all genes necessary for their own transposition (e.g., Ac element in corn).
Nonautonomous elements: Lack essential genes and require autonomous elements for movement (e.g., Ds element in corn).
Biological Significance of Transposons
Transposons can increase genetic variability, cause mutations, and contribute to genome evolution. They may also carry beneficial genes, such as antibiotic resistance in bacteria. The "selfish DNA theory" suggests that TEs persist because they can replicate within the genome, while another view is that they may provide evolutionary advantages.
Structure and Compaction of Eukaryotic Chromosomes
Levels of Chromatin Organization
Eukaryotic DNA is highly compacted to fit within the nucleus. The DNA-protein complex is called chromatin, which can be divided into euchromatin (less condensed, transcriptionally active) and heterochromatin (highly condensed, transcriptionally inactive).
Nucleosome: The basic unit of chromatin, consisting of DNA wrapped around a histone octamer.
Histones: Basic proteins (H2A, H2B, H3, H4, and H1) that package and order DNA into nucleosomes.
30 nm fiber: A higher-order structure formed by the association of nucleosomes.
Radial loop domains: Loops of chromatin attached to the nuclear matrix, further compacting the DNA.
Higher-Order Chromosome Structure
During cell division, chromosomes become highly condensed, forming metaphase chromosomes. This compaction involves the formation of radial loop domains and the action of protein complexes such as condensin and cohesin, which help organize and segregate chromosomes.
Condensin: Promotes chromosome condensation during mitosis and meiosis.
Cohesin: Holds sister chromatids together until anaphase.
SMC proteins: Structural maintenance of chromosomes proteins involved in chromosome architecture.
Summary Table: Abundance of Transposable Elements in Selected Species
Species | Percentage of Genome Composed of TEs |
|---|---|
Frog (Xenopus laevis) | 77% |
Corn (Zea mays) | 60% |
Human (Homo sapiens) | 45% |
Mouse (Mus musculus) | 40% |
Fruit fly (Drosophila melanogaster) | 20% |
Nematode (Caenorhabditis elegans) | 12% |
Yeast (Saccharomyces cerevisiae) | 4% |
Bacterium (Escherichia coli) | 0.3% |
Key Equations
Supercoiling linking number:
Negative supercoiling:
Additional info:
Some images and diagrams referenced in the notes are not included here but are available in the original source material. The notes above provide a comprehensive overview of chromosome structure, organization, and the role of transposable elements, suitable for college-level genetics students.
