 Back
BackCH16: Regulation of Gene Expression in Prokaryotes: The lac and trp Operons
Study Guide - Smart Notes

Regulation of Gene Expression in Prokaryotes
Overview of Gene Regulation
Gene expression in prokaryotes, such as E. coli, is tightly regulated to ensure that cellular resources are used efficiently. Bacteria respond to environmental changes by turning genes on or off, allowing them to produce only the proteins needed at any given time. This regulation occurs at the level of transcription and can involve feedback inhibition or gene regulation mechanisms.
Inducible enzymes: Produced only when specific substrates are present.
Constitutive enzymes: Produced continuously, regardless of environmental conditions.
Repressible enzymes: Not produced when a specific molecule is present.
Regulation may be under negative control (expression occurs unless shut off by a regulator) or positive control (expression occurs only if stimulated by a regulator).
The lac Operon: An Inducible System
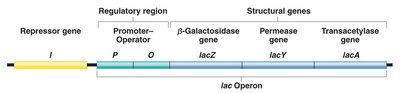
Structure and Function of the lac Operon
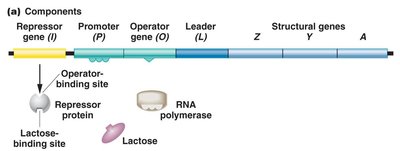
The lac operon in E. coli is a classic example of an inducible gene system. It consists of three structural genes (lacZ, lacY, lacA) and an upstream regulatory region (operator and promoter). These genes are transcribed as a single polycistronic mRNA, allowing coordinated expression.
lacZ: Encodes β-galactosidase, which cleaves lactose into glucose and galactose.
lacY: Encodes permease, which facilitates lactose entry into the cell.
lacA: Encodes transacetylase, involved in removing toxic by-products.
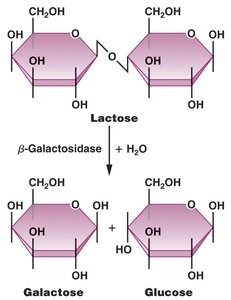
β-Galactosidase Activity
β-Galactosidase catalyzes the hydrolysis of lactose into glucose and galactose, enabling the cell to utilize lactose as an energy source.
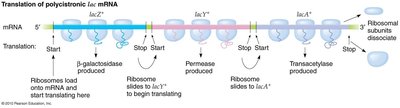
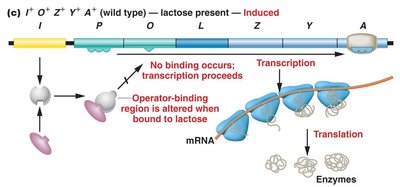
Polycistronic mRNA and Translation
All three structural genes are transcribed together, resulting in a polycistronic mRNA that is translated to produce the three enzymes.
Regulation of the lac Operon: Negative Control
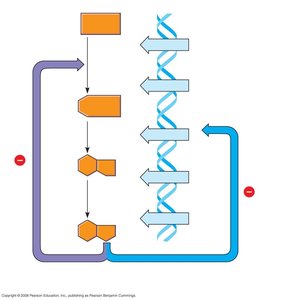
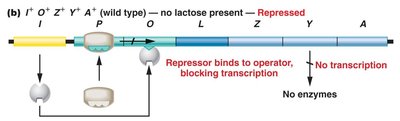
The lac operon is regulated by the lacI gene, which produces a repressor protein. In the absence of lactose, the repressor binds to the operator, blocking transcription. When lactose is present, it binds to the repressor, causing a conformational change that prevents the repressor from binding the operator, allowing transcription to proceed.
Negative control: Transcription occurs only when the repressor is not bound to the operator.
Genetic Proof of the Operon Model
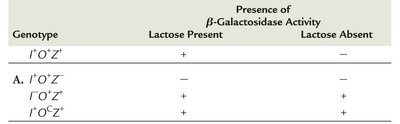
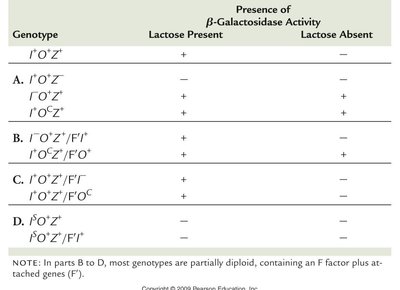
Genetic experiments using partial diploid merozygotes provided evidence for the operon model. The lacI gene produces a trans-acting product, while the operator region regulates transcription but does not encode a protein. The operator must be adjacent to the structural genes to regulate their expression.
Genotype | Lactose Present | Lactose Absent |
|---|---|---|
I+O+Z+ | + | − |
I−O+Z+ | + | + |
I+OcZ+ | + | + |

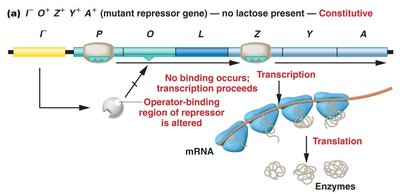

Mutations Affecting lac Operon Regulation
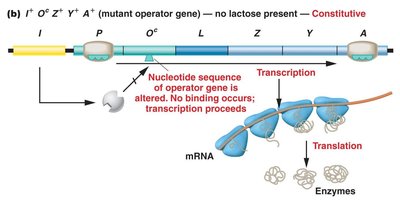
Mutations in the repressor gene (lacI) or operator (lacO) can lead to constitutive expression, where transcription occurs regardless of lactose presence.
lacI− mutation: Repressor cannot bind operator; transcription proceeds.
lacOc mutation: Operator sequence altered; repressor cannot bind; transcription proceeds.


Positive Control: Catabolite Repression and CAP
Role of Catabolite-Activating Protein (CAP)
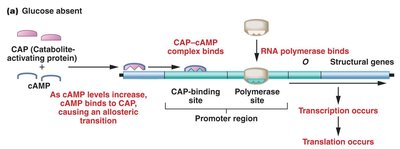
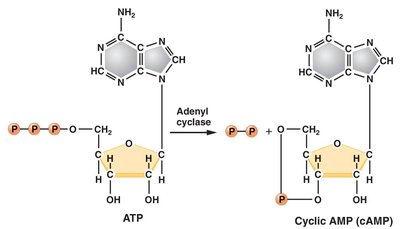
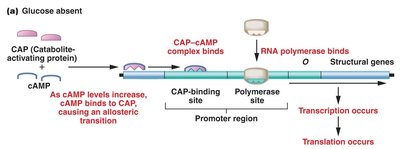
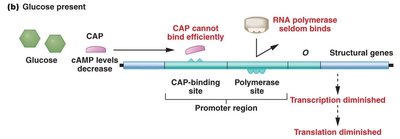
CAP exerts positive control over the lac operon. In the absence of glucose, cAMP levels rise, and the CAP-cAMP complex binds to the CAP-binding site, facilitating RNA polymerase binding and transcription. When glucose is present, cAMP levels fall, CAP cannot bind efficiently, and transcription is diminished.
Catabolite repression: Glucose inhibits expression of the lac operon.
CAP-cAMP complex: Required for maximal transcription of the lac operon.
The trp Operon: A Repressible System
Structure and Function of the trp Operon
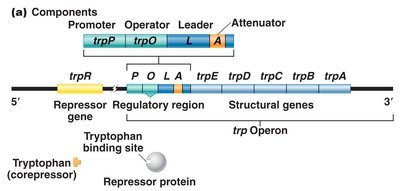
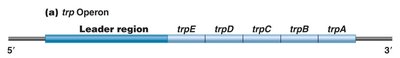
The trp operon in E. coli is a repressible gene system responsible for the biosynthesis of tryptophan. It consists of five structural genes (trpE, trpD, trpC, trpB, trpA) and a regulatory region (promoter, operator, leader, attenuator).
trpR: Encodes the repressor protein.
trpO: Operator region.
trpP: Promoter region.
trpE-trpA: Structural genes for tryptophan biosynthesis.
Regulation of the trp Operon
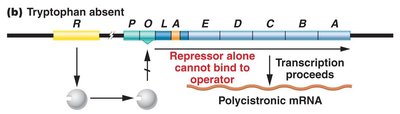
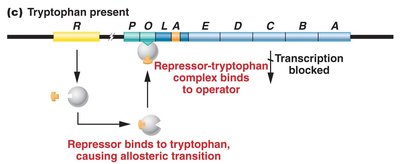
In the absence of tryptophan, the repressor cannot bind the operator, and transcription proceeds. When tryptophan is present, it acts as a corepressor, binding to the repressor and enabling it to bind the operator, blocking transcription.
Repressible system: Transcription is blocked in the presence of tryptophan.
Attenuation Mechanism
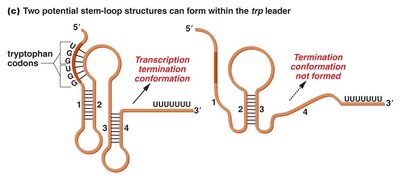
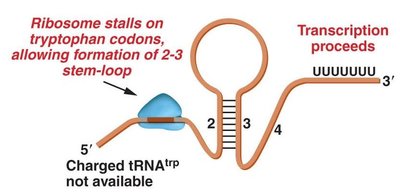
Attenuation is a form of repression in which transcription of the trp operon is greatly reduced but not entirely prevented. The leader region of the operon can form different stem-loop structures depending on tryptophan availability. In the presence of tryptophan, a terminator hairpin forms, halting transcription. In its absence, an antiterminator hairpin forms, allowing transcription to proceed.
Leader region: Contains two tryptophan codons.
Attenuator: Sequence that can form stem-loop structures to regulate transcription.

Key Terms and Concepts
Operator: Regulatory region to which repressor protein binds.
Operon: Adjacent genes serving one function that are transcribed and regulated together.
Polycistronic mRNA: Single mRNA molecule encoding multiple proteins.
Inducer: Molecule that inactivates the repressor to allow transcription (e.g., lactose).
Corepressor: Molecule required for the repressor to bind the operator (e.g., tryptophan).
Summary Table: lac Operon Activity Under Different Conditions
Lactose | Glucose | lac Operon Activity |
|---|---|---|
Absent | Absent | Low |
Present | Absent | High |
Absent | Present | Low |
Present | Present | Low |
Additional info: These notes expand on the original content by providing definitions, context, and examples for each regulatory mechanism, as well as summarizing the genetic and biochemical evidence supporting the operon model.
