 Back
BackRegulation of Gene Expression in Eukaryotes
Study Guide - Smart Notes

Regulation of Gene Expression in Eukaryotes
Overview of Eukaryotic Gene Regulation
Gene expression in eukaryotes is a highly regulated process, allowing cells to produce gene products at specific times, in specific cell types, and in response to environmental changes. Unlike prokaryotes, eukaryotic gene regulation is more complex due to the larger genome, chromosomal organization, spatial separation of transcription and translation, mRNA processing, RNA stability, and cellular differentiation.
Temporal regulation: Genes are expressed at particular developmental stages or in response to signals.
Spatial regulation: Genes are expressed in specific cell types or tissues.
Environmental response: Gene expression can change in response to external stimuli.
Chromosome Organization and Chromatin Modifications
Chromosome Territories and Nuclear Architecture
During interphase, each chromosome occupies a distinct region in the nucleus called a chromosome territory. These territories are separated by interchromosomal domains, which are regions with little or no DNA. Chromosome structure is dynamic, and transcriptionally active genes are often relocated to the edges of these territories, where they can interact with transcription factors in specialized regions called transcription factories.
Chromatin Structure and Remodeling
Eukaryotic DNA is packaged with histone and nonhistone proteins to form chromatin. Chromatin structure can be remodeled to regulate gene expression by making DNA more or less accessible to transcription machinery.
Nucleosome: The basic unit of chromatin, consisting of DNA wrapped around histone proteins.
Chromatin remodeling: The dynamic modification of chromatin architecture to allow access to condensed genomic DNA.
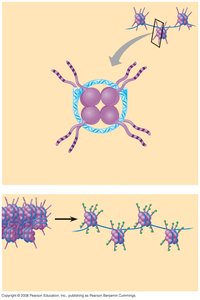
Mechanisms of Chromatin Remodeling
Chromatin can be remodeled in several ways:
Altering nucleosome composition: Incorporation of histone variants (e.g., H2A.Z, H3.3) at promoters of active genes.
Histone modification: Covalent addition or removal of chemical groups (acetyl, phosphate, methyl) to histone tails, affecting chromatin structure and gene accessibility.
Nucleosome repositioning: Sliding or restructuring nucleosomes to expose or hide DNA regions.
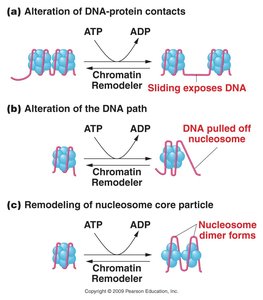
Histone Modifications
Acetylation: Addition of acetyl groups by histone acetyltransferases (HATs) neutralizes positive charges on lysines, loosening DNA-histone interaction and promoting gene activation. Histone deacetylases (HDACs) remove acetyl groups, leading to gene repression.
Phosphorylation: Addition of phosphate groups by kinases to serine or histidine residues, introducing negative charges and altering chromatin structure.
Methylation: Addition of methyl groups by methyltransferases to lysine or arginine residues, often associated with gene repression.
The combination of these modifications constitutes the histone code, which determines gene activation or silencing.
DNA Methylation
DNA methylation involves the addition of methyl groups to cytosine residues, typically in CpG dinucleotides. Heavily methylated promoter regions (CpG islands) are associated with gene silencing. DNA methylation can inhibit transcription factor binding and recruit repressive chromatin remodeling complexes.
Example: The inactive X chromosome in females is more heavily methylated than the active X chromosome.
Regulation at Cis-Acting Sites
Cis-Acting Elements
Cis-acting elements are DNA sequences that regulate the expression of genes located on the same chromosome. Major types include:
Promoters: Regions immediately upstream of genes that serve as binding sites for transcription machinery.
Enhancers: DNA elements that increase transcription rates, can be located far from the gene.
Silencers: DNA elements that repress transcription initiation.
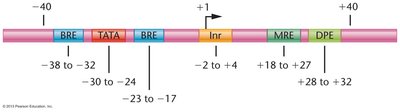
Promoter Structure and Function
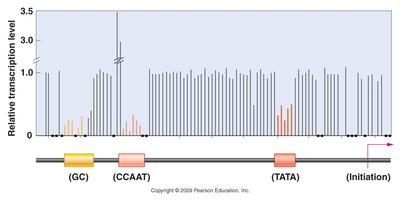
Promoters are composed of core elements (e.g., TATA box, BRE, Inr, DPE) and proximal promoter elements (e.g., CAAT box, GC box) that enhance basal transcription. Mutational analyses demonstrate the importance of specific promoter sequences for transcription efficiency.
Enhancers and Silencers
Enhancers and silencers are modular DNA sequences that can be located upstream, downstream, or within a gene. Enhancers increase, while silencers decrease, the rate of transcription initiation by interacting with specific transcription factors.
Transcription Factors and the Transcription Initiation Complex
Transcription Factors
Transcription factors are proteins that bind to cis-acting elements to regulate gene expression. They can act as activators or repressors and often have tissue-specific or temporally regulated expression. Many require structural modification for activation and may compete for DNA binding.
DNA-binding domain: Recognizes specific DNA sequences.
Trans-activating domain: Interacts with other proteins to activate or repress transcription.
Common DNA-binding motifs include helix-turn-helix, zinc finger, and leucine zipper.
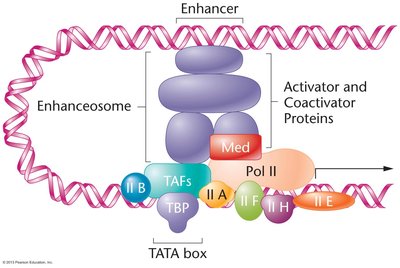
Formation of the Transcription Initiation Complex
Basal (general) transcription factors are required for the assembly of the transcription initiation complex at the promoter. TFIID, containing the TATA-binding protein (TBP), is the first to bind, followed by other factors and RNA polymerase II. Activators and coactivators can form an enhanceosome that interacts with the initiation complex to modulate transcription rates.
Posttranscriptional Gene Regulation
Alternative RNA Splicing
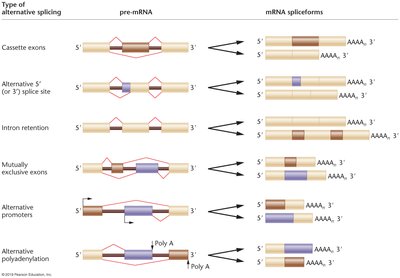
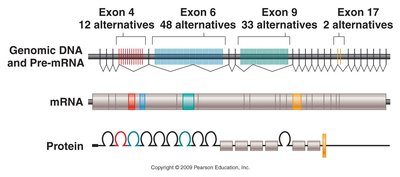
Alternative splicing allows a single gene to produce multiple mRNA and protein isoforms by varying the combination of exons included in the mature mRNA. This increases protein diversity without increasing gene number.
Types of alternative splicing: Cassette exons, alternative splice sites, intron retention, mutually exclusive exons, alternative promoters, and alternative polyadenylation.
Control of mRNA Stability
The steady-state level of mRNA in a cell is determined by the rates of transcription and degradation. mRNA stability is influenced by sequences in the 5' and 3' untranslated regions (UTRs), the length of the poly(A) tail, and the presence of specific binding proteins.
Degradation pathways: Shortening of the poly(A) tail, removal of the 5' cap, and endonucleolytic cleavage (e.g., nonsense-mediated decay).

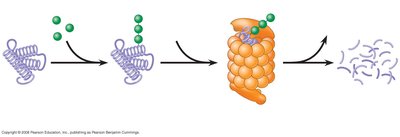
Protein Stability and Posttranslational Modification
After translation, proteins may undergo various modifications that affect their stability, localization, and function. Proteins can be tagged with ubiquitin and degraded by the proteasome, a large protein complex responsible for regulated protein turnover.
Posttranslational modifications: N-terminal modification, phosphorylation, methylation, acetylation, glycosylation, cleavage, and complex formation with metals.
RNA Silencing and RNA Interference (RNAi)
Mechanisms of RNA-Induced Gene Silencing
Short RNA molecules, such as siRNAs and miRNAs, regulate gene expression posttranscriptionally by repressing translation or triggering mRNA degradation. This process is known as RNA interference (RNAi) and involves several key proteins:
Drosha: Processes primary miRNA in the nucleus.
Dicer: Cleaves double-stranded RNA into ~22 nucleotide fragments.
RISC (RNA-induced silencing complex): Binds siRNA or miRNA and mediates mRNA cleavage or translational repression.
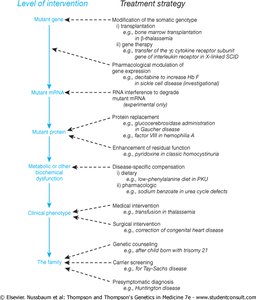
RNAi technology is being developed as a therapeutic strategy to silence disease-causing genes, such as mutant alleles in dominant genetic disorders.
Summary Table: Key Mechanisms of Eukaryotic Gene Regulation
Level of Regulation | Mechanism | Effect |
|---|---|---|
Chromatin Remodeling | Histone modification, nucleosome repositioning, DNA methylation | Alters DNA accessibility for transcription |
Transcriptional | Cis-acting elements, transcription factors, enhanceosome formation | Regulates initiation and rate of transcription |
Posttranscriptional | Alternative splicing, mRNA stability, RNA silencing | Determines mRNA and protein diversity and abundance |
Translational/Posttranslational | Protein modification, ubiquitin-mediated degradation | Controls protein function and turnover |
