 Back
BackEukaryotic Gene Expression and Regulation: Structured Study Notes
Study Guide - Smart Notes

Gene Regulation in Eukaryotes
Overview of Eukaryotic Gene Expression
Eukaryotic gene expression is a highly regulated process that determines when, where, and how much of a gene product is produced. Regulation occurs at multiple levels, but transcriptional control is a primary mechanism.
Transcription factors are proteins that bind to specific DNA sequences to regulate gene expression.
Regulatory DNA sequences include promoters, enhancers, silencers, and insulators.
Gene expression is influenced by chromatin modifications, such as histone modification and DNA methylation.

Transcription Activators and Repressors
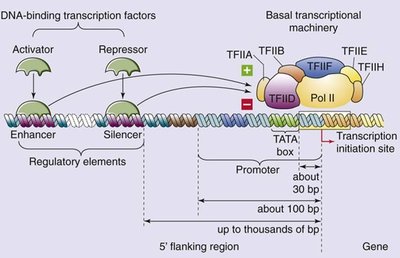
Transcription activators and repressors are key regulatory proteins that modulate gene expression by binding to specific DNA elements.
Activator: A protein that turns genes ON by binding to enhancer DNA or proximal-promoter elements, increasing transcription.
Enhancer: A regulatory DNA sequence that activators bind to, stimulating transcription from a distance.
Repressor: A protein that turns genes OFF by binding to silencer DNA, decreasing transcription.
Silencer: A regulatory DNA sequence that repressors bind to, inhibiting transcription initiation.

Regulation of Transcription Initiation
The initiation of transcription is regulated by the interaction of multiple transcription factors and regulatory DNA elements.
Enhancers and silencers can be located upstream, downstream, or within genes, modulating transcription from a distance.
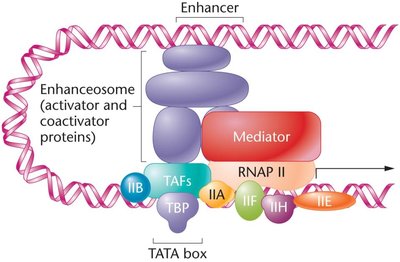
DNA looping brings activators, repressors, and general transcription factors to the promoter vicinity, often mediated by the Mediator complex.
The combined effects of multiple transcription regulators determine the final rate of transcription initiation.

Combinatorial Control of Gene Activation
Combinatorial control refers to the regulation of gene expression by the combined action of multiple transcription factors.
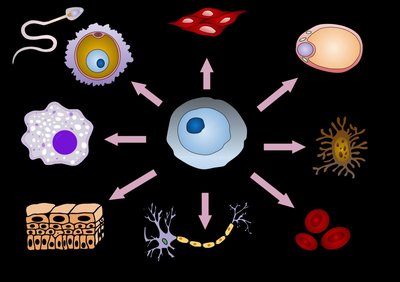
Different combinations of transcription factors (TFs) determine when, where, and how strongly a gene is expressed.
Identical genomes can produce different outcomes due to differential gene expression, allowing cells to specialize.
Control elements activate transcription only when the appropriate activator proteins are present.

Regulatory Control Elements: Promoters, Enhancers, Silencers, Insulators
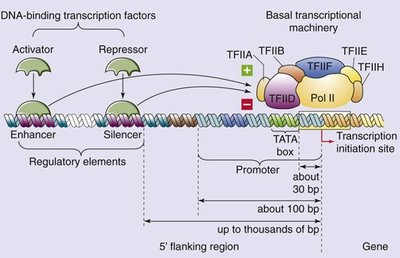
Regulatory control elements are DNA sequences that serve as binding sites for protein factors and are essential for accurate and regulated transcription.
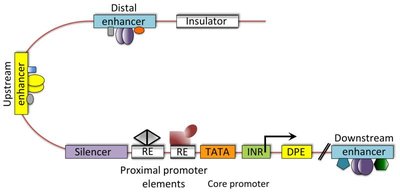
Promoters: Contain core and proximal-promoter elements; core promoter determines accurate initiation, proximal elements modulate efficiency.
Enhancers: Increase transcription levels, can act from a distance.
Silencers: Decrease transcription levels, can act from a distance.
Insulators: DNA sequences that block or insulate the effect of enhancers, preventing them from influencing non-target genes.
Mutation Analysis of Regulatory Elements
Mutations within regulatory elements can significantly affect transcription levels.
Mutations in core promoter elements (e.g., TATA box) can reduce transcription substantially.
Mutations in proximal-promoter elements (e.g., CAAT, GC boxes) can modulate basal transcription levels.
Reporter gene assays are used to study the effects of mutations or deletions in regulatory elements.
Reporter Gene Assays
Reporter gene assays are experimental tools used to monitor gene expression.
A reporter gene is attached to a regulatory sequence (promoter or enhancer) of another gene.
By deleting or mutating control elements, researchers can assess their role in gene expression.
If deletion causes a reduction in expression, the element likely activates transcription; if deletion causes an increase, the element likely represses transcription.
Cis-Acting Elements and Trans-Acting Factors
Cis-acting elements: DNA sequences located on the same molecule as the gene they regulate; act locally.
Trans-acting factors: Proteins encoded elsewhere in the genome; can diffuse and act on target cis-acting DNA sequences.
Experimental Analysis: Nuclear Extract vs Purified System
Nuclear extract contains most proteins from cell nuclei, including specific transcription factors.
Purified system contains only RNA Pol II and general transcription factors.
High transcription levels in nuclear extract but low in purified system indicate the requirement for additional specific factors.
Essential promoter elements (e.g., TATA box) are required for basal transcription.
Transcriptional Regulation in Yeast: The GAL Gene System
The GAL gene system in yeast is a classic model for studying eukaryotic gene regulation.
Four structural genes (GAL1, GAL2, GAL7, GAL10) are required for galactose metabolism.
Three regulatory genes (GAL3, GAL4, GAL80) control expression of structural genes.
GAL4 is a transcriptional activator; GAL80 is a repressor; GAL3 is a signal transducer/sensor.
In the absence of galactose, GAL80 binds GAL4 and blocks activation. In the presence of galactose, GAL3 inhibits GAL80, allowing GAL4 to activate transcription.
Mutation Analysis in the GAL System
Mutations in GAL4 (activator) or GAL3 (sensor) can prevent activation of GAL genes.
Mutations in GAL80 (repressor) can lead to constitutive expression of GAL genes.
Mutations in UASG (upstream activation sequence) or the TATA box can reduce or abolish transcription.
Summary Table: Regulatory Elements and Their Functions
Element | Function | Effect of Mutation |
|---|---|---|
Promoter (Core) | Initiates transcription | Reduces transcription if mutated |
Enhancer | Stimulates transcription | Reduces transcription if mutated |
Silencer | Represses transcription | Increases transcription if mutated |
Insulator | Blocks enhancer effects | Loss of boundary, misregulation |
UASG (Yeast) | Activates GAL genes | Reduces GAL gene expression if mutated |
Key Equations and Methods
Quantification of transcriptional output: RT-qPCR, RNA-seq, Reporter gene assays
Example equation for gene expression quantification:
Posttranscriptional Regulation in Eukaryotes
Alternative splicing determines which RNA spliceforms are translated.
Gene expression is regulated by mRNA stability and degradation.
Noncoding RNAs play diverse roles in posttranscriptional regulation.
mRNA localization and translation initiation are highly regulated.
Posttranslational modifications regulate protein activity.
Epigenetic Regulation
Histone modification and DNA methylation influence gene expression.
Epigenetic changes can be heritable and affect cell differentiation.
Learning Objectives
Understand the organization of eukaryotic cells and gene regulation at multiple levels.
Describe the influence of chromatin modifications on gene expression.
Explain the role of cis-acting elements and trans-acting factors in transcription initiation.
Analyze mutation effects on regulatory elements using reporter gene assays.
Apply knowledge of the GAL gene system in yeast to understand eukaryotic gene regulation.
