 Back
BackRegulation of Gene Expression in Eukaryotes: Mechanisms and Molecular Strategies
Study Guide - Smart Notes

Regulation of Gene Expression in Eukaryotes
Overview of Eukaryotic Gene Regulation
Eukaryotic gene expression is a highly regulated process, allowing cells to produce gene products at specific times, in specific cell types, and in response to environmental changes. Unlike prokaryotes, eukaryotes possess complex regulatory mechanisms due to their larger genome, chromosomal organization, spatial separation of transcription and translation, mRNA processing, RNA stability, and cellular differentiation.
Temporal regulation: Genes are expressed at particular times during development or in response to stimuli.
Spatial regulation: Expression occurs in specific cell types or tissues.
Environmental regulation: Gene expression adapts to external conditions.
Levels of Gene Regulation
Gene regulation in eukaryotes can occur at any step from DNA to protein product, including:
Chromatin remodeling
Transcriptional regulation
Posttranscriptional regulation (RNA processing, mRNA stability)
Translational regulation
Posttranslational modification
Chromosome Organization and Chromatin Modifications
Chromosome Territories and Nuclear Architecture
During interphase, each chromosome occupies a distinct domain within the nucleus, known as a chromosome territory. This spatial organization facilitates gene regulation by cycling transcriptionally active genes to the edges of territories, where they can interact with transcription factors in "transcription factories." Channels between chromosomes, called interchromosomal domains, contain little or no DNA.
Chromatin Structure and Remodeling
Eukaryotic DNA is packaged with histone and nonhistone proteins to form chromatin. Chromatin can be remodeled to regulate gene expression in two general ways:
Modification of nucleosomes
Modification of DNA
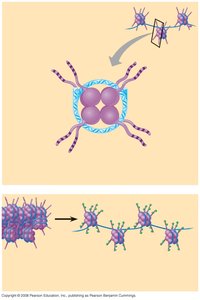
Nucleosome Remodeling Mechanisms
Nucleosomal chromatin can be remodeled by:
Altering nucleosome composition: Variant histones (e.g., H2A.Z, H3.3) flank promoters of active genes.
Histone modification: Covalent modifications of histone tails, including acetylation, phosphorylation, and methylation.
Repositioning nucleosomes: Remodeling complexes (e.g., SWI/SNF) shift nucleosomes to expose regulatory regions.
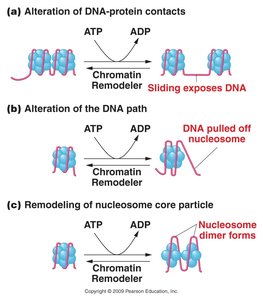
Histone Modifications
Acetylation: Addition of acetyl groups by histone acetyltransferases (HATs) to lysine residues, neutralizing positive charge and opening chromatin for transcription. Histone deacetylases (HDACs) remove acetyl groups, leading to chromatin condensation and gene repression.
Phosphorylation: Addition of phosphate groups by kinases to serine/histidine residues, introducing negative charge.
Methylation: Addition of methyl groups by methyltransferases to lysine/arginine residues, often linked to gene repression.
Histone code: The pattern of covalent modifications determines gene activation or silencing.
DNA Methylation
DNA methylation involves the addition of methyl groups to cytosine bases, typically at CpG dinucleotides. Heavily methylated promoter regions (CpG islands) are associated with gene silencing. Methylation can inhibit transcription factor binding and recruit repressive chromatin remodeling complexes.
Transcriptional Regulation: Cis-Acting Elements and Transcription Factors
Cis-Acting Elements
Cis-acting elements are DNA sequences that regulate the expression of genes located on the same chromosome. Key cis-acting elements include:
Promoters: Recognition sites for transcription machinery, located adjacent to regulated genes.
Enhancers: Modular DNA sequences that increase transcription rates, located at variable distances from the gene.
Silencers: DNA elements that repress transcription initiation.
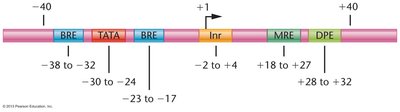
Promoter Structure and Function
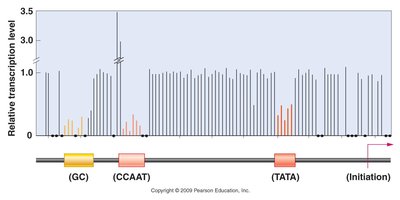
Promoters consist of core elements (e.g., TATA box, BRE, Inr, MRE, DPE) that bind specific initiation proteins. Proximal promoter elements (e.g., CAAT boxes, GC boxes) enhance basal transcription levels by binding transcription factors.
Enhancers and Silencers
Enhancers contain multiple short DNA sequences that increase transcription rates and can be located upstream, downstream, or within the gene. Silencers repress transcription initiation. Both function by interacting with transcription factors and regulatory proteins.
Transcription Factors and Their Functional Domains
Transcription factors are proteins that regulate gene expression by binding to cis-acting sites. They possess two main domains:
DNA-binding domain: Recognizes specific DNA sequences.
Trans-activating domain: Activates or represses transcription via protein-protein interactions.
Common DNA-binding motifs include:
Helix-turn-helix
Zinc finger
Leucine zipper
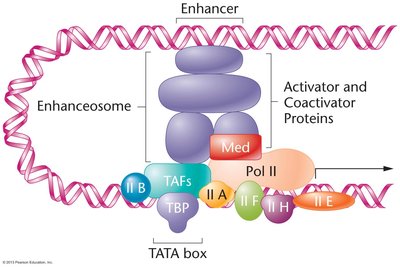
Transcription Initiation Complex
Basal (general) transcription factors assemble at the promoter, enabling RNA polymerase II binding. TFIID, containing the TATA binding protein (TBP), is the first to bind the TATA box, followed by other factors and RNA polymerase II.
Enhanceosome Formation and Regulation
Transcription activators and coactivators form a complex called the enhanceosome, which interacts with general transcription factors to enhance transcription initiation. Activators/repressors can also modify chromatin structure or directly interact with transcription factors to regulate initiation rates.
Posttranscriptional Regulation
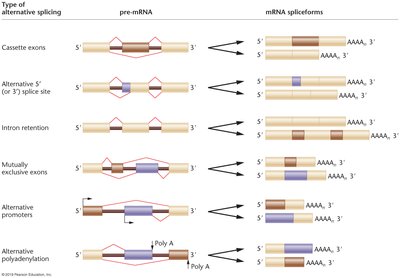
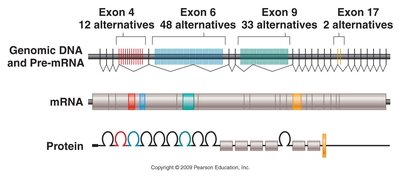
Alternative RNA Splicing
Alternative splicing generates different protein isoforms from the same pre-mRNA by varying the set and order of exons incorporated into the mature mRNA. This increases protein diversity and functional complexity.
Control of mRNA Stability
The steady-state level of mRNA in a cell is determined by the rate of transcription and degradation. Eukaryotic mRNA is generally more stable than prokaryotic mRNA, with stability influenced by sequences in the 5' and 3' untranslated regions (UTRs).
Poly-A tail shortening: Reduces mRNA stability, leading to degradation.
Decapping: Removal of the 5' cap decreases stability.
Endonucleolytic cleavage: Internal cleavage (e.g., nonsense-mediated decay) exposes mRNA ends to exonucleases.

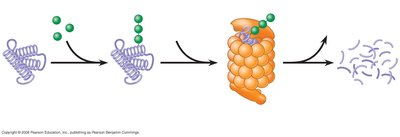
Protein Stability and Posttranslational Modification
After translation, proteins undergo various modifications, including cleavage, phosphorylation, methylation, acetylation, glycosylation, and complexing with metals. These modifications are critical for protein function. Proteasomes degrade proteins tagged with ubiquitin, regulating protein levels and activity.
RNA Silencing and RNA Interference (RNAi)
RNA-Induced Gene Silencing
Short RNA molecules (siRNAs and miRNAs) regulate gene expression by repressing translation and triggering mRNA degradation. This sequence-specific regulation is known as RNA interference (RNAi).
siRNAs: Arise from viral infection or exogenous sources, processed by Dicer.
miRNAs: Endogenous noncoding RNAs that negatively regulate gene expression.
RISC: RNA-induced silencing complex, containing Argonaut proteins, mediates mRNA destruction.
Therapeutic Applications of RNAi
RNAi technology is being developed as a pharmaceutical agent to reduce expression of dominant mutant genes or abnormal gene products. Promising results have been observed in reducing the severity of diseases such as HIV, influenza, and Huntington's disease.
Summary Table: Mechanisms of Eukaryotic Gene Regulation
Level | Mechanism | Key Features |
|---|---|---|
Chromatin Remodeling | Histone modification, nucleosome repositioning, DNA methylation | Regulates accessibility of DNA to transcription machinery |
Transcriptional Regulation | Cis-acting elements, transcription factors, enhanceosome | Controls initiation and rate of transcription |
Posttranscriptional Regulation | Alternative splicing, mRNA stability | Determines mRNA isoforms and longevity |
Translational Regulation | RNA silencing (RNAi) | Represses translation, triggers mRNA degradation |
Posttranslational Modification | Protein processing, proteasome degradation | Modifies protein function and stability |
