 Back
BackDNA Repair Systems, Transposable Elements, and the Genetic Code: Study Notes for Genetics Students
Study Guide - Smart Notes

DNA Repair Systems
Overview of DNA Repair
DNA repair systems are essential for maintaining the integrity of genetic material in all organisms. They counteract both spontaneous and induced DNA damage, preventing genetic diseases and cancer. Defective DNA repair mechanisms are linked to increased cancer susceptibility.
Mismatch Repair (MMR): Corrects base pair mismatches not fixed by DNA polymerase proofreading. Involves endonuclease, exonuclease, DNA polymerase, and ligase. Defective MMR is associated with hereditary colon cancer.
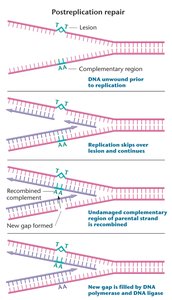
Post-Replication Repair: Repairs gaps left when DNA damage blocks replication. Uses recombination with the undamaged template to fill gaps, allowing replication to finish despite DNA damage.
SOS Repair: Unique to bacteria. Induces expression of multiple genes, including error-prone polymerases, to allow survival under extensive DNA damage. Results in random nucleotide insertions.
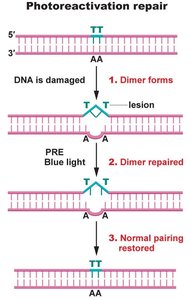
Photoactivation Repair: Cleaves thymidine dimers caused by UV radiation using photoreactivation enzyme (PRE), activated by blue light. Not found in humans.
Base Excision Repair: Corrects minor defects affecting single bases, such as uracil in DNA. DNA glycosylase removes the base, AP endonuclease cleaves the backbone, and DNA polymerase and ligase repair the gap.
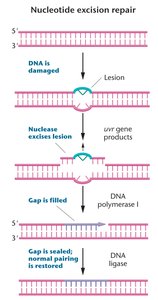
Nucleotide Excision Repair (NER): Repairs bulky lesions that distort the double helix. Removes a 13-28 bp region and fills the gap using DNA polymerase and ligase.
Xeroderma Pigmentosum: Human disease caused by defective NER, leading to skin cancer susceptibility. NER involves damage recognition proteins, helicases, nucleases, and DNA polymerases.
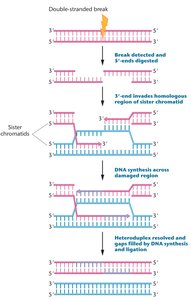
Double-Strand Break Repair: Repairs double-stranded DNA breaks via homologous recombination or nonhomologous end joining. Defects predispose to hereditary breast and ovarian cancers.
Transposable Elements (Transposons)
Discovery and Biological Impact
Transposable elements are DNA sequences that can move within the genome. Discovered in maize, they cause genetic variation and can revert mutant phenotypes to wild type.
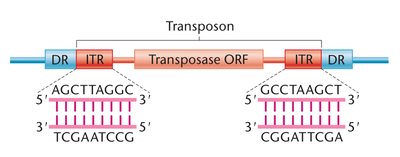
DNA Transposons: Move without an RNA intermediate. Contain inverted terminal repeats (ITRs) recognized by transposase enzyme. Use a cut-and-paste mechanism.


Types of DNA Transposons:
Autonomous elements: Can transpose independently; mutations are unstable.
Nonautonomous elements: Require autonomous elements for transposition; insertions are stable unless autonomous elements are present.
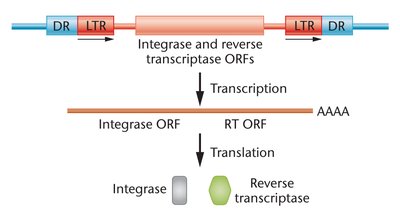
Retrotransposons: Move via an RNA intermediate using a copy-and-paste mechanism. Resemble retroviruses.
Human Transposable Elements: LINEs and SINEs constitute 35% of the human genome. LINEs are 1-6 kb with 850,000 copies; SINEs are 100-500 bp with 1,500,000 copies. Active transposition can cause disease.
The Genetic Code and Transcription
Flow of Genetic Information
Genetic information flows from DNA to mRNA to protein. mRNA acts as an unstable intermediate, transferring information from the nucleus to the cytosol. Protein synthesis uses the genetic code to convert nucleotide sequences into proteins.
Properties of the Genetic Code
Linear and Triplet Code: mRNA sequence is read in triplets (codons), each specifying an amino acid or stop signal.
Unambiguous and Degenerate: Each codon specifies only one amino acid, but some amino acids are specified by multiple codons.
Start and Stop Codons: One start codon (AUG) and three stop codons (UAA, UAG, UGA).
Continuous and Nonoverlapping: Codons are read sequentially without overlap.
Nearly Universal: The code is conserved across most organisms, with minor exceptions.
Experimental Deciphering of the Genetic Code
Frameshift Mutations: Showed the code is a triplet.
Synthetic mRNAs: Used to determine which codons specify which amino acids.
Triplet Binding Assay: Identified codon assignments using labeled tRNAs and ribosomes.
Repeating Copolymers: Used to assign codons to amino acids by synthesizing mRNAs with repeating sequences.
The Wobble Hypothesis
The first two bases of a codon are more critical for tRNA recognition than the third, allowing one tRNA to pair with multiple codons. This reduces the number of tRNA species needed.
Universality and Ordered Nature of the Genetic Code
Universality: The code is conserved in almost all organisms, with rare exceptions.
Ordered Nature: Chemically similar amino acids share the middle base in their codons, buffering the effects of mutations.
Transcription Mechanisms
Transcription in Prokaryotes and Eukaryotes
RNA Polymerase: Catalyzes RNA synthesis using NTPs. Does not require a primer. Synthesizes RNA 5' to 3'.
Promoters: DNA sequences upstream of the transcription start site. Contain consensus sequences (TATAAT at -10, TTGACA at -35 in bacteria).
Initiation, Elongation, Termination: Initiation involves promoter recognition, elongation involves RNA synthesis, and termination involves unique sequences signaling the end of transcription.
Eukaryotic Transcription: Occurs in the nucleus, involves three RNA polymerases, chromatin remodeling, and extensive cis- and trans-acting factors.
Post-Transcriptional mRNA Processing in Eukaryotes
5' Cap Addition: 7-methylguanosine cap protects mRNA and aids translation initiation.
Intron Splicing: Introns are removed, exons are spliced together.
PolyA Tail Addition: Stabilizes mRNA and is important for export and translation.
Intron Functions and Splicing Mechanisms
Alternative Splicing: Allows production of multiple proteins from one gene.
Spliceosomal Intron Removal: Involves snRNAs and snRNPs, forming a lariat structure.
RNA Editing: Enzymatic alteration of mRNA base sequence post-transcription.
Translation and Protein Synthesis
tRNA Structure and Function
tRNAs: Adapt codons in mRNA to the correct amino acid. Anticodons complement mRNA codons.
Structure: Cloverleaf stem-loop with modified bases. 5'-CCA-3' at the 3' end is the amino acid attachment site.
Charging: Aminoacyl-tRNA synthetases covalently link amino acids to tRNAs.
Translation Mechanisms
Initiation: mRNA associates with the small ribosomal subunit. Initiator tRNA and initiation factors are required.
Elongation: Charged tRNAs occupy A and P sites. Peptide bonds are formed by rRNA in the large subunit.
Termination: Release factors recognize stop codons and dissociate the complex.
Ribosome Structure
Composition: 50-80 proteins and 3-4 rRNAs. Subunits named by Svedberg coefficient.
Prokaryotic vs Eukaryotic Ribosomes: Prokaryotes have smaller ribosomes and coupled transcription/translation; eukaryotes have larger ribosomes and spatial separation.
Regulation of Gene Expression in Bacteria
Gene Regulation Overview
Gene regulation controls transcription, determining how much mRNA is made, when, and in which cells. It allows cells to conserve energy, differentiate, and respond to environmental changes.
Types of Regulation
Inducible Systems: Genes are transcribed only when an inducer is present.
Repressible Systems: Transcription stops when a specific substance is present.
Negative Control: Transcription occurs unless shut off by a repressor.
Positive Control: Transcription occurs only if an activator stimulates RNA production.
Operons
Definition: Genomic regions transcribing polycistronic mRNAs, containing promoter, operator, and structural genes.
lac Operon: Negative control by lac repressor; positive control by CAP. Regulates lactose metabolism.
trp Operon: Repressible system under negative control. Regulates tryptophan synthesis.
Additional Regulatory Mechanisms
Attenuation: Leader sequence regulates transcription.
Riboswitches: mRNA secondary structures regulate transcription in response to metabolites.
sRNAs: Small noncoding RNAs regulate translation of other genes.
CRISPR/Cas: Prokaryotic adaptive immunity system, destroys invading phage DNA.
