 Back
BackGene Expression and Regulation in Bacteria: Mechanisms and Operon Models
Study Guide - Smart Notes

Gene Expression and Regulation in Bacteria
Overview of Gene Regulation
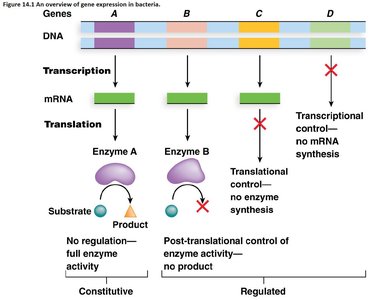
Gene expression in bacteria is tightly regulated to ensure that proteins are produced only when needed. This regulation occurs at multiple levels, including transcription, translation, and post-translation. The primary control point is transcription, but bacteria also utilize translational and post-translational mechanisms to fine-tune gene expression.
Constitutive expression: Genes that are always expressed, such as those encoding ribosomal proteins.
Regulated expression: Genes whose expression is controlled in response to environmental or cellular signals.
Regulatory proteins: Proteins that interact with DNA to increase or decrease transcription.
Mechanisms of regulation: Include negative and positive control, attenuation, translational repression, and feedback inhibition.
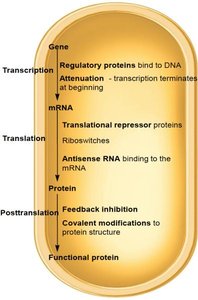
Mechanisms of Gene Expression Regulation
Negative and Positive Control of Transcription
Transcriptional regulation is achieved through the action of regulatory proteins that bind to specific DNA sequences near target genes. These proteins can act as repressors (negative control) or activators (positive control).
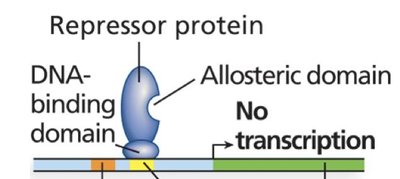
Negative control: A repressor protein binds to an operator sequence, blocking RNA polymerase and preventing transcription.
Positive control: An activator protein binds to a regulatory sequence, facilitating RNA polymerase binding and promoting transcription.
Negative Control: Repressors and Inducers
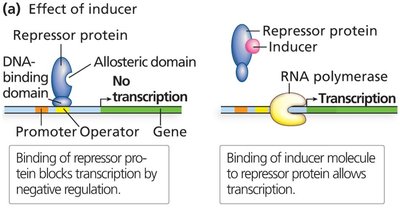
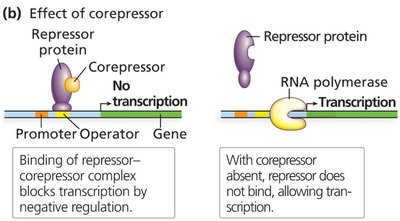
Repressor proteins have a DNA-binding domain and an allosteric domain. The allosteric domain binds regulatory molecules (inducers or corepressors) that alter the repressor's ability to bind DNA.
Inducers inactivate repressors, allowing transcription to proceed.
Corepressors activate repressors, blocking transcription.
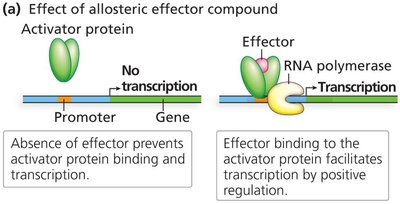
Positive Control: Activators and Effectors
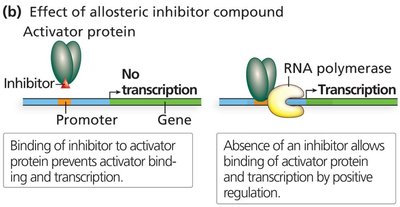
Activator proteins bind to activator binding sites and enhance RNA polymerase recruitment.
Allosteric effectors activate the DNA-binding ability of activators.
Inhibitors prevent activator binding, reducing transcription.
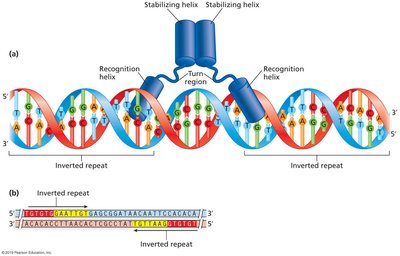
DNA-Protein Interactions: The Helix-Turn-Helix Motif
Many bacterial regulatory proteins use the helix-turn-helix (HTH) motif to recognize and bind specific DNA sequences, often at inverted repeats. The recognition helix fits into the major groove of DNA, while the stabilizing helix supports the interaction.
Operons: Coordinated Gene Regulation
Definition and Structure of Operons
An operon is a cluster of genes under the control of a single promoter and regulatory region, transcribed as a polycistronic mRNA. Operons allow bacteria to coordinate the expression of genes involved in the same metabolic pathway.
Regulatory region: Contains promoter, operator, and activator binding sites.
Structural genes: Encode proteins with related functions.
The lac Operon: An Inducible System
Function and Components
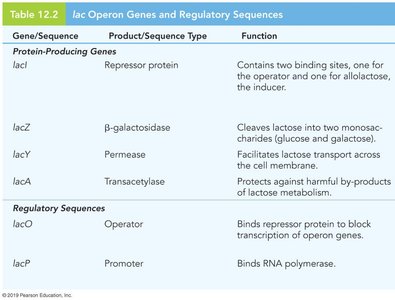
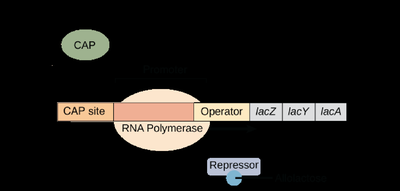
The lac operon in E. coli enables the bacterium to metabolize lactose when glucose is absent. It consists of three structural genes (lacZ, lacY, lacA), a promoter, an operator, and a CAP binding site. The lacI gene encodes the repressor protein but is not part of the operon itself.
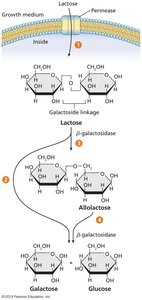
lacZ: Encodes β-galactosidase, which cleaves lactose into glucose and galactose.
lacY: Encodes permease, which transports lactose into the cell.
lacA: Encodes transacetylase, which detoxifies by-products of lactose metabolism.
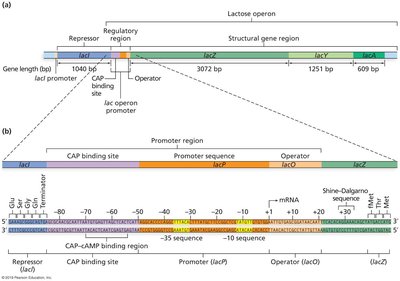
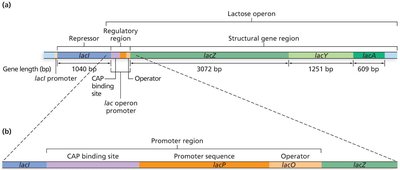
Regulation of the lac Operon
The lac operon is regulated by both negative and positive control mechanisms:
Negative control: The lac repressor (encoded by lacI) binds to the operator, blocking transcription in the absence of lactose.
Induction: When lactose is present, a small amount is converted to allolactose, which binds the repressor and inactivates it, allowing transcription.
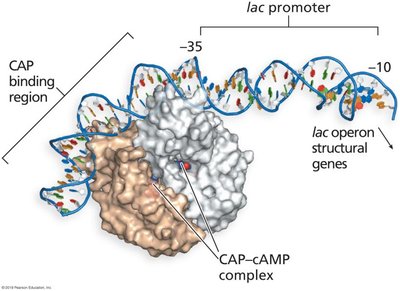
Positive control: When glucose is absent, cAMP levels rise, forming a CAP-cAMP complex that binds the promoter and enhances RNA polymerase binding.
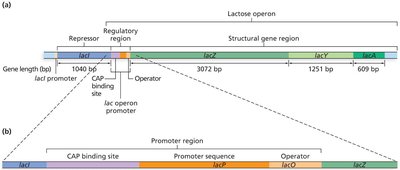
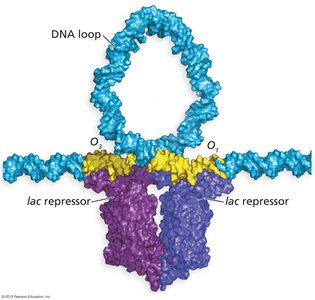
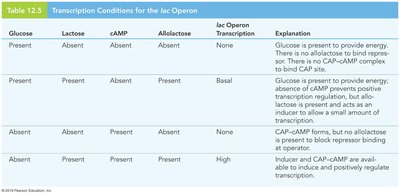
lac Operon Transcriptional States
Glucose | Lactose | cAMP | Allolactose | lac Operon Transcription | Explanation |
|---|---|---|---|---|---|
Present | Absent | Absent | Absent | None | CAP-cAMP not formed, repressor bound |
Present | Present | Absent | Present | Basal | Repressor removed, but no positive regulation |
Absent | Absent | Present | Absent | None | CAP-cAMP present, but repressor bound |
Absent | Present | Present | Present | High | Inducer and CAP-cAMP both present |


The trp Operon: A Repressible and Attenuated System
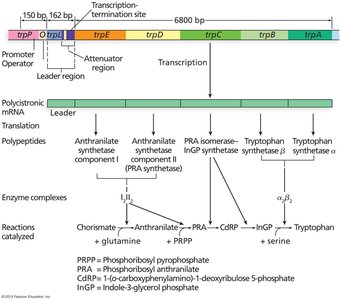
Function and Structure
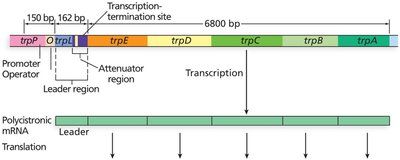
The trp operon in E. coli encodes enzymes for tryptophan biosynthesis. It is regulated by both repression (negative control) and attenuation (fine-tuning of transcription termination).
Structural genes: trpE, trpD, trpC, trpB, trpA
Regulatory regions: Promoter, operator, leader (trpL) with attenuator
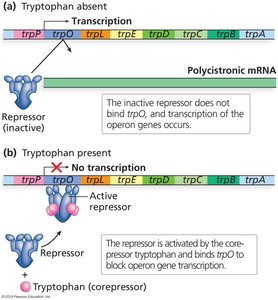
Negative Control: Repression by Tryptophan
When tryptophan is abundant, it acts as a corepressor, binding to the trp repressor protein and activating it.
The active repressor binds the operator, blocking transcription.
When tryptophan is scarce, the repressor is inactive and transcription proceeds.
Attenuation: Fine-Tuning trp Operon Expression
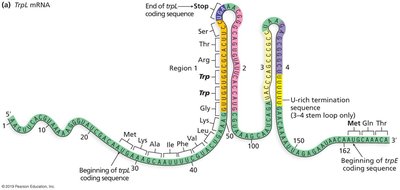
Attenuation is a regulatory mechanism that uses the coupling of transcription and translation in bacteria to fine-tune gene expression. The leader region (trpL) contains sequences that can form alternative stem-loop structures in the mRNA, influencing whether transcription continues or terminates prematurely.
Low tryptophan: Ribosome stalls at trp codons, allowing formation of the 2-3 antiterminator stem-loop, so transcription continues.
High tryptophan: Ribosome quickly translates leader peptide, allowing formation of the 3-4 terminator stem-loop, causing transcription termination.
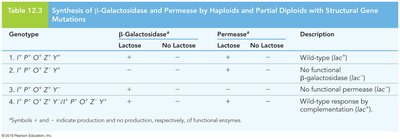
Mutations Affecting Operon Regulation
Structural vs. Regulatory Mutations
Structural gene mutations: Affect the function of the encoded enzymes, impacting the metabolic pathway directly.
Regulatory region mutations: Can cause constitutive expression (always on) or permanent repression (always off) by disrupting protein binding sites.
Operator mutations: May prevent repressor binding, leading to unregulated gene expression.
Allosteric site mutations: May prevent inducer or corepressor binding, locking the operon in an 'off' state.
Transcriptional Regulation in Archaea
Archaea share many regulatory features with bacteria, including operon organization and the use of repressor and activator proteins. However, some regulatory mechanisms are more similar to those found in eukaryotes.
Translational Regulation in Bacteria
Although less common than transcriptional regulation, bacteria can regulate gene expression at the translational level. Mechanisms include:
Translation repressor proteins: Bind near the Shine-Dalgarno sequence, blocking ribosome binding.
Small RNAs (sRNAs): Can activate or repress translation by base-pairing with mRNA.
Summary: Key Points
Gene expression in bacteria is regulated at multiple levels, with transcriptional control being the most prevalent.
Operons allow coordinated regulation of functionally related genes.
The lac operon is an inducible system controlled by both negative and positive regulation.
The trp operon is a repressible and attenuated system, allowing fine-tuned control of biosynthetic pathways.
Mutations in operon regions can have profound effects on gene expression and cellular metabolism.
