 Back
Back(W 6 Slides) RNA and Protein Technologies in Genetics: Transcriptomics, cDNA, Microarrays, and Proteomics
Study Guide - Smart Notes

RNA Technologies and Transcriptomics
Central Dogma of Molecular Biology
The central dogma describes the flow of genetic information within a biological system. It explains how DNA is transcribed into RNA, which is then translated into protein. This process is fundamental to understanding gene expression and regulation.
Replication: DNA makes a copy of itself.
Transcription: DNA is used as a template to synthesize RNA.
Translation: RNA is used as a template to synthesize proteins.
Reverse Transcription: RNA can be reverse-transcribed back into DNA (e.g., by retroviruses).

Equation:
Types of RNA Involved in Protein Synthesis
Multiple RNA types play distinct roles in the process of protein synthesis:
mRNA (messenger RNA): Carries genetic instructions from DNA to ribosomes for protein synthesis.
tRNA (transfer RNA): Transports amino acids to ribosomes and matches them to the coded mRNA message.
rRNA (ribosomal RNA): Forms the core of ribosome's structure and catalyzes protein synthesis.
snRNA (small nuclear RNA): Involved in pre-mRNA splicing.
Other non-coding RNAs: Includes siRNA, miRNA, lncRNA, etc., with regulatory and structural roles.

Relationship Between RNA and Protein Sequences
RNA acts as the intermediary between DNA and proteins. The sequence of nucleotides in mRNA determines the sequence of amino acids in a protein, which in turn affects protein abundance and function.
mRNA: Specifies the amino acid sequence of a polypeptide chain.
tRNA: Each tRNA is linked to a specific amino acid and helps incorporate it into the growing polypeptide.
snRNA: Assists in processing pre-mRNA into mature mRNA.
Transcriptome and Transcriptomics
The Transcriptome
The transcriptome is the complete set of RNA transcripts produced by the genome under specific circumstances or in a specific cell. It includes mRNAs, tRNAs, rRNAs, and various non-coding RNAs. The transcriptome is dynamic and varies with time, environment, tissue, and cell type.
Regulation: Changes in the transcriptome are controlled by cis- and trans-regulatory elements, such as repressors, activators, and transcription factors.
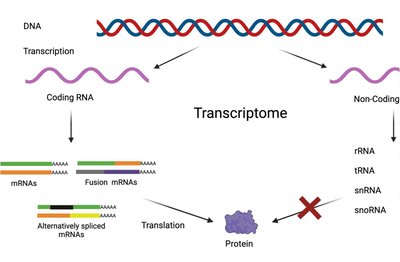
Transcriptomics
Transcriptomics is the study of the transcriptome, focusing on RNA sequences, structures, functions, locations, expression levels, and degradation. It is also known as expression profiling, as it examines all RNA transcripts expressed in a cell.
Omics Hierarchy: Transcriptomics is part of the broader 'omics' studies, which include genomics (DNA), proteomics (proteins), and metabolomics (metabolites).

Transcriptomics Aims
Catalogue all species of transcripts, including mRNAs, noncoding RNAs, and small RNAs.
Determine the transcriptional structure of genes (start sites, splicing, modifications).
Quantify changing expression levels of each transcript during development and under different conditions.

cDNA: Complementary DNA
Definition and Synthesis
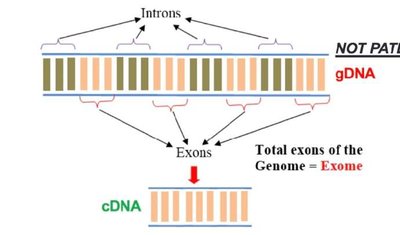
cDNA (complementary DNA) is a DNA copy synthesized from an mRNA template via the process of reverse transcription, catalyzed by the enzyme reverse transcriptase. cDNA is used in various molecular biology applications because it represents only the expressed genes (exons) of an organism.
Relation to DNA and mRNA: cDNA is synthesized from mature mRNA, which means it lacks introns present in genomic DNA (gDNA).

Steps of cDNA Synthesis
Isolation of mRNA from cells.
Hybridization with oligo-dT primers to bind the polyA tail of mRNA.
Reverse transcription to synthesize the first cDNA strand.
Removal of RNA template using RNase.
Synthesis of the second DNA strand by DNA polymerase.
Comparison: Genomic DNA vs. cDNA
Feature | Genomic DNA (gDNA) | cDNA |
|---|---|---|
Contains introns | Yes | No |
Represents | Entire genome | Expressed genes only |
Source | Genomic extraction | Reverse transcription from mRNA |
Importance of cDNA
Gene expression studies
Cloning and recombinant DNA technology
Quantitative PCR (qPCR)
Microarrays
Microarrays
Principle and Workflow
Microarrays are high-throughput techniques used to analyze the expression of thousands of genes simultaneously. They rely on nucleic acid hybridization, where cDNA samples are labeled with fluorescent dyes and hybridized to DNA probes fixed on a solid surface.
Each spot on the array contains a known DNA sequence (probe).
Fluorescently labeled cDNA from samples hybridizes to complementary probes.
Fluorescence intensity at each spot indicates the expression level of the corresponding gene.

Applications of Microarrays
Transcriptome profiling
Gene expression analysis
Disease diagnosis and biomarker discovery
Drug discovery and pharmacogenomics
Limitations of Microarrays
Limited to known sequences (requires prior knowledge of gene sequences).
Cross-hybridization and non-specific binding can occur.
Quantification of gene expression can be challenging.
RNA Sequencing (RNA-Seq)
Principle and Workflow
RNA-Seq is a next-generation sequencing (NGS) technique that determines the presence and quantity of RNA in a biological sample by sequencing cDNA derived from RNA. It provides comprehensive information about gene expression, alternative splicing, and allele-specific expression.
Isolation of RNA from cells.
Conversion of RNA to cDNA.
Fragmentation and preparation of cDNA for sequencing.
Sequencing of cDNA fragments.
Assembly and analysis of sequence reads into RNA transcripts.
Applications of RNA-Seq
Transcriptome profiling
Gene expression quantification
Alternative splicing detection
Disease diagnosis and biomarker discovery
Drug discovery and pharmacogenomics
Proteomics
Definition and Scope
Proteomics is the large-scale study of the proteome, the complete set of proteins produced by a cell, tissue, or organism. It analyzes protein structures, functions, interactions, and modifications. The proteome is dynamic and more complex than the genome due to alternative splicing and post-translational modifications (PTMs).
Categories of Proteomics:
Expression Proteomics: Studies protein expression levels.
Functional Proteomics: Identifies protein functions and interactions.
Structural Proteomics: Studies the 3D structure of proteins.
SDS-PAGE in Proteomics
Sodium Dodecyl Sulphate-Polyacrylamide Gel Electrophoresis (SDS-PAGE) is a technique used to separate proteins based on their size and charge. Proteins are denatured and given a uniform negative charge by SDS, then separated in a polyacrylamide gel under an electric field.
Determines protein purity and abundance.
Prepares proteins for further analysis (e.g., mass spectrometry, Western blotting).
Limitations: Not suitable for low-abundance proteins, cannot resolve proteins with similar molecular weights, and denatures proteins (disrupting interactions).
Mass Spectrometry
Mass spectrometry is used to identify proteins by measuring the mass-to-charge ratio of peptide fragments. It provides a protein 'fingerprint' that can be matched to databases for identification.
Affinity Capturing
Affinity capturing isolates specific proteins from complex mixtures using ligands (often antibodies) bound to beads. It is highly specific and preserves protein interactions, making it useful for studying protein complexes and post-translational modifications.
Applications: Protein purification, protein-protein interaction studies, PTM analysis, drug target identification.
Advantages: High specificity, preservation of interactions, compatibility with downstream analyses.
Disadvantages: Potential for false positives, harsh elution conditions, cost.
Protein Microarrays
Protein microarrays immobilize thousands of purified proteins on a solid surface for simultaneous analysis. They are used for diagnostics, proteomic research, drug discovery, and antibody characterization. Unlike DNA microarrays, they detect proteins and their interactions.
Workflow: Surface preparation, protein immobilization, blocking, sample incubation, detection, and imaging.
Summary Table: Microarrays vs. RNA-Seq
Feature | Microarrays | RNA-Seq |
|---|---|---|
Requirement for prior knowledge | Yes | No |
Detection of novel transcripts | No | Yes |
Quantification accuracy | Moderate | High |
Alternative splicing detection | Limited | Comprehensive |
Additional info: This guide integrates content from textbook chapters 13 and 20, focusing on RNA and protein technologies, and is suitable for exam preparation in a college-level genetics course.
