 Back
BackTranscription: DNA to RNA in Prokaryotes and Eukaryotes
Study Guide - Smart Notes

Genetic Material and the Central Dogma
Qualities of Genetic Material
The genetic material must possess several essential qualities to fulfill its biological role:
Accurate Replication: Ensures progeny cells and organisms inherit identical genetic information from the parent.
Stable Information Storage: Maintains the instructions for cellular form, structure, function, development, and behavior.
Information Transfer: Facilitates the flow of genetic information from DNA to RNA to protein.
DNA is the primary genetic material in most organisms, encoding the instructions for protein synthesis.
The Central Dogma of Molecular Biology
The central dogma describes the directional flow of genetic information:
DNA (sequence of nucleotides) is transcribed into RNA (sequence of nucleotides).
RNA is translated into protein (sequence of amino acids).
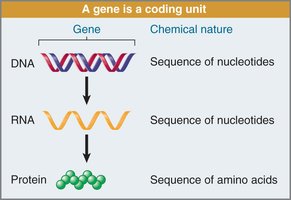
Crick (1956) proposed that genetic information moves from DNA through an RNA intermediate to produce proteins.
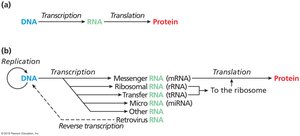
Today, it is recognized that multiple types of RNA are involved in protein synthesis, and information can sometimes flow backwards (e.g., reverse transcription in retroviruses).
RNA Structure and Types
RNA Nucleotides and Chemical Properties
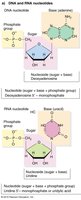
RNA is a nucleic acid composed of nucleotides, each containing:
Ribose sugar (with a 2' OH group)
Phosphate group (attached at 5' carbon)
Nitrogenous base (attached at 1' carbon)
RNA contains uracil (U) instead of thymine (T).
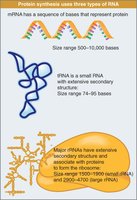
Major Types of RNA in Protein Synthesis
tRNA (transfer RNA): Small, stable, extensive secondary structure; delivers amino acids to ribosome.
rRNA (ribosomal RNA): Large, stable, forms ribosome core; catalyzes peptide bond formation.
mRNA (messenger RNA): Diverse, less stable; carries genetic code from DNA to ribosome.

Gene Structure and Transcription
Gene Organization on Chromosomes
Genes are located on chromosomes, which are long double-stranded DNA helices. Each gene occupies a defined position and is separated by noncoding DNA sequences.
Transcription: DNA to RNA
Transcription is the process by which an RNA transcript is synthesized from a DNA template. The enzyme responsible is RNA polymerase.
Transcription starts at a defined position (+1 base).
Transcription stops at a defined position.
Only one DNA strand (template strand) is used for RNA synthesis.
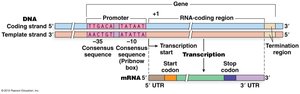
Transcript Structure
Transcripts may contain untranslated regions (UTRs) at their 5' and 3' ends, which do not code for protein.
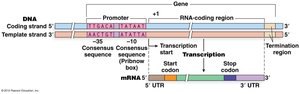
Promoter Regions and Initiation
The promoter region, located upstream of the gene, determines:
Where transcription starts
Which DNA strand is used
Direction of transcription
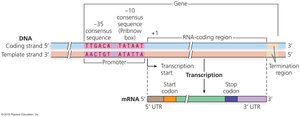
RNA Polymerase and Transcription Mechanism
RNA Polymerase Function
RNA polymerase synthesizes RNA in the 5' to 3' direction, using NTPs (ATP, GTP, CTP, UTP) as substrates. Unlike DNA polymerase, it does not require a primer.
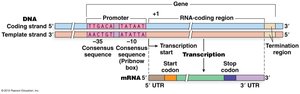
Base Pairing Rules
G pairs with C
U pairs with A (no T in RNA)
RNA polymerase forms a phosphodiester bond between the 3' OH of the growing RNA strand and the 5' phosphate of the incoming nucleotide.
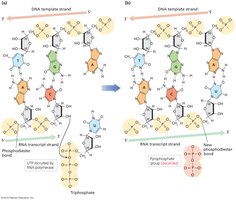
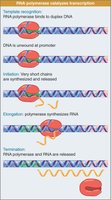
Stages of Transcription
Initiation: RNA polymerase binds to the promoter.
Elongation: RNA chain is extended.
Termination: Transcription stops and RNA is released.
Transcription in Prokaryotes
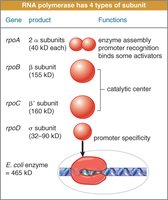
Bacterial RNA Polymerase and Sigma Factor
Bacterial RNA polymerase consists of multiple subunits. The sigma factor directs the core enzyme to the promoter region.
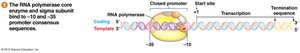
Bacterial Promoters and Consensus Sequences
Bacterial promoters have two consensus sequences:
-10 site: TATAAT
-35 site: TTGACA

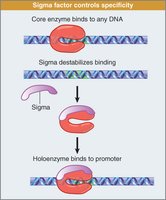
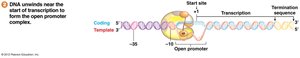
Transcription Initiation and Complex Formation
Transcription initiation involves formation of a closed complex (sigma factor binds promoter), followed by local unwinding to form an open complex.
Transcription Elongation and Termination
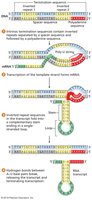
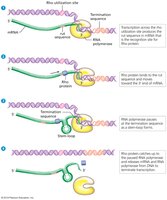
After initiation, sigma factor dissociates, and the core polymerase extends the mRNA. Termination occurs at specific sequences:
Intrinsic termination: RNA hairpin structure causes polymerase to fall off.
Rho-dependent termination: Rho protein binds RNA and interacts with polymerase to terminate transcription.
Transcription in Eukaryotes
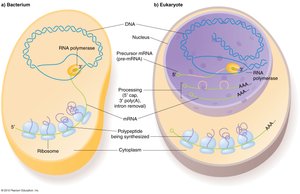
Key Differences from Prokaryotes
Protein-coding sequences are a minority of DNA.
Genes are larger, interrupted by introns, and more complex.
Transcription occurs in the nucleus; mRNA is exported to cytoplasm.
Three RNA polymerases (I, II, III) instead of one.
DNA is wrapped around nucleosomes to form chromatin.
Eukaryotic RNA Polymerases
RNA polymerase I: Synthesizes rRNA in nucleolus.
RNA polymerase II: Synthesizes mRNA.
RNA polymerase III: Synthesizes tRNA and other small RNAs.
Transcription Initiation in Eukaryotes
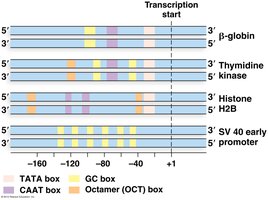
General Transcription Factors (GTFs) are required for accurate initiation. The TATA box (consensus sequence TATAA) is a common promoter element.
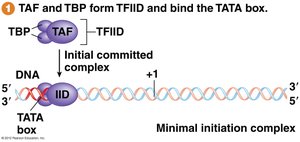
TFIID (contains TBP) binds TATA box.
TFIIB is recruited next.
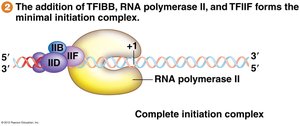
TFIIF and RNA pol II are recruited.
TFIIE and TFIIH join to form the preinitiation complex (PIC).
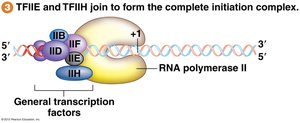
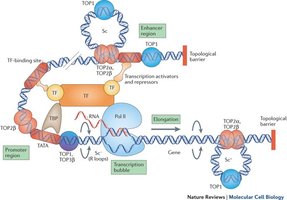
Transcription Elongation and Topoisomerase Function
RNA polymerase II moves forward, synthesizing mRNA. Topoisomerases relieve DNA supercoiling ahead of the transcription machinery.
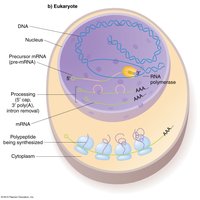
mRNA Processing in Eukaryotes
Modifications of Pre-mRNA
5' capping: Addition of 7-methyl-guanosine to the 5' end protects mRNA from degradation.
PolyA tail: Addition of a string of adenines to the 3' end facilitates export and stability.
Splicing: Removal of introns by the spliceosome; exons are joined to form mature mRNA.
Alternative Splicing
Some genes undergo alternative splicing, producing different proteins from the same gene by including or excluding specific exons.
Example: In one cell type, exons 1/2/3/4 are spliced together; in another, exons 1/2/3/5/6 are spliced together.
Additional info: Alternative splicing increases protein diversity and is regulated by cell type and developmental stage.
----------------------------------------
