 Back
BackThe Genetic Code and Transcription: Deciphering the Flow of Genetic Information
Study Guide - Smart Notes

The Genetic Code and Transcription
Introduction to the Flow of Genetic Information
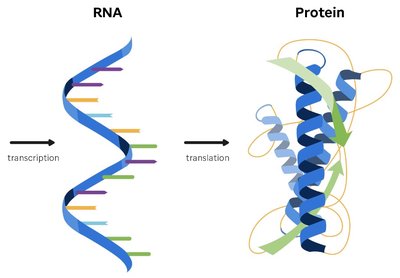
The central dogma of molecular genetics describes the flow of genetic information from DNA to RNA to protein. This process involves transcription, where DNA is used as a template to synthesize messenger RNA (mRNA), and translation, where mRNA is decoded to build proteins. The genetic code is the set of rules by which information encoded in mRNA is translated into proteins.
The Genetic Code: Properties and Structure
The genetic code is a set of instructions that determines how the nucleotide sequence of mRNA is converted into the amino acid sequence of proteins. It is composed of codons, which are sequences of three nucleotides that correspond to specific amino acids or stop signals during protein synthesis.
Linear Form: The code is read in a linear fashion along the mRNA.
Triplet Codons: Each codon consists of three ribonucleotides.
Unambiguous: Each codon specifies only one amino acid or stop signal.
Degenerate: Most amino acids are encoded by more than one codon.
Start and Stop Codons: There is one start codon (AUG) and three stop codons (UAA, UAG, UGA).
Continuous and Nonoverlapping: Codons are read one after another without overlap.
Nearly Universal: The code is conserved across almost all organisms, with minor exceptions.
Experimental Deciphering of the Genetic Code
Understanding the genetic code required a series of experiments using synthetic mRNAs and cell-free translation systems. These experiments demonstrated the triplet nature of the code and allowed scientists to assign specific codons to amino acids.
Frameshift Mutations: Insertions or deletions of one or two nucleotides cause frameshifts, altering the reading frame. Insertions or deletions of three nucleotides do not shift the frame, supporting the triplet nature of the code.
Synthetic mRNAs: Artificial mRNAs of known composition were used to direct protein synthesis in vitro.
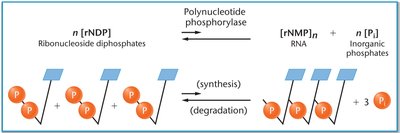
Polynucleotide Phosphorylase: This enzyme was used to synthesize RNA without a DNA template, allowing the creation of homopolymers and heteropolymers for code analysis.
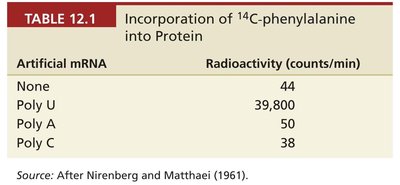
Homopolymers and the First Codons Deciphered
Homopolymers are synthetic mRNAs composed of only one type of nucleotide. When used in cell-free systems, they directed the incorporation of a single type of amino acid, revealing the codons for phenylalanine (UUU), lysine (AAA), and proline (CCC).

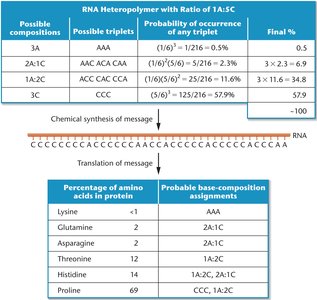
Mixed Heteropolymers and Codon Assignment
Mixed heteropolymers, composed of two or more types of nucleotides in defined ratios, allowed the statistical prediction of codon frequencies and the assignment of additional codons to amino acids.
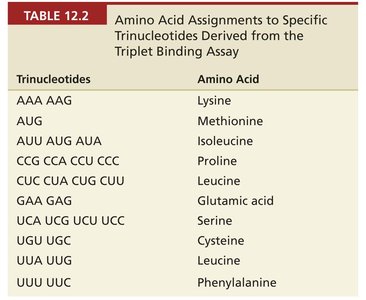
The Triplet Binding Assay
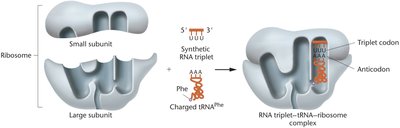
The triplet binding assay was a key technique for assigning codons to amino acids. In this assay, synthetic RNA triplets, charged tRNAs, and ribosomes were combined. If the tRNA matched the codon, the complex was retained on a filter, allowing identification of codon-amino acid pairs.
Repeating Copolymers and Further Deciphering
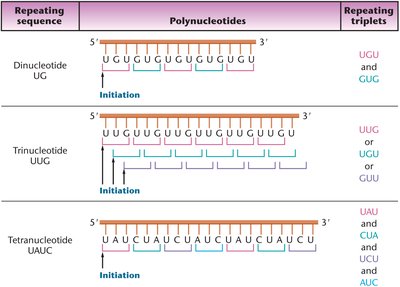
Repeating copolymers, synthesized with defined repeating sequences (di-, tri-, or tetra-nucleotides), produced mRNAs that encoded repeating sets of codons. This approach allowed the assignment of additional codons to amino acids.
The Complete Genetic Code
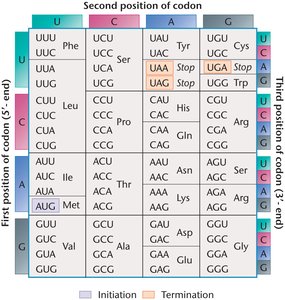
The complete genetic code table shows all 64 codons and their corresponding amino acids or stop signals. Most amino acids are specified by multiple codons, except tryptophan (Trp) and methionine (Met), which have only one codon each. The start codon (AUG) encodes methionine, and the stop codons (UAA, UAG, UGA) signal termination of translation.

The Wobble Hypothesis
The wobble hypothesis explains how a single tRNA can recognize multiple codons. The first two bases of the codon are most important for specifying the amino acid, while the third base is less constrained, allowing non-standard base pairing. This flexibility reduces the number of tRNAs needed to read all codons.
Wobble Position: The third base of the codon (3' end of mRNA) can pair with multiple bases in the tRNA anticodon.
Minimum tRNAs: At least 30 different tRNAs can accommodate the 61 sense codons.
Universality and Exceptions of the Genetic Code
The genetic code is nearly universal, conserved across almost all organisms and viruses. However, some exceptions exist, such as in mitochondria and certain protozoans, where specific codons may encode different amino acids or stop signals.
Colinearity: The linear sequence of codons in a gene corresponds directly to the linear sequence of amino acids in the protein.
5' to N-Terminus: The 5' end of the gene corresponds to the N-terminus of the protein.
Ordered Nature of the Genetic Code
The genetic code is ordered such that chemically similar amino acids often share the same middle base in their codons. This organization buffers the effects of mutations, as changes in the first base often result in amino acids with similar properties, minimizing the impact on protein function.
Example: Codons with U in the second position often encode hydrophobic amino acids (Leu, Ile, Val), while those with C in the second position encode hydrophilic amino acids (Ser, Thr).
