 Back
BackMutation and DNA Repair Mechanisms in Genetics
Study Guide - Smart Notes

Mutation: Causes and Types
Spontaneous vs. Induced Mutations
Mutations are heritable changes in the DNA sequence. They can arise spontaneously due to natural cellular processes or be induced by external factors such as chemicals or radiation.
Spontaneous mutations: Occur naturally during DNA replication or as a result of normal metabolic activities. These are rare events but are the primary source of genetic variation.
Induced mutations: Result from exposure to mutagens, which are physical or chemical agents that increase the mutation rate above the spontaneous background level.
Organism | Character | Locus | Rate* |
|---|---|---|---|
riophage T2 | Lysis inhibitBacteion | r+ → r | 1 × 10−8 |
Escherichia coli | Lactose fermentation | lac+ → lac | 2 × 10−8 |
Zea mays | Shrunken seeds | sh+ → sh | 1 × 10−7 |
Drosophila melanogaster | White eye | w+ → w | 1 × 10−6 |
Mus musculus | Piebald coat | p+ → p | 2 × 10−6 |

Additional info: Mutation rates are typically measured per gene per generation and are influenced by the organism and the specific locus.
taneSpoous Mutations: Mechanisms
Replication Errors and Slippage
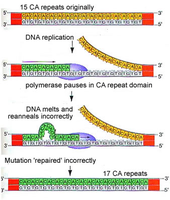
DNA replication is not perfectly accurate. DNA polymerase can incorporate incorrect nucleotides, and some errors escape proofreading, leading to point mutations. Replication slippage, especially in repetitive sequences, can cause insertions or deletions (indels), which are common in regions called mutation hot spots.
Replication slippage: Occurs when DNA polymerase slips on the template strand, causing loops that result in extra or missing nucleotides in the new strand.
Hot spots: Regions with repetitive sequences are more prone to slippage and mutation.
Clinical relevance: Diseases such as Fragile-X syndrome and Huntington disease are associated with repeat expansions due to replication slippage.
Tautomeric Shifts
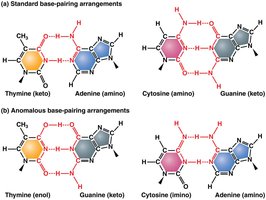
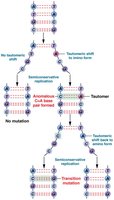
Nitrogenous bases in DNA can exist in alternative chemical forms called tautomers. Tautomeric shifts can cause non-standard base pairing during replication, leading to transition mutations.
Tautomers: Alternate forms of purines and pyrimidines that differ in the position of protons and bonding.
Effect: Tautomeric shifts can result in A–C or G–T base pairing instead of the normal A–T and G–C pairs.
Outcome: If not corrected, these mispairings become permanent mutations after subsequent rounds of replication.
DNA Base Damage: Depurination and Deamination
Spontaneous chemical changes can damage DNA bases, leading to mutations if not repaired.
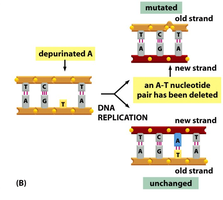
Depurination: Loss of a purine base (adenine or guanine) from the DNA, creating an apurinic site. During replication, this site may be skipped, resulting in a deletion.
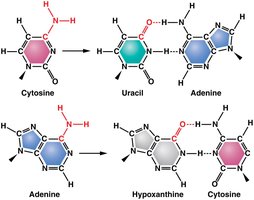
Deamination: Removal of an amino group from cytosine or adenine, converting cytosine to uracil or adenine to hypoxanthine. This alters base pairing and can result in transition mutations (e.g., A–T to G–C).
Oxidative Damge and Free Radicals
Reactive oxygen species (ROS), such as superoxides, hydroxyl radicals, and hydrogen peroxide, are by-products of normal metabolism and can damage DNA. Free radicals can alter bases, break phosphod Cellular respiration, radiation, and environmental toxins.
Prevention: Antioxidants in the diet can reduce oxidative damage.
Induced Mutations: Chemical and Physical Agents
Mutagens
Mutagens are agents that increase the frequency of mutations. They include chemicals, radiation, and biological agents.
Chemical mutagens: Base analogs, alkylating agents, intercalating agents, and adduct-forming agents.
Physical mutagens: Ultraviolet (UV) light and ionizing radiation (X-rays, gamma rays, cosmic rays).
Base Analogs
Base analogs are chemicals that resemble DNA bases and can be incorporated into DNA during replication. They often increase the rate of tautomeric shifts and mispairing.
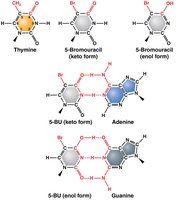
Example: 5-Bromouracil (5-BU) is an analog of thymine and can pair with guanine, leading to transition mutations.
Alkylating Agents
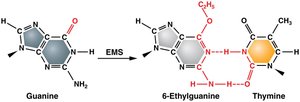
Alkylating agents add alkyl groups to DNA bases, altering their base-pairing properties and causing transition mutations.
Example: Ethyl methanesulfonate (EMS) and mustard gas are common alkylating agents used in genetics research.
Intercalating Agents
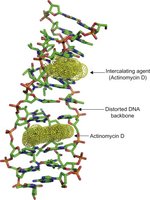
Intercalating agents are planar molecules that insert between DNA base pairs, distorting the double helix and causing insertions or deletions during replication.
Example: Ethidium bromide and acridine dyes are classic intercalating agents.
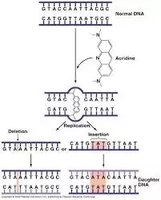
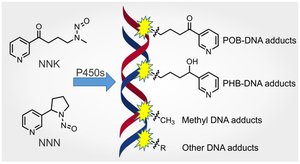
Adduct-Forming Agents
Adduct-forming agents covalently bind to DNA, altering its structure and interfering with replication and repair. Examples include acetaldehyde (from cigarette smoke) and heterocyclic amines (from cooked meats).
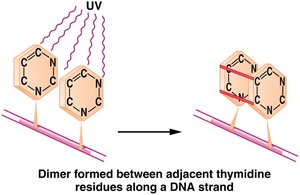
Ultraviolet (UV) Light
UV light induces the formation of pyrimidine dimers, especially thymine dimers, which distort the DNA helix and block replication.
Mechanism: Adjacent pyrimidines (usually thymines) become covalently linked, preventing normal base pairing.
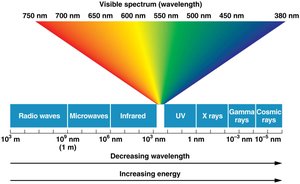
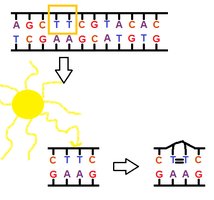
Ionizing Radiation
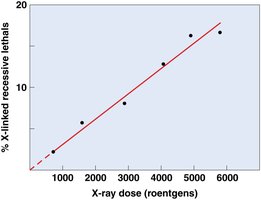
Ionizing radiation (X-rays, gamma rays, cosmic rays) has enough energy to break DNA strands and cause base modifications, deletions, and chromosomal rearrangements.
Effect: Can cause single- and double-strand breaks, leading to mutations or cell death.
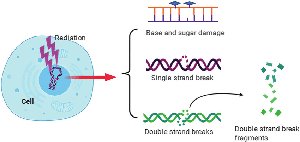
DNA Repair Mechanisms
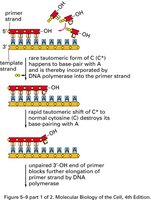
Proofreading and Mismatch Repair
DNA polymerase has proofreading activity that removes incorrectly paired nucleotides during replication. If errors escape proofreading, mismatch repair systems detect and correct them based on strand discrimination (e.g., methylation patterns in bacteria).
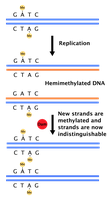
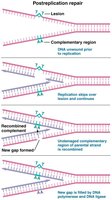
Postreplication Repair
Postreplication repair mechanisms, such as those involving the RecA protein in bacteria, use homologous recombination to fill gaps left by unrepaired lesions after replication.
SOS Repair System
The SOS repair system in E. coli is an emergency response to extensive DNA damage. It allows DNA replication to continue past lesions but is error-prone and can introduce mutations.
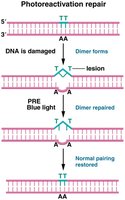
Photoreactivation Repair
Photoreactivation repair (PRE) uses light energy to directly reverse thymine dimers. This system is present in some organisms but absent in humans.
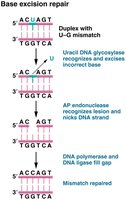
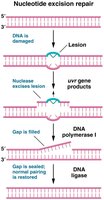
Base and Nucleotide Excision Repair
Excision repair mechanisms remove damaged bases or nucleotides and fill the gap with newly synthesized DNA.
Base excision repair (BER): Removes single damaged bases using DNA glycosylases.
Nucleotide excision repair (NER): Removes bulky lesions that distort the DNA helix, such as thymine dimers.
Xeroderma Pigmentosum (XP)
XP is a rare genetic disorder caused by defects in NER pathways. Affected individuals are extremely sensitive to UV light and have a high risk of skin cancer due to inability to repair thymine dimers.

Double-Strand Break Repair
Double-strand breaks (DSBs) are highly dangerous and can lead to chromosomal rearrangements or cell death. Two main pathways repair DSBs:
Homologous recombination repair: Uses a homologous sequence as a template for accurate repair, usually during late S or G2 phase.
Nonhomologous end joining (NHEJ): Directly ligates broken DNA ends without a template, which can be error-prone.
p
Definition and Effects
Transposable elements are DNA sequences that can move within or betweeTransposable Elements (Transosons)n genomes. They can disrupt gene function, cause chromosomal rearrangements, and contribute to genetic diversity and evolution.
Autonomous elements: Can move independently.
Nonautonomous elements: Require the presence of autonomous elements for movement.
Examples in Model Organisms
Ac-Ds system in maize: Discovered by Barbara McClintock, these elements cause variegated kernel color in corn.
P elements in Drosophila: Used as genetic tools for creating transgenic flies and studying gene function.
Human genome: Contains LINEs and SINEs, which are types of transposable elements. About half of the human genome is composed of transposable elements.
Additional info: Transposons can cause mutations by inserting into coding or regulatory regions, and their movement is a significant source of genetic variation.
