Back
BackRegulation of Gene Expression: The trp Operon and Eukaryotic Mechanisms
Study Guide - Smart Notes

Regulation of Gene Expression in Prokaryotes and Eukaryotes
Overview
Gene expression is tightly regulated in both prokaryotic and eukaryotic cells to ensure that genes are expressed only when needed. This regulation is essential for cellular function, adaptation to environmental changes, and development. In prokaryotes, gene regulation is often linked to metabolic needs, while eukaryotes employ multiple, complex regulatory mechanisms at various levels.
The Tryptophan (trp) Operon in E. coli
Structure and Function
The trp operon in Escherichia coli is a classic example of a repressible gene system. It consists of five contiguous structural genes (trpE, trpD, trpC, trpB, trpA) that encode enzymes required for the biosynthesis of tryptophan. These genes are transcribed as a single polycistronic mRNA.
Promoter (trpP): Site where RNA polymerase binds to initiate transcription.
Operator (trpO): Binding site for the trp repressor protein.
Leader and Attenuator: Regulatory sequences upstream of the structural genes that further modulate expression.
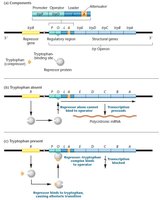
Mechanism of Regulation
In the absence of tryptophan: The repressor protein is inactive and cannot bind to the operator, allowing transcription of the operon and synthesis of tryptophan.
In the presence of tryptophan: Tryptophan acts as a corepressor, binding to the repressor protein and causing an allosteric change. The repressor-tryptophan complex binds to the operator, blocking transcription.
This negative feedback mechanism ensures that tryptophan is synthesized only when it is not available from the environment.
Comparison of Gene Regulation: Prokaryotes vs. Eukaryotes
Key Differences
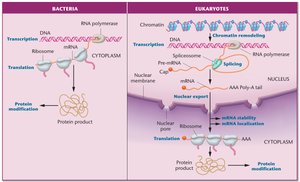
Prokaryotes: Gene regulation is primarily at the transcriptional level and is closely linked to metabolic needs. Transcription and translation occur simultaneously in the cytoplasm.
Eukaryotes: Regulation occurs at multiple levels, including chromatin structure, transcription, RNA processing, and translation. Transcription occurs in the nucleus, and mRNA is processed (capping, splicing, polyadenylation) before export to the cytoplasm for translation.
Regulation of Gene Expression in Eukaryotes
Chromatin Structure and Modifications
Eukaryotic DNA is packaged into chromatin, which consists of DNA wrapped around histone proteins to form nucleosomes. Chromatin structure can regulate gene accessibility:
Closed chromatin: DNA is tightly associated with histones, inhibiting access by transcription machinery.
Open chromatin: Chromatin remodeling and histone modifications (e.g., acetylation, methylation, phosphorylation) can relax chromatin, allowing transcription to proceed.
Histone acetyltransferases (HATs) add acetyl groups to histones, increasing transcription, while histone deacetylases (HDACs) remove them, repressing transcription.
DNA Methylation
DNA methylation, typically at cytosine residues in CpG islands, is a heritable epigenetic modification that represses gene expression. High levels of methylation in promoter regions are associated with gene silencing.
Transcriptional Regulation: Promoters, Enhancers, and Silencers
Transcription initiation in eukaryotes requires the interaction of multiple regulatory DNA elements and proteins:
Promoters: DNA sequences where RNA polymerase and general transcription factors bind to initiate transcription. Core promoters specify the transcription start site.
Proximal-promoter elements: Located upstream of the core promoter, these elements enhance basal transcription levels (e.g., CAAT box, GC box).
Enhancers: Cis-acting elements that increase transcription rates and can function at a distance from the gene.
Silencers: Cis-acting elements that repress transcription, often in a tissue- or time-specific manner.
Types of Core Promoters
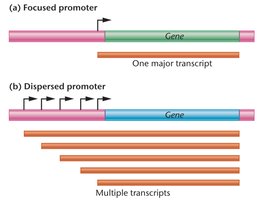
Focused core promoters: Direct transcription initiation at a single site; common in lower eukaryotes.
Dispersed core promoters: Direct initiation at multiple weak sites over a region; common in higher eukaryotes.
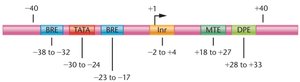
Core-Promoter Elements
Focused promoters contain several conserved elements:
Initiator (Inr) element
TATA box
TFIIB recognition element (BRE)
Motif ten element (MTE)
Downstream promoter element (DPE)
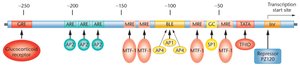
Promoter and Enhancer Interactions
Promoters and enhancers contain binding sites for transcription factors and regulatory proteins. The human metallothionein 2A (MT2A) gene is an example where multiple cis-acting elements and transcription factors regulate both basal and induced transcription in response to heavy metals and stress hormones.
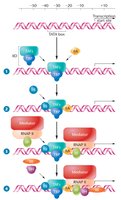
Pre-Initiation Complex (PIC) Formation
The assembly of the pre-initiation complex (PIC) is a critical step in transcription initiation. The PIC includes RNA polymerase II and general transcription factors (GTFs), which assemble at the core promoter. The first step is the binding of TFIID to the TATA box via the TATA-binding protein (TBP).
Mechanisms of Transcription Activation and Repression
Transcription activators and repressors modulate gene expression by interacting with enhancers, silencers, and the transcription machinery. DNA looping allows distant regulatory elements to interact with the promoter region, facilitating or inhibiting PIC assembly and transcription initiation.
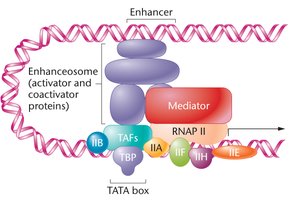
Coactivators and Enhanceosomes
Coactivators serve as bridges between activators bound to enhancers and the general transcription factors at the promoter. Enhanceosomes are large complexes of activators and coactivators that direct transcription activation. Repressor proteins at silencer elements decrease the rate of PIC assembly and RNA polymerase release.
Post-Transcriptional Regulation: Alternative Splicing
Alternative Splicing and Isoforms
Alternative splicing of pre-mRNA allows a single gene to produce multiple protein variants (isoforms) by including or excluding different exons. This process increases protein diversity and allows for tissue- or developmental stage-specific protein functions.
Example: The Drosophila Mhc gene encodes a motor protein with isoforms that differ in contractile properties, affecting wing-beat frequency.
Protein Degradation: Ubiquitin-Mediated Pathway
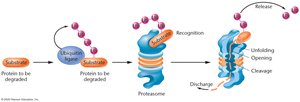
Ubiquitin-Proteasome System
Protein degradation is an important regulatory mechanism. Ubiquitin ligase enzymes attach ubiquitin molecules to substrate proteins, marking them for degradation by the proteasome. The proteasome recognizes ubiquitinated proteins, removes the ubiquitin tags, unfolds the protein, and cleaves it into small peptides.