 Back
BackRegulation of Gene Expression in Prokaryotes: The lac and trp Operons
Study Guide - Smart Notes

Regulation of Gene Expression in Prokaryotes
Overview of Gene Regulation
Gene expression in prokaryotes, such as E. coli, is tightly regulated to ensure that cellular resources are used efficiently. Bacteria respond to environmental changes by turning genes on or off, allowing them to produce only the proteins needed at any given time. Regulation occurs at the level of transcription and can be achieved through feedback inhibition or gene regulation mechanisms.
Inducible enzymes: Produced only when specific substrates are present.
Constitutive enzymes: Produced continuously, regardless of environmental conditions.
Repressible enzymes: Not produced when a specific molecule is present.
Regulation may be under negative control (expression occurs unless shut off by a regulator) or positive control (transcription occurs only if stimulated by a regulator).
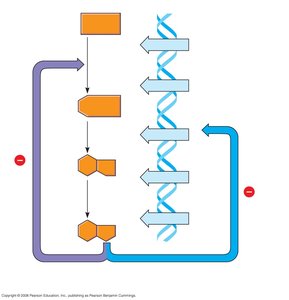
The Operon Model
Definition and Structure
An operon is a cluster of functionally related genes under coordinated control by a single regulatory region. The regulatory "switch" is the operator, usually located within the promoter. The operon includes the operator, promoter, and structural genes.
Operator: DNA segment where a repressor protein binds to regulate gene expression.
Promoter: DNA region where RNA polymerase binds to initiate transcription.
Structural genes: Genes encoding proteins with related functions.
The lac Operon in E. coli
Components and Function
The lac operon is an inducible system regulating lactose metabolism. It consists of three structural genes (lacZ, lacY, lacA) and an upstream regulatory region (operator and promoter). All three genes are transcribed as a single unit, resulting in a polycistronic mRNA.
lacZ: Encodes β-galactosidase, which converts lactose to glucose and galactose.
lacY: Encodes permease, facilitating lactose entry into the cell.
lacA: Encodes transacetylase, involved in removing toxic by-products.
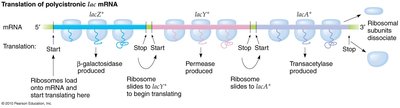
Regulatory Regions
The operator interacts with a specific repressor protein to regulate gene expression, while the promoter is the binding site for RNA polymerase. The lacI gene produces a repressor molecule that regulates transcription of the structural genes. The repressor is allosteric, meaning it changes shape and activity upon binding to another molecule (lactose).
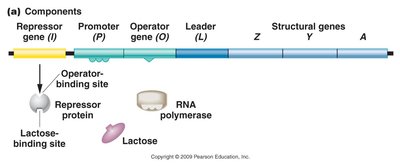
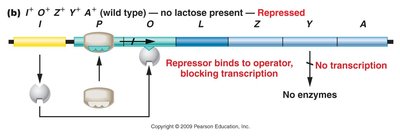
Negative Control of the lac Operon
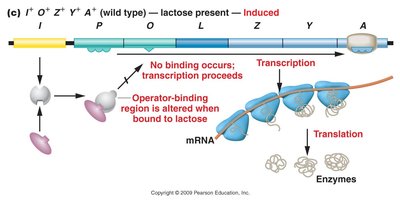
Transcription occurs only when the repressor fails to bind to the operator region. In the absence of lactose, the repressor binds to the operator, blocking transcription. When lactose is present, it binds to the repressor, causing a conformational change that prevents the repressor from binding the operator, allowing transcription to proceed.
Mutations Affecting lac Operon Regulation
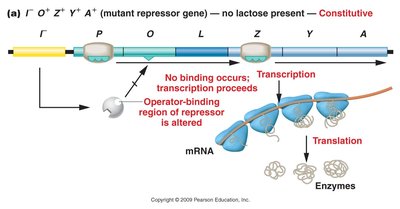
Mutations in the repressor gene or operator gene can lead to constitutive expression (continuous transcription regardless of lactose presence) or permanent repression.
Mutant repressor gene: Repressor cannot bind operator; transcription proceeds.
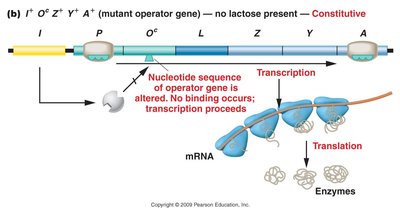
Mutant operator gene: Operator cannot bind repressor; transcription proceeds.
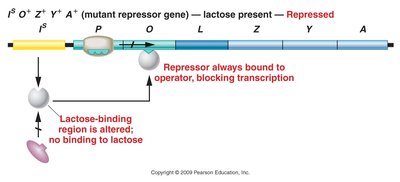
Mutant repressor (lactose-binding site): Repressor always binds operator; transcription is blocked even in presence of lactose.
Positive Control: Catabolite Repression and CAP
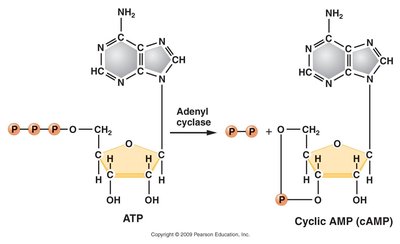
The catabolite-activating protein (CAP) exerts positive control over the lac operon. In the presence of glucose, CAP does not bind the promoter efficiently, and transcription is repressed. CAP requires cyclic adenosine monophosphate (cAMP) to bind the promoter. Glucose inhibits adenylyl cyclase, reducing cAMP levels and preventing CAP binding.
Maximal expression: Occurs when repressor is not bound and CAP is bound at the CAP-binding site.
The trp Operon in E. coli
Components and Function
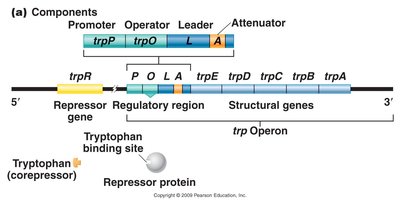
The trp operon is a repressible gene system regulating tryptophan biosynthesis. It consists of five structural genes (trpE, trpD, trpC, trpB, trpA), a regulatory region (promoter, operator, leader, attenuator), and a repressor gene (trpR).
Corepressor: Tryptophan acts as a corepressor, enabling the repressor protein to bind the operator and inhibit transcription.
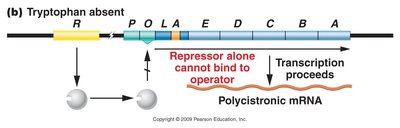
Regulation of the trp Operon
In the absence of tryptophan, the repressor cannot bind the operator, and transcription proceeds. In the presence of tryptophan, the repressor binds the operator, blocking transcription.

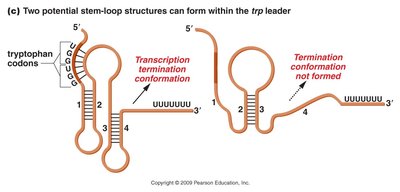
Attenuation Mechanism
Even when the trp operon is repressed, transcription initiation can occur, producing a leader sequence. Attenuation is a form of repression where transcription is greatly reduced but not entirely prevented. The leader region can form different hairpin structures depending on tryptophan availability:
Presence of tryptophan: Terminator hairpin forms, halting transcription.
Absence of tryptophan: Antiterminator hairpin forms, allowing transcription to proceed.

Key Terms and Concepts
Operator: Regulatory region to which repressor protein binds.
Operon: Adjacent genes serving one function, transcribed and regulated together.
Polycistronic mRNA: Single mRNA molecule encoding multiple proteins.
Inducer: Molecule (e.g., lactose) that inactivates the repressor and allows transcription.
Corepressor: Molecule (e.g., tryptophan) that activates the repressor to inhibit transcription.
Summary Table: Regulation of lac Operon
Lactose | Glucose | lac Operon Activity |
|---|---|---|
Absent | Absent | Low |
Present | Absent | High |
Absent | Present | Low |
Present | Present | Low |
Equations
Conversion of ATP to cAMP:
Transcriptional regulation:
Attenuation mechanism:
Example: In the lac operon, a nonsense mutation in the lacZ gene results in loss of β-galactosidase activity, preventing lactose metabolism.
