 Back
BackGenetic Analysis and Mapping in Bacteria and Bacteriophages
Study Guide - Smart Notes

Genetic Analysis and Mapping in Bacteria and Bacteriophages
Introduction
This chapter explores the specialized methods used for genetic analysis in bacteria and bacteriophages, focusing on gene transfer mechanisms, mapping strategies, and the evolutionary implications of lateral gene transfer. Bacteria and their viruses (bacteriophages) serve as powerful model systems for understanding fundamental genetic processes due to their simplicity, rapid growth, and genetic tractability.
Specialized Methods for Genetic Analysis of Bacteria
Advantages of Bacteria in Genetic Studies
Genome simplicity: Bacteria have fewer genes and smaller genomes compared to eukaryotes, facilitating genetic analysis.
Haploid genomes: Mutations are directly observable since only one gene copy is present.
Short generation times: Bacterial populations can double in minutes, allowing rapid experimentation.
Large numbers of progeny: Enables detection of rare genetic events.
Ease of propagation: Bacteria are easy and inexpensive to culture.
Numerous heritable differences: Mutants are easily created, identified, and manipulated.
Bacterial Culture and Growth Analysis
Bacteria reproduce by binary fission, producing genetically identical progeny.
Growth occurs in liquid or solid media containing essential nutrients (carbon, nitrogen, water, etc.).
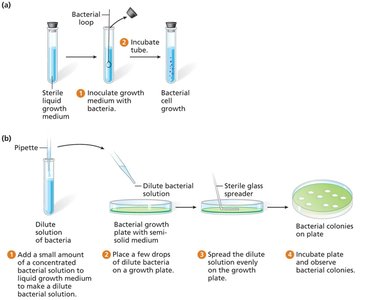
Types of Media
Minimal medium: Contains only essential nutrients (e.g., glucose, nitrogen, water, inorganic salts).
Prototrophs: Bacteria that can grow on minimal medium; they have no mutations blocking biosynthetic pathways.
Auxotrophs: Mutant bacteria that require additional nutrients (e.g., amino acids) due to biosynthetic defects; grow only on complete or supplemented media.
Complete medium: Contains all compounds required for growth and reproduction.
Plating and Replica Plating
Different media are used to distinguish bacterial genotypes based on growth patterns.
Replica plating: Technique to transfer cells from one plate to another, allowing identification of auxotrophs by their inability to grow on minimal medium.
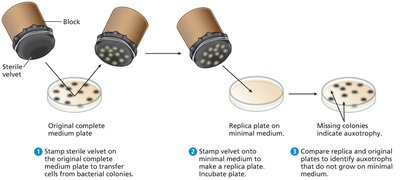
Bacterial Genomes and Plasmids
Bacterial genomes are typically a single, covalently closed circular DNA molecule containing essential genes.
Most bacteria also carry plasmids: small, circular, double-stranded DNA molecules with nonessential genes.
Plasmids can be high-copy-number (replicate independently) or low-copy-number (depend on the chromosome for replication).
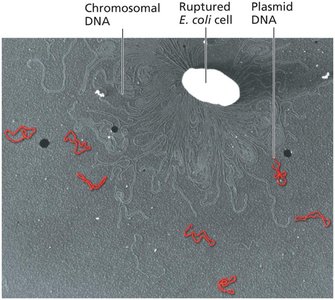
Types of Plasmids
F (fertility) plasmid: Carries genes for its own transfer between cells.
R (resistance) plasmid: Contains antibiotic resistance genes.
Plasmids are important tools in recombinant DNA technology.
Bacterial Gene Transfer Mechanisms
Overview of Gene Transfer
Conjugation: Direct transfer of DNA from donor to recipient via cell-to-cell contact.
Transformation: Uptake of free DNA from the environment by a recipient cell.
Transduction: Transfer of DNA from one bacterium to another via a bacteriophage (virus).
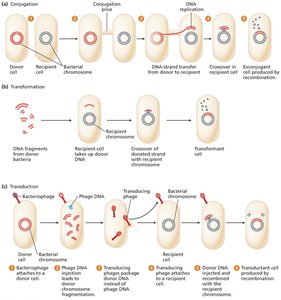
Bacterial Conjugation
Donor cells (F+) possess the F factor; recipient cells (F−) lack it.
Conjugation is mediated by genes on the F plasmid, which direct formation of a conjugation pilus and DNA transfer machinery.
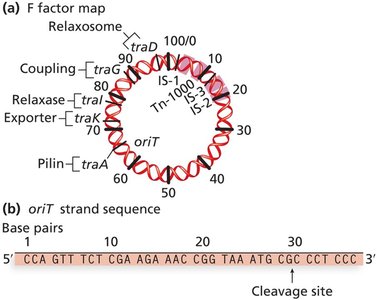
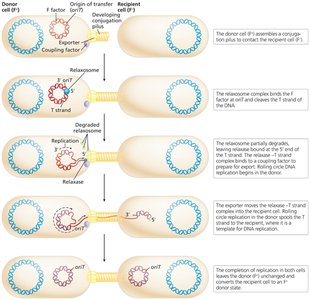
The relaxosome complex initiates transfer by nicking the F factor at the origin of transfer (oriT).
DNA is transferred via rolling circle replication, resulting in both cells containing the F factor.
Hfr Strains and Chromosome Transfer
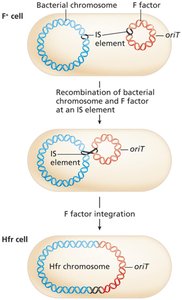
In Hfr (high frequency of recombination) strains, the F factor integrates into the bacterial chromosome at IS elements.
During conjugation, Hfr cells transfer chromosomal genes to recipients in a linear order starting from oriT.
Complete transfer of the chromosome is rare; only a segment is usually transferred.
Homologous recombination incorporates donor DNA into the recipient chromosome.
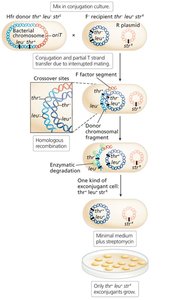
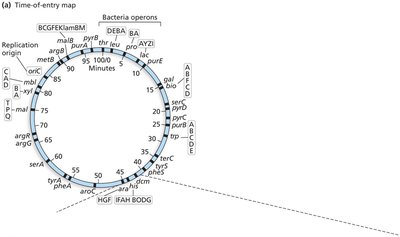
Selection and Mapping by Interrupted Mating
Selective media are used to identify exconjugants with specific genotypes.
Interrupted mating: Conjugation is stopped at intervals to determine the order and timing of gene transfer (time-of-entry mapping).
Genes closer to oriT are transferred earlier and more frequently.
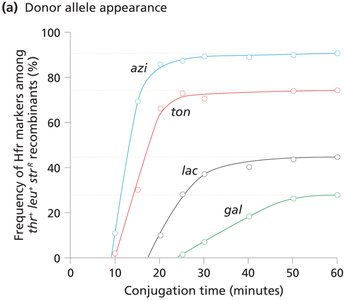
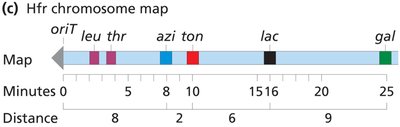
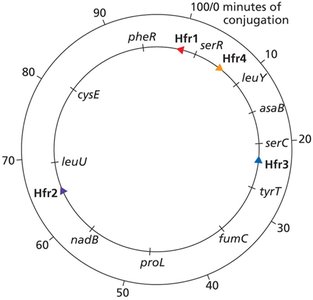
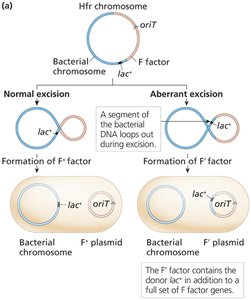
F' (F Prime) Factors and Partial Diploids
F' factors are formed by aberrant excision of the F factor from the Hfr chromosome, carrying additional bacterial genes.
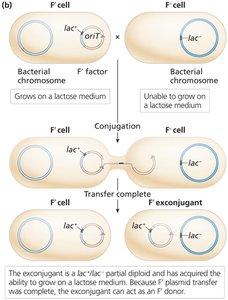
Conjugation with F' donors creates partial diploids (merodiploids), useful for studying gene regulation and dominance.
Bacterial Transformation
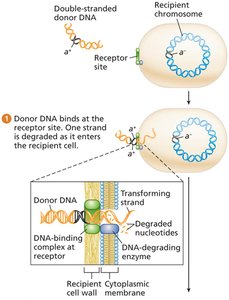
Mechanism of Transformation
Recipient cells take up DNA fragments from lysed donor cells in the environment.
One DNA strand is degraded during uptake; the other aligns with the recipient chromosome and may recombine, forming a heteroduplex.
After replication and cell division, one daughter cell is a transformant (genetically altered), the other is unchanged.
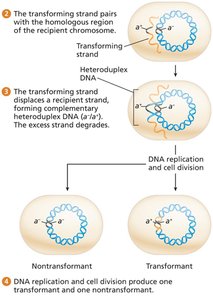
Mapping by Transformation
Transforming DNA is usually short (~100 kb), allowing mapping of closely linked genes.
Cotransformation: Simultaneous transformation of two or more genes; frequency reflects gene proximity.
Bacterial Transduction
Mechanism of Transduction
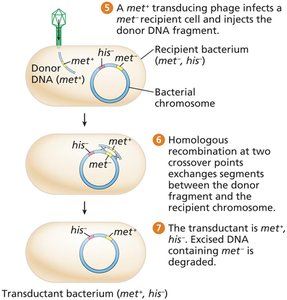
Transduction: Transfer of bacterial DNA by bacteriophages (viruses that infect bacteria).
Integration of donor DNA into the recipient chromosome by homologous recombination produces a transductant.
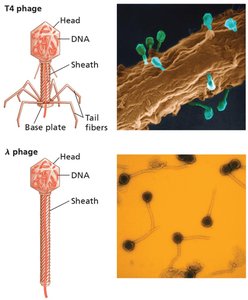
Bacteriophage Structure and Life Cycles
Bacteriophages have an icosahedral head, protein sheath, and sometimes tail fibers.
Two main life cycles: Lytic (host cell lysis and phage release) and Lysogenic (phage DNA integrates as a prophage).
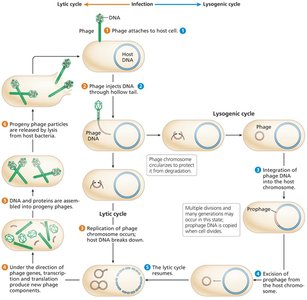
Generalized Transduction
Generalized transducing phages (e.g., P1) package random bacterial DNA fragments during assembly.
These phages inject donor DNA into new recipients, where recombination can occur.
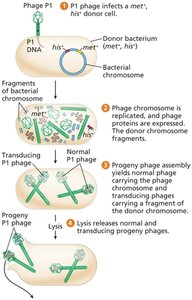
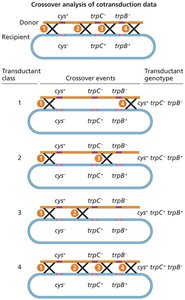
Cotransduction and Mapping
Genes close together are more likely to be cotransduced.
Cotransduction frequency is used to determine gene order and distance.
Two-step selection screens for cotransductants: select for one marker, then screen for the second.
Transductant Class | Transductant Genotype | Number |
|---|---|---|
1 | cys+ trpC− trpB− | 139 |
2 | cys+ trpC− trpB+ | 18 |
3 | cys+ trpC+ trpB+ | 141 |
4 | cys+ trpC+ trpB− | 4 |
TOTAL | 302 | |
Specialized Transduction
Temperate phages integrate at specific att sites in the host genome.
Imprecise excision can package adjacent bacterial genes into phage particles (specialized transducing phages).
Only genes near the integration site are transferred.
Lateral Gene Transfer (LGT) and Genome Evolution
Definition and Importance
Lateral gene transfer (LGT): Movement of genetic material between unrelated organisms, not by descent.
LGT contributes significantly to bacterial and archaeal genome content and evolution.
Genes for cell surface proteins, DNA-binding proteins, and pathogenicity are often acquired by LGT.
Identification of LGT
LGT regions, or genomic islands, are identified by sequence features distinct from the rest of the genome (e.g., unusual G-C content, similarity to distant species).
Medical Relevance
LGT enables rapid adaptation, such as acquisition of antibiotic resistance and pathogenicity islands.
Examples include E. coli strains with genes for toxins or host invasion acquired via LGT.
Summary Table: Bacterial Gene Transfer Mechanisms
Mechanism | Key Features | Genetic Outcome |
|---|---|---|
Conjugation | Direct cell-to-cell contact, F plasmid, rolling circle replication | Transfer of plasmid or chromosomal genes; formation of exconjugants |
Transformation | Uptake of free DNA from environment | Stable integration of donor DNA; formation of transformants |
Transduction | Phage-mediated DNA transfer | Generalized: random genes; Specialized: genes near att site |
Additional info: This chapter provides foundational knowledge for advanced topics such as recombinant DNA technology, genomics, and the molecular basis of bacterial evolution and pathogenesis.
