 Back
BackChapter 10
Study Guide - Smart Notes

Mutation: Overview and Types
Definition and Importance
Mutation refers to a heritable and permanent change in the DNA sequence of an organism's genome. Mutations are the original source of genetic diversity, fueling evolutionary change. Although most mutations are random and often deleterious, they are essential for adaptation and evolution.
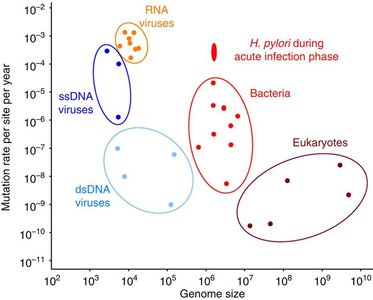
Mutation rates are generally low, varying among organisms and genes.
Factors such as genome size and life cycle affect mutation rates.
Mutations can occur due to errors in DNA replication, spontaneous chemical changes, or damage followed by incorrect repair.
Types of Mutations
Point mutations: Change in one or a few nucleotides.
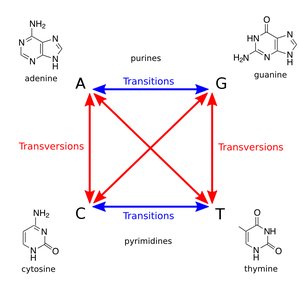
Transition mutations: Purine replaces purine (A ↔ G) or pyrimidine replaces pyrimidine (C ↔ T).
Transversion mutations: Purine replaces pyrimidine or vice versa (A/G ↔ C/T).
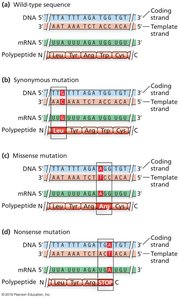
Point Mutations in Coding Regions
Point mutations can have different effects depending on their location and nature:
Silent (synonymous): No change in amino acid.
Missense: Change in amino acid.
Nonsense: Change to a stop codon.
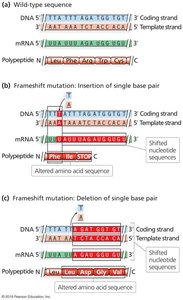
Frameshift Mutations
Frameshift mutations result from the insertion or deletion of nucleotides, altering the reading frame of the gene. These can introduce premature stop codons and are a subset of indels (insertions-deletions).
Not all indels cause frameshifts; only those not in multiples of three nucleotides.
Regulatory Mutations
Some point mutations affect gene regulation rather than protein sequence:
Promoter mutations: Alter consensus sequence, affecting transcription.
Splicing mutations: Affect intron-exon boundaries, possibly retaining introns in mRNA.
Cryptic splice sites: Create new splicing sites.
Polyadenylation mutations: Block poly-A tail addition.
Forward Mutation and Reversion
Forward mutation: Wild-type allele becomes mutant.
Reverse mutation (reversion): Mutant allele returns to wild-type or near wild-type.
True reversion: Second mutation restores original sequence.
Suppressor mutation: New mutation suppresses effects of earlier mutation, rescuing phenotype.
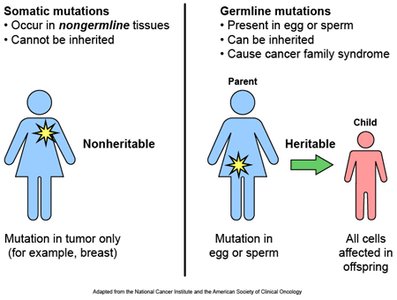
Somatic vs. Germ Line Mutations
Mutations can occur in somatic or germ-line cells:
Somatic mutations: Occur in non-reproductive tissues; not inherited.
Germ-line mutations: Occur in reproductive cells; can be passed to offspring.
Mechanisms of Mutation: Spontaneous vs. Induced
Spontaneous Mutations
Spontaneous mutations arise without exposure to external mutagens, mainly due to errors in DNA replication or spontaneous chemical changes.
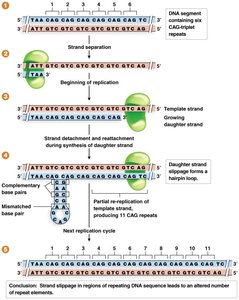
Repeat mutations: Alterations in DNA repeat number via strand slippage.
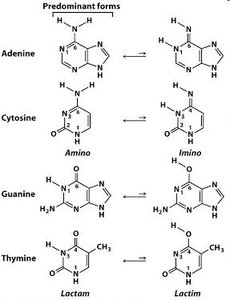
Tautomeric shifts: Temporary changes in base structure cause mispairing.
Depurination: Loss of purine base, leaving apurinic site.
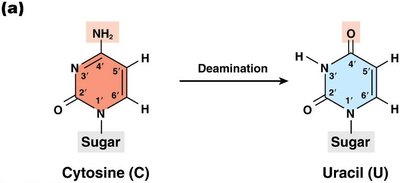
Deamination: Loss of amino group, often converting cytosine to uracil.
Trinucleotide Repeat Disorders
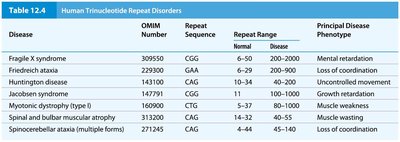
Trinucleotide repeat disorders are caused by expansions of repeat sequences beyond a threshold, leading to diseases such as Fragile X syndrome and Huntington disease.
Disease | Repeat Sequence | Repeat Range (Normal) | Repeat Range (Disease) | Principal Disease Phenotype |
|---|---|---|---|---|
Fragile X syndrome | CGG | 6-50 | 200-2000 | Mental retardation |
Friedreich ataxia | GAA | 6-33 | 200-2000 | Loss of coordination |
Huntington disease | CAG | 10-34 | 40-100 | Uncontrolled movement |
Myotonic dystrophy (type I) | CTG | 5-37 | 80-1500 | Muscle weakness |
Spinal and bulbar muscular atrophy | CAG | 12-30 | 40-62 | Muscle wasting |
Spinocerebellar ataxia | CAG | 4-44 | 45-140 | Loss of coordination |

Induced Mutations
Induced mutations result from interactions with physical, chemical, or biological agents (mutagens).
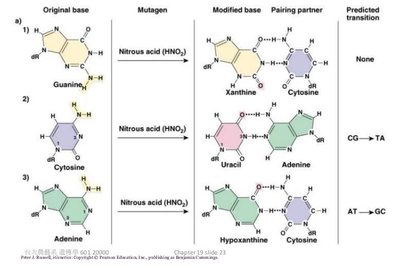
Deaminating agents: Remove amine groups, causing mispairing.
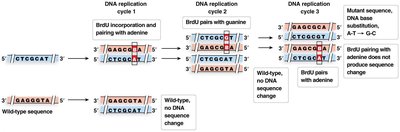
Nucleotide base analogs: Mimic normal bases, causing mispairing.
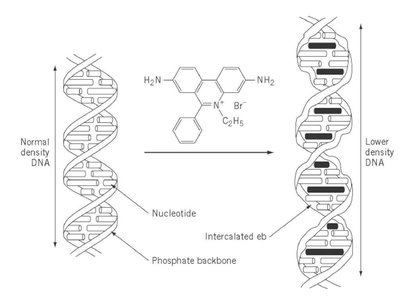
Intercalating agents: Insert between base pairs, causing frameshifts.
Oxidizing, alkylating, and hydroxylating agents: Modify bases, causing mispairing or strand breaks.
Physical Mutagens
Physical mutagens such as UV light and ionizing radiation cause DNA damage:
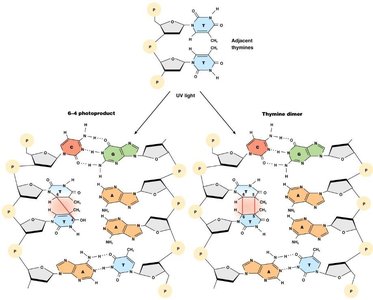
UV irradiation: Causes pyrimidine dimers and photoproducts.
X-rays and radioactive materials: Cause single- and double-strand breaks.
DNA Repair Mechanisms
Base Excision Repair (BER)
BER corrects single-base damage by removing the damaged base and replacing it with the correct nucleotide.
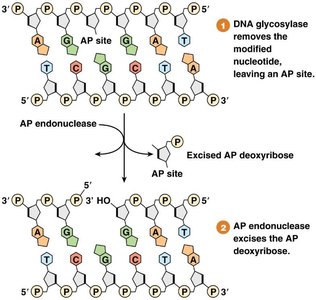
DNA glycosylase removes the base, leaving an apurinic/apyrimidinic (AP) site.
AP endonuclease removes the sugar-phosphate backbone.
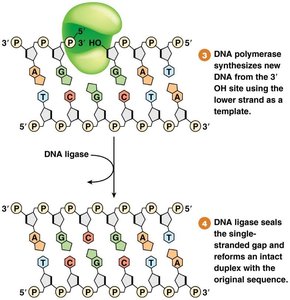
DNA polymerase and ligase fill the gap and seal the backbone.
Mismatch Repair (MMR)
MMR corrects errors that escape proofreading during DNA replication. Enzymes distinguish the original strand by methylation and remove the mismatched nucleotide.
Exonuclease digests through the mismatch.
DNA polymerase fills the gap, ligase seals it, and Dam methylase methylates the repaired strand.
Nucleotide Excision Repair (NER)
NER removes bulky adducts and covalent modifications caused by UV light, excising a short stretch of nucleotides from the damaged strand.
DNA polymerase and ligase fill and seal the gap.
Photoreactive Repair
Photoreactive repair directly reverses pyrimidine dimers using photolyase and visible light. This mechanism is present in bacteria, single-celled eukaryotes, plants, and some animals, but not humans.
Double-Strand Break Repair
Double-strand breaks are repaired by two main mechanisms:
Nonhomologous end joining (NHEJ): Joins broken ends, often resulting in mutations.
Synthesis-dependent strand annealing (SDSA): Uses sister chromatid as template for error-free repair.
Homologous Recombination and Gene Conversion
Homologous Recombination
Homologous recombination is the exchange of genetic material between homologous DNA molecules, important for double-strand break repair and meiosis.
In bacteria, occurs via RecBCD pathway.
In eukaryotes, occurs during prophase I of meiosis.
Double-Strand Break Model of Meiotic Recombination
Recombination is initiated by a double-strand break, followed by strand invasion, D-loop formation, and Holliday junction resolution.
Resolution of Holliday junctions can result in gene conversion or recombinant chromosomes.
Gene Conversion
Gene conversion occurs when mismatches in heteroduplex DNA are repaired, resulting in conversion of one allele to another and homozygosity.
The Ames Test: Mutagenicity Assay
Principle and Application
The Ames test is used to assess the mutagenicity of compounds using mutant strains of Salmonella typhimurium that cannot synthesize histidine. Reversion mutations restore the ability to grow without histidine, indicating mutagenicity.
Detects both substitution and frameshift mutations.
Reverse mutations indicate a compound is mutagenic.
Summary Table: Mutation Types and Effects
Mutation Type | Definition | Effect |
|---|---|---|
Point mutation | Change in one or few nucleotides | Silent, missense, or nonsense |
Frameshift mutation | Insertion/deletion altering reading frame | Premature stop codon, altered protein |
Regulatory mutation | Change in promoter, intron, UTR | Altered gene expression |
Repeat mutation | Expansion/contraction of repeat sequences | Trinucleotide repeat disorders |
Key Equations and Concepts
Mutation rate:
Transition vs. Transversion:
Additional info: Academic context and explanations have been expanded for clarity and completeness. Only directly relevant images and tables have been included to reinforce key concepts.
