 Back
BackGene Expression and Regulation in Eukaryotes
Study Guide - Smart Notes

Gene Expression and Regulation in Eukaryotes
Introduction to Eukaryotic Gene Regulation
Gene expression in eukaryotes is regulated at multiple levels, allowing for precise control over cellular function, development, and response to environmental cues. This regulation is more complex than in prokaryotes due to the presence of multiple cell types, developmental stages, and the physical separation of transcription and translation.
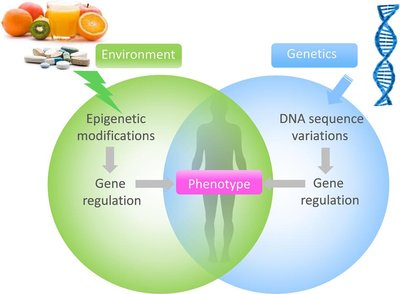
Phenotype results from the interaction between genotype (DNA sequence) and environmental factors, often mediated by epigenetic modifications.
Gene regulation occurs at transcriptional, post-transcriptional, translational, and post-translational levels.
Epigenetics and Environmental Influence
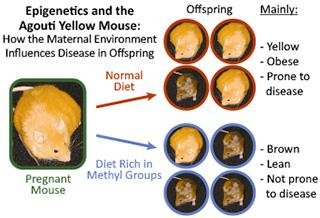
Epigenetic modifications are heritable changes in gene activity that do not alter the DNA sequence but affect how genes are read by cells. Environmental factors, such as diet, can induce epigenetic changes that influence phenotype.
Agouti mice provide a classic example: maternal diet rich in methyl groups leads to offspring with different phenotypes due to epigenetic silencing of the agouti gene.
Epigenetic changes can turn genes on or off, affecting traits such as obesity and disease susceptibility.
Levels of Gene Regulation in Eukaryotes
Overview of Regulatory Mechanisms
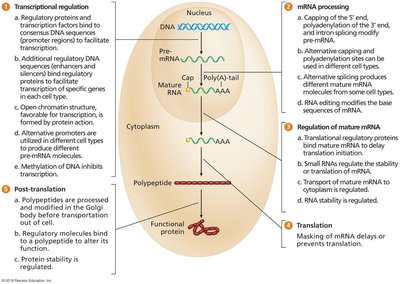
Gene expression in eukaryotes is regulated at several key stages:
Transcriptional regulation: Control of when and how much a gene is transcribed.
RNA processing: Modifications such as capping, splicing, and polyadenylation of pre-mRNA.
Regulation of mature mRNA: Stability and localization of mRNA molecules.
Translational regulation: Control of when and how much protein is synthesized from mRNA.
Post-translational regulation: Modifications to proteins after translation, affecting their activity and stability.
Complexity of Eukaryotic Gene Regulation
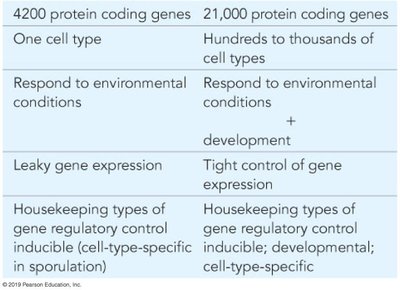
Eukaryotic gene regulation is more complex than in prokaryotes due to several factors:
Greater number of genes and cell types
Physical separation of transcription (nucleus) and translation (cytoplasm)
Developmental and environmental regulation
Tight control of gene expression
E. coli | Human | |
|---|---|---|
Protein coding genes | 4,200 | 21,000 |
Cell types | One | Hundreds to thousands |
Gene expression control | Leaky | Tight |
Regulatory control | Housekeeping, inducible | Housekeeping, developmental, cell-type-specific |
Transcriptional Regulation: Regulatory Sequences and Transcription Factors
Core Promoter Elements
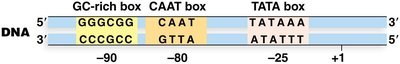
Core promoter regions are DNA sequences upstream of genes that are essential for the initiation of transcription. Key elements include:
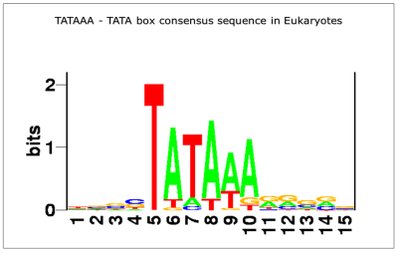
TATA box (Goldberg-Hogness box): 5'-TATAAA-3', located around -25 bp from the transcription start site.
CAAT box: Found near -80 bp.
GC-rich box: Consensus 5'-GGGCGG-3', located at -90 bp or further upstream.
Enhancers, Silencers, and Cis-Regulatory Elements
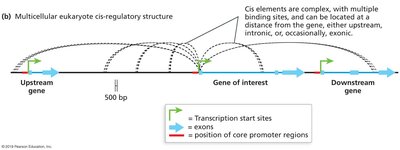
Enhancer and silencer regions are cis-acting regulatory sequences that can be located far from the genes they regulate. They can be upstream, downstream, or even within the gene.
Enhancers increase transcription; silencers decrease transcription.
These elements are often highly conserved and relatively short.
Transcription Factors and Regulatory Proteins
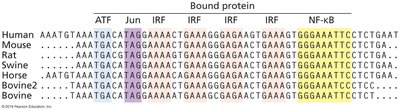
Transcription factors are trans-acting proteins that bind to specific DNA sequences to regulate gene expression. Examples include ATFs, Jun/AP-1, IRFs, and NF-κB.
Transcription factors can act on multiple chromosomes and genes.
They interact with cis-acting elements to control transcription.
Bound protein | ATF | Jun | IRF | NF-κB |
|---|---|---|---|---|
Human | AAATGTA | TGA | GGGAA | GGGAA |
Mouse | AAATGTA | TGA | GGGAA | GGGAA |
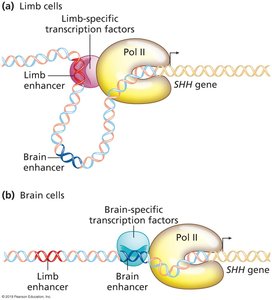
Cell-Type Specific Regulation: Example of SHH Gene
The sonic hedgehog (SHH) gene is regulated by cell-type specific transcription factors, leading to different expression patterns in limb and brain cells.
Limb-specific enhancers and transcription factors activate SHH in limb cells.
Brain-specific enhancers and transcription factors activate SHH in brain cells.
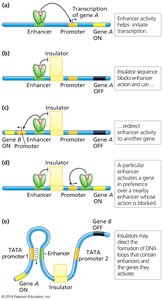
Insulator Sequences
Insulator sequences are cis-acting elements that protect genes from inappropriate regulatory signals by blocking or redirecting enhancer activity. They can also facilitate DNA loop formation, bringing distant regulatory elements into proximity with promoters.
Insulators bind regulatory proteins to insulate specific genes.
They can block or redirect enhancer-promoter interactions.
Epigenetic Regulation
Chromatin Structure and Remodeling
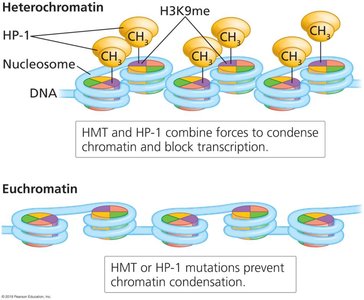
Epigenetic modifications alter chromatin structure, affecting gene accessibility and transcription. Chromatin can exist as:
Heterochromatin: Tightly packed, transcriptionally inactive.
Euchromatin: Loosely packed, transcriptionally active.
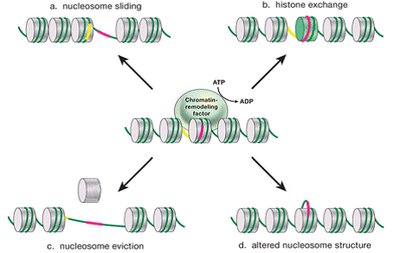
Chromatin Remodeling Complexes
Chromatin remodelers are protein complexes that use ATP to alter nucleosome structure, allowing access to DNA for transcription factors and polymerases. Mechanisms include nucleosome sliding, histone exchange, and nucleosome eviction.
Examples: SWI/SNF and NuRD complexes.
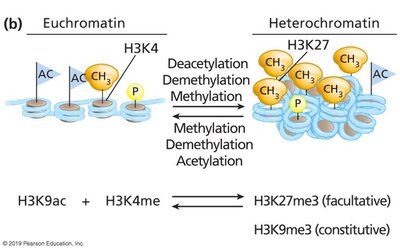
Chemical Modifications of Histones
Histone proteins can be chemically modified by acetylation and methylation, influencing chromatin structure and gene expression.
Acetylation (e.g., H3K9ac) is associated with euchromatin and active transcription.
Methylation (e.g., H3K27me3, H3K9me3) is associated with heterochromatin and gene silencing.
H3K4me is also associated with active chromatin.
DNA Methylation and CpG Islands
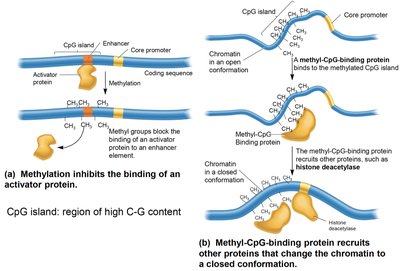
Methylation of cytosine residues in CpG dinucleotides can inhibit gene expression by blocking activator binding or recruiting proteins that condense chromatin. CpG islands are regions of high C-G content, often found near promoters.
Unmethylated CpG islands promote open chromatin and active transcription.
Methylated CpG islands lead to gene silencing.
Genomic Imprinting
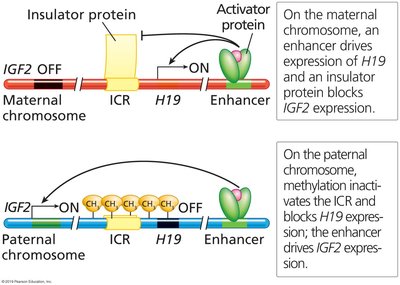
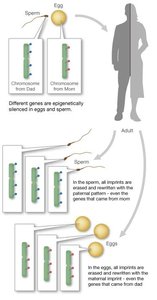
Genomic imprinting is an epigenetic phenomenon where only one parental allele of a gene is expressed. Imprints are erased and re-established in gametes each generation. Key examples include the IGF2 and H19 genes on human chromosome 11.
On the maternal chromosome, H19 is active and IGF2 is silenced by an insulator protein.
On the paternal chromosome, methylation inactivates the insulator, allowing IGF2 expression and silencing H19.
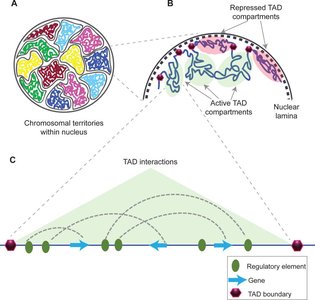
Topologically Associating Domains (TADs)
3D Genome Organization
Within the nucleus, chromosomes are organized into distinct regions called topologically associating domains (TADs). TADs are segments of DNA that preferentially interact with themselves, influencing gene regulation by spatially organizing regulatory elements and genes.
TADs can range from tens of thousands to millions of base pairs.
They help bring enhancers, promoters, and other regulatory elements into close proximity.
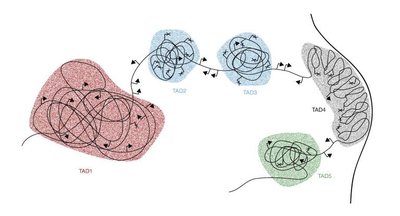
Mapping TADs: Hi-C Method
The Hi-C sequencing method is used to map 3D genome interactions by cross-linking DNA, digesting it, ligating fragments in close proximity, and sequencing the ligation products to create a contact map.
Hi-C reveals the spatial organization of the genome and identifies TAD boundaries.
RNA-Based Gene Silencing
RNA Interference (RNAi)
RNA interference (RNAi) is a mechanism of gene silencing mediated by small non-coding RNAs, such as siRNAs and miRNAs. These RNAs bind to complementary mRNA targets, leading to mRNA degradation or inhibition of translation.
RNAi can regulate endogenous genes or defend against viruses and transposable elements.
Key proteins: Drosha, Dicer, Argonaute, and RISC (RNA-induced silencing complex).
MicroRNAs (miRNAs)
miRNAs are processed from primary transcripts (pri-miRNAs) to precursor miRNAs (pre-miRNAs) in the nucleus, then to mature miRNA duplexes in the cytoplasm. The mature miRNA is loaded into RISC, which mediates gene silencing.
miRNAs can block translation or promote mRNA degradation.
Applications of RNAi
RNAi is used as a research tool to knock down gene expression and study gene function. It also has therapeutic potential for diseases caused by overexpression or abnormal expression of specific genes.
Post-Translational Regulation
Protein Modifications
After translation, proteins often undergo modifications that affect their function, localization, and stability. These include:
Addition of chemical groups (e.g., methyl, phosphate)
Amino acid modifications
Polypeptide cleavage
Interaction with other proteins
Addition of complex molecules (e.g., carbohydrates, lipids)
Feedback Inhibition
Feedback inhibition is a regulatory mechanism where the end product of a metabolic pathway inhibits an enzyme involved earlier in the pathway, thus controlling the rate of product synthesis. This is important in pathways such as the shikimate pathway for aromatic amino acid synthesis.
Summary of Key Points
Gene expression in eukaryotes is regulated at multiple levels: transcriptional, post-transcriptional, translational, and post-translational.
Regulatory sequences, transcription factors, and epigenetic modifications play central roles in controlling gene activity.
3D genome organization and RNA-based mechanisms add further layers of regulation.
Understanding these processes is essential for studying development, disease, and biotechnology applications.
