 Back
BackGene Mutation, DNA Repair, and Homologous Recombination: Study Notes
Study Guide - Smart Notes

Gene Mutation, DNA Repair, and Homologous Recombination
Introduction
This chapter explores the molecular mechanisms underlying gene mutation, the cellular processes that repair DNA damage, and the role of homologous recombination in maintaining genome integrity. Understanding these processes is fundamental to genetics, as mutations are the source of genetic variation and can lead to genetic diseases.
Mutations: Types and Mechanisms
Mutation Rates and Randomness
Gene mutations are rare, random events that alter DNA sequence.
Mutation rate can be measured by counting mutations affecting a phenotype or by determining the frequency per base pair.
Mutation rates vary among organisms and genes; mutation hotspots are genes with elevated mutation rates, often due to large gene size.
The Luria-Delbrück fluctuation test provided experimental proof that mutations arise randomly, not adaptively.
Types of Mutations
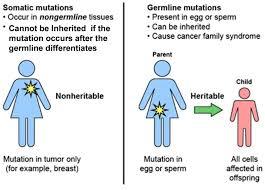
Germ-line mutations occur in gametes and are heritable; somatic mutations occur in non-gamete cells and are not inherited.
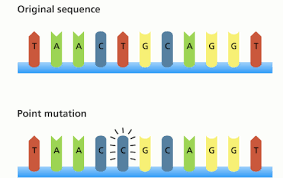
Point mutations involve changes at a specific nucleotide position.
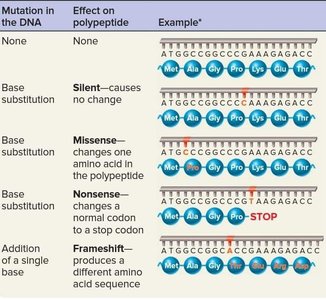
Base-pair substitution mutations include:
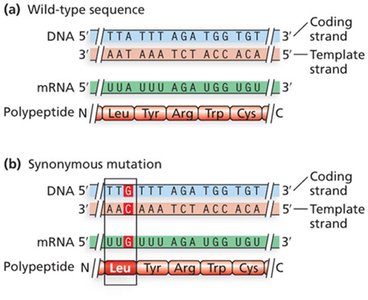
Synonymous (silent) mutations: Do not alter the amino acid sequence.
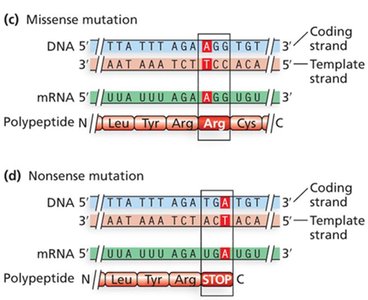
Missense mutations: Change one amino acid in the protein.
Nonsense mutations: Convert a codon to a stop codon, truncating the protein.
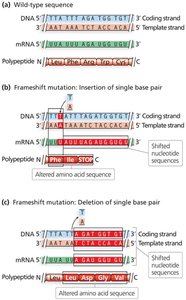
Frameshift mutations: Insertions or deletions that alter the reading frame, often producing nonfunctional proteins.
Regulatory mutations: Affect gene expression without altering the protein sequence (e.g., promoter, splicing, polyadenylation mutations).
Forward mutations convert wild-type alleles to mutant alleles; reversions restore wild-type or near wild-type function.
Regulatory Mutations
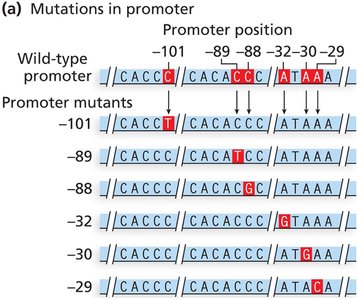
Promoter mutations: Alter consensus sequences, affecting transcription initiation and gene expression levels.
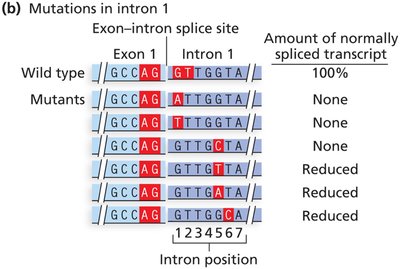
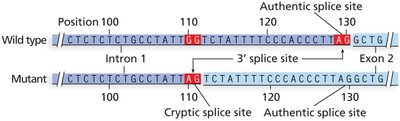
Splicing mutations: Affect intron removal, potentially leading to abnormal mRNA and protein products.
Cryptic splice sites: New splice sites created by mutation, leading to aberrant mRNA processing.
Polyadenylation mutations: Disrupt mRNA 3' end processing, reducing protein production.

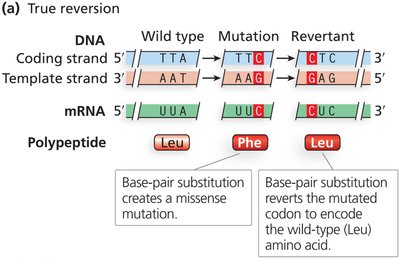
Forward Mutation and Reversion
True reversion: Second mutation restores the original DNA sequence.
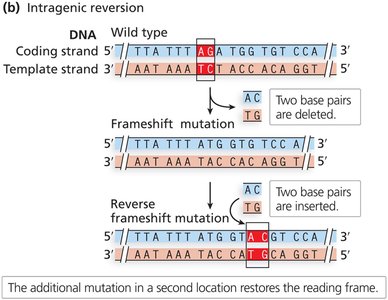
Intragenic reversion: Second mutation elsewhere in the same gene restores function.
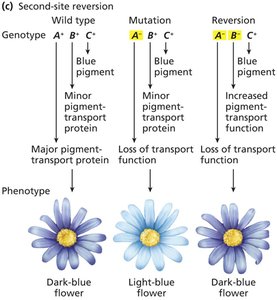
Second-site reversion (suppressor mutation): Mutation in a different gene compensates for the original mutation.
Origins of Gene Mutations
Spontaneous Mutations
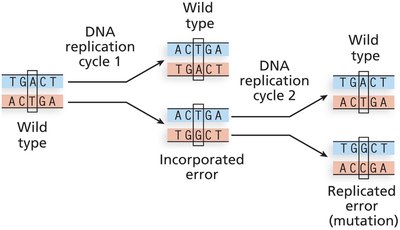
Occur without exposure to mutagens, mainly due to errors in DNA replication or spontaneous chemical changes.
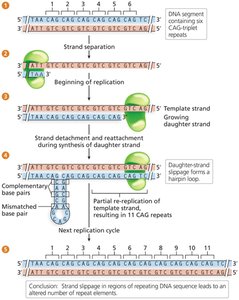
Strand slippage during replication can cause insertions or deletions, especially in regions with repeated sequences.
Trinucleotide repeat expansion disorders are caused by increased numbers of repeats due to strand slippage.
Mispaired nucleotides (non–Watson-Crick base pairing) can lead to mutations if not repaired.
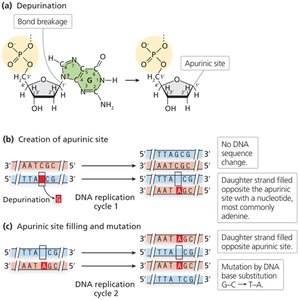
Depurination: Loss of a purine base, creating an apurinic site; if unrepaired, can result in mutation.
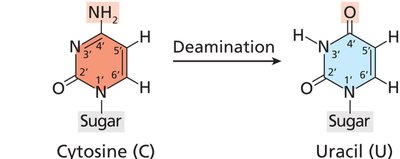
Deamination: Loss of an amino group from a base (e.g., cytosine to uracil); can lead to base substitutions if not repaired.
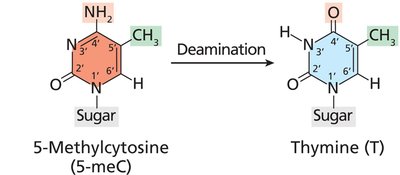
Deamination of methylated cytosine produces thymine, leading to G-C to A-T transitions if unrepaired.
Induced Mutations: Chemical and Physical Agents
Chemical Mutagens
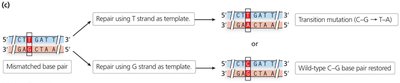
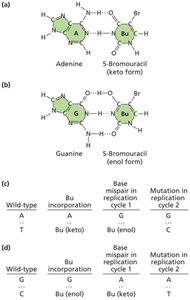
Nucleotide base analogs: Chemicals resembling DNA bases (e.g., 5-bromodeoxyuridine) can be incorporated into DNA, causing transition mutations.
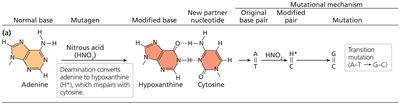
Deaminating agents: Remove amino groups from bases (e.g., nitrous acid), leading to base substitutions.
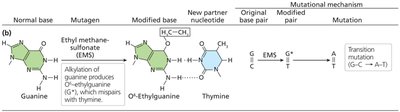
Alkylating agents: Add bulky groups to bases (e.g., ethyl methanesulfonate), distorting DNA and causing transitions.
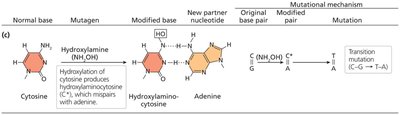
Hydroxylating agents: Add hydroxyl groups (e.g., hydroxylamine), causing C-G to T-A transitions.
Oxidizing agents: Cause transversion mutations by modifying bases.
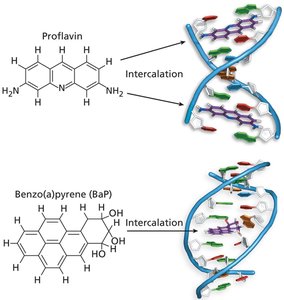
Intercalating agents: Insert between base pairs, causing frameshift mutations (e.g., proflavin, acridine orange).
Mutagen | Type of Agent | Mutagenic Event |
|---|---|---|
2-Aminopurine | Nucleotide base analog | Transition mutation |
5-Bromodeoxyuridine | Nucleotide base analog | Transition mutation |
Ethyl methanesulfonate | Alkylating agent | Transition mutation |
Hydroxylamine | Hydroxylating agent | Transition mutation |
Nitrous oxide | Deaminating agent | Transition mutation |
Oxygen radicals | Oxidizing agent | Transversion mutation |
Acridine orange | Intercalating agent | Frameshift mutation |
Proflavin | Intercalating agent | Frameshift mutation |
Radiation-Induced DNA Damage
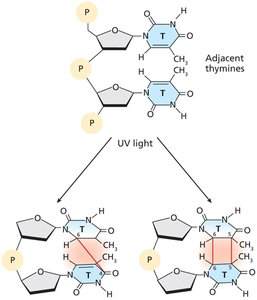
UV irradiation causes formation of photoproducts such as thymine dimers and 6-4 photoproducts, which distort DNA structure and block replication.
Unrepaired photoproducts can lead to mutations and are associated with skin cancer.
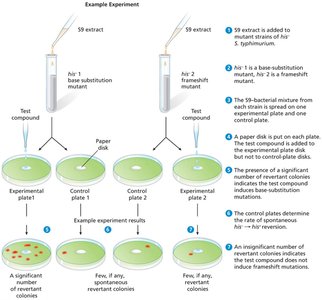
The Ames Test
The Ames test is used to assess the mutagenicity of chemical compounds by measuring the rate of reversion mutations in bacteria.
Bacteria are exposed to the test compound and mammalian liver enzymes; an increase in revertant colonies indicates mutagenicity.
DNA Repair Mechanisms
Direct Repair Systems
Proofreading by DNA polymerase corrects errors during replication.
Photoreactive repair (not in humans): Photolyase enzyme uses visible light to break bonds in UV-induced photoproducts.
Base excision repair (BER): DNA glycosylases remove damaged bases, AP endonuclease creates a nick, and DNA polymerase and ligase fill and seal the gap.
Nucleotide excision repair (NER): Removes segments containing bulky lesions (e.g., UV damage); DNA polymerase and ligase restore the strand.
Mismatch repair: Detects and repairs mismatches missed by DNA polymerase, using methylation to distinguish old and new strands.
Repair System | Main Function |
|---|---|
Photoreactive repair | Removes UV-induced photoproducts |
Base excision repair (BER) | Removes incorrect/damaged bases |
Nucleotide excision repair (NER) | Removes bulky DNA lesions |
Mismatch repair | Removes base-pair mismatches |
DNA Damage Signaling and the p53 Pathway
Cells use signaling systems to detect DNA damage and activate repair or cell cycle arrest.
ATM protein activates the p53 repair pathway, which can pause the cell cycle or trigger apoptosis if damage is irreparable.
Mutations in DNA repair genes (e.g., p53) increase cancer risk (e.g., Li-Fraumeni syndrome).
Translesion DNA Synthesis and Double-Strand Break Repair
Translesion DNA Synthesis (SOS Repair)
When replication is blocked by DNA damage, translesion DNA polymerases (error-prone) can synthesize DNA across lesions.
In E. coli, the SOS response involves RecA protein and pol V, allowing replication to continue at the cost of increased mutations.
Double-Strand Break Repair
Double-strand breaks (DSBs) are severe lesions that can cause chromosome instability and cell death.
Nonhomologous end joining (NHEJ): Error-prone repair mechanism that joins broken DNA ends, often causing mutations.
Synthesis-dependent strand annealing (SDSA): Error-free repair using a homologous template, typically a sister chromatid.
Summary Table: Types of Mutations and Repair Mechanisms
Mutation Type | Cause | Repair Mechanism |
|---|---|---|
Base substitution | Replication error, chemical mutagen | Proofreading, mismatch repair, BER, NER |
Frameshift | Strand slippage, intercalating agent | NER, mismatch repair |
Thymine dimer | UV irradiation | Photoreactive repair, NER |
Double-strand break | Ionizing radiation, replication fork collapse | NHEJ, SDSA |
Key Terms
Mutation: A heritable change in the DNA sequence.
Mutagen: An agent that increases the frequency of mutations.
Proofreading: The ability of DNA polymerase to remove incorrectly paired nucleotides during replication.
Apoptosis: Programmed cell death, often triggered by irreparable DNA damage.
