 Back
BackThe Genetic Code and Transcription: Deciphering the Flow of Genetic Information
Study Guide - Smart Notes

The Flow of Genetic Information
DNA, RNA, and Protein Synthesis
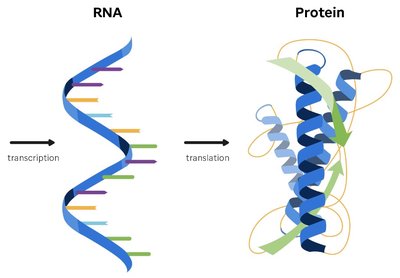
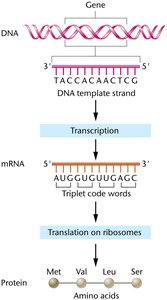
The central dogma of molecular genetics describes the flow of genetic information from DNA to RNA to protein. DNA stores genetic information, which is transcribed into messenger RNA (mRNA), an unstable intermediate that transfers information from the nucleus to the cytosol. Protein synthesis uses the genetic code to convert nucleotide sequence information into proteins.
The Genetic Code
Properties of the Genetic Code
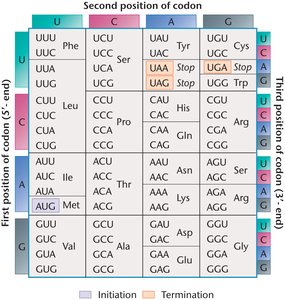
The genetic code is a set of rules used to translate the nucleotide sequence of mRNA into the amino acid sequence of proteins. It is composed of codons, each consisting of three ribonucleotide 'letters'. The code is unambiguous (each codon specifies only one amino acid or stop signal), degenerate (multiple codons can specify the same amino acid), continuous (codons are read sequentially), nonoverlapping (each nucleotide is part of only one codon), and nearly universal (with minor exceptions in mitochondria and some organisms).
Start codon: AUG (methionine)
Stop codons: UAA, UAG, UGA
Number of codons: 64 possible codons for 20 amino acids
Triplet Nature of the Code
Experimental evidence from virus frameshift mutations showed that the genetic code is read in triplets. Insertions or deletions of one or two nucleotides cause frameshift mutations, while changes of three nucleotides do not alter the reading frame.
Deciphering the Genetic Code
Producing Synthetic mRNAs
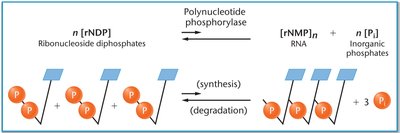
Polynucleotide phosphorylase is an enzyme used to create synthetic mRNAs in vitro. It can synthesize RNA without a template, and the probability of incorporating a specific ribonucleotide depends on its concentration.
Cell-Free Protein Synthesis
Proteins were synthesized in a cell-free system using E. coli extracts containing ribosomes, tRNAs, amino acids, and other molecules necessary for translation. Synthetic mRNAs and radiolabeled amino acids were added to track incorporation into proteins.
Homopolymer Experiments
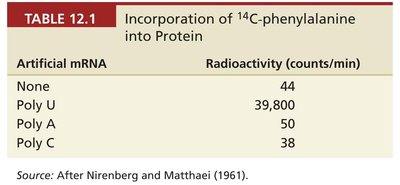
Homopolymers (polyU, polyA, polyC, polyG) were used to decipher the first codons. Each homopolymer encoded only one type of codon, resulting in the incorporation of a single amino acid type into the protein.
Artificial mRNA | Radioactivity (counts/min) |
|---|---|
None | 44 |
Poly U | 39,800 |
Poly A | 50 |
Poly C | 38 |

Mixed Heteropolymer Experiments
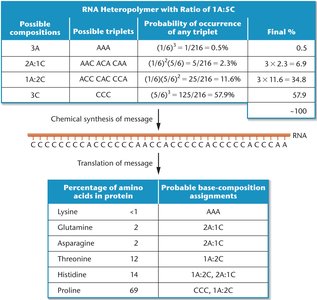
Mixed heteropolymers were created using two or more ribonucleotides. The frequency of a particular codon was predicted based on the relative concentrations of ribonucleotides. This allowed manipulation of codon composition and analysis of amino acid incorporation.
Possible compositions | Possible triplets | Probability of occurrence | Final % |
|---|---|---|---|
3A | AAA | (1/6)^3 = 0.5% | 0.5 |
2A:1C | AAC, ACA, CAA | (1/6)^2(5/6) = 2.3% | 6.9 |
1A:2C | ACC, CAC, CCA | (1/6)(5/6)^2 = 11.6% | 34.8 |
3C | CCC | (5/6)^3 = 57.9% | 57.9 |
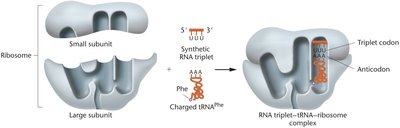
Triplet Binding Assay
The triplet binding assay involved synthesizing triplet mRNAs of known sequence, labeling a single amino acid, and mixing with ribosomes and charged tRNAs. The ribosome complex containing the triplet mRNA and appropriate tRNA was retained on a filter, allowing identification of codon assignments.
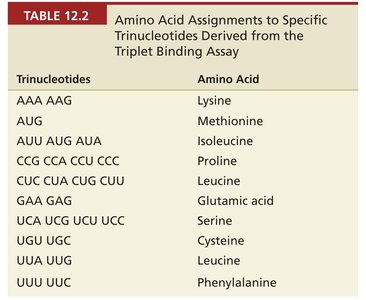
Trinucleotides | Amino Acid |
|---|---|
AAA, AAG | Lysine |
AUG | Methionine |
AUU, AUG, AUA | Isoleucine |
CCG, CCA, CCU, CCC | Proline |
CUC, CUA, CUG, CUU | Leucine |
GAA, GAG | Glutamic acid |
UCA, UCG, UCU, UCC | Serine |
UGU, UGC | Cysteine |
UUA, UUG | Leucine |
UUU, UUC | Phenylalanine |
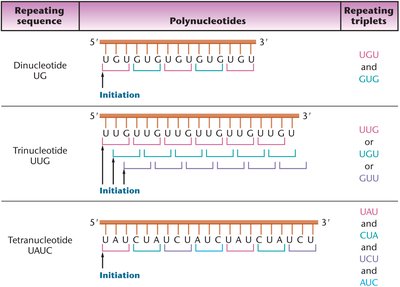
Repeating Copolymer Experiments
Khorana synthesized long mRNA molecules consisting of short repeating sequences (di-, tri-, and tetra-ribonucleotides). These repeating copolymers allowed further deciphering of codon assignments.
The Complete Genetic Code
Codon Assignments and Degeneracy
Most amino acids are represented by two to six codons, except tryptophan (Trp) and methionine (Met), which are specified by only one codon. AUG encodes methionine and initiates all eukaryotic polypeptide chains. UAA, UAG, and UGA are termination signals that do not encode an amino acid or bind a tRNA.

The Wobble Hypothesis
Flexibility in Codon-Anticodon Pairing
The wobble hypothesis states that the first two ribonucleotides of a codon are more critical than the third, which is less spatially constrained. This allows the anticodon of one tRNA to pair with more than one codon sequence, enabling a minimum of 30 tRNA species to accommodate the 61 amino acid codons.
Base at First Position (5′-end) of tRNA | Base at Third Position (3′-end) of mRNA |
|---|---|
A | U |
C | G |
G | C or U |
U | A or G |
I | A, U, or C |
Universality and Ordered Nature of the Genetic Code
Conservation and Colinearity
The genetic code is conserved in almost all organisms and viruses, with exceptions for a small number of codons in mitochondria, ciliate protozoans, and one bacterial species. The linear sequence of codons in a gene corresponds precisely with the linear amino acid sequence of the protein (colinearity). The 5’ end of a gene corresponds with the N-terminus of the protein.
Ordered Nature and Mutation Buffering
Chemically similar amino acids share the middle base in the different triplets encoding them. For example, U is present in the second position of codons specifying hydrophobic amino acids (Leu, Ile, Val), while C is present in the second position of codons specifying hydrophilic amino acids (Ser, Thr). This ordered code buffers the potential effect of mutation on protein function, as mutations in the first base often result in a change to an amino acid with similar chemical properties.

Summary Table: Deciphering the Genetic Code
Method | Main Purpose | Key Findings |
|---|---|---|
Homopolymer Synthesis | Assign codons to amino acids | UUU = phenylalanine, AAA = lysine, CCC = proline |
Mixed Heteropolymer Synthesis | Analyze codon composition | Codon frequency based on nucleotide ratios |
Triplet Binding Assay | Direct codon assignment | Unambiguous and degenerate code |
Repeating Copolymer Synthesis | Assign codons using repeating sequences | Further codon assignments |
