 Back
BackTranslation and Protein Structure: From Genes to Functional Proteins
Study Guide - Smart Notes

Translation: Protein Synthesis in Cells
Polyribosomes (Polysomes)
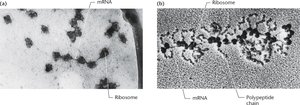
Polyribosomes, or polysomes, are complexes of a single mRNA molecule with multiple ribosomes attached, each actively translating the mRNA into a polypeptide chain. This arrangement allows for the simultaneous synthesis of several copies of a protein from one mRNA, greatly increasing the efficiency of protein production.
Structure: Multiple ribosomes spaced along the length of an mRNA strand.
Function: Enable rapid and efficient translation of proteins.
Example: Polyribosomes are visible in electron micrographs of cells actively synthesizing proteins, such as hemoglobin in reticulocytes.
X-Ray Diffraction Analysis of Ribosomes
X-ray diffraction analysis has enabled scientists to visualize the atomic structure of ribosomes, including the arrangement of their subunits and the binding sites for mRNA and tRNAs. These studies have revealed that ribosomes undergo conformational changes during translation and that their RNA components make direct contact with tRNA molecules.
Key Sites: A (aminoacyl), P (peptidyl), and E (exit) sites are critical for translation.
Functional Dynamics: Ribosome shape changes during different stages of translation.
Translation in Eukaryotes vs. Prokaryotes
Translation is more complex in eukaryotes than in prokaryotes due to differences in ribosome structure, mRNA processing, and cellular compartmentalization.
Ribosomes: Eukaryotic ribosomes are larger and more stable.
mRNA Processing: Eukaryotic mRNAs are capped at the 5′ end with 7-methylguanosine and have a poly-A tail at the 3′ end.
Location: Transcription occurs in the nucleus, while translation occurs in the cytoplasm in eukaryotes.
The Kozak Sequence
The Kozak sequence is a conserved nucleotide motif surrounding the start codon (AUG) in eukaryotic mRNAs. It enhances the efficiency of translation initiation by helping the ribosome recognize the correct start site.
Consensus Sequence: A/GNNG (where N is any nucleotide).
Function: Increases translation initiation efficiency.
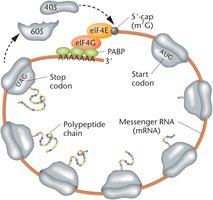
Closed-Loop Translation in Eukaryotes
In eukaryotes, the 5′ cap and 3′ poly-A tail of mRNA interact via protein factors to form a closed-loop structure. This configuration protects mRNA from degradation and facilitates efficient ribosome recycling, allowing multiple rounds of translation.
Key Proteins: eIF4E binds the cap, eIF4G acts as a scaffold, and PABPs bind the poly-A tail.
Advantages: Prevents translation of degraded mRNA and enhances translation efficiency.
One-Gene:One-Enzyme and One-Gene:One-Polypeptide Hypotheses
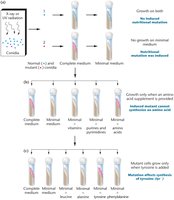
Beadle and Tatum's Experiments
Beadle and Tatum demonstrated that genes control the synthesis of enzymes by studying nutritional mutants in the bread mold Neurospora. Their work led to the formulation of the one-gene:one-enzyme hypothesis, which was later refined to the one-gene:one-polypeptide hypothesis as it became clear that not all proteins are enzymes and many proteins are composed of multiple polypeptide chains.
Key Findings: Mutations in single genes can disrupt specific enzymatic pathways.
Modern View: Each gene encodes a single polypeptide chain, which may function alone or as part of a larger protein complex.
Protein Structure and Folding
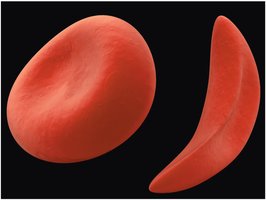
Sickle-Cell Anemia: A Case Study in Protein Mutation
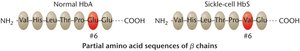
Sickle-cell anemia is a genetic disorder caused by a single amino acid substitution in the β-globin chain of hemoglobin. This mutation leads to abnormal hemoglobin structure, causing red blood cells to become sickle-shaped under low oxygen conditions.
Genetics: Recessive inheritance; homozygotes are affected, heterozygotes are carriers.
Biochemical Basis: Normal hemoglobin (HbA) and sickle-cell hemoglobin (HbS) differ by one amino acid at position 6 (glutamic acid to valine).
Phenotypic Effect: Altered hemoglobin causes red blood cells to deform, leading to various health complications.
Polypeptides and Proteins
Proteins are polymers of amino acids, assembled as polypeptide chains on ribosomes. The sequence and chemical properties of amino acids determine the protein's structure and function.
Polypeptides: Linear chains of amino acids; precursors to functional proteins.
Proteins: Folded polypeptides with specific three-dimensional structures.
Amino Acids: Structure and Classification
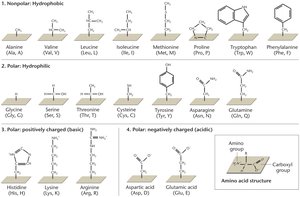
Amino acids are the building blocks of proteins. Each amino acid contains a central carbon atom bonded to an amino group, a carboxyl group, a hydrogen atom, and a variable R (side) group. The properties of the R group determine the amino acid's characteristics and its role in protein structure.
Classes of R Groups:
Nonpolar (hydrophobic)
Polar (hydrophilic)
Positively charged (basic)
Negatively charged (acidic)
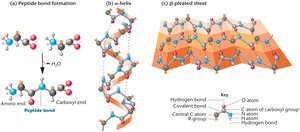
Peptide Bond Formation
A peptide bond forms between the amino group of one amino acid and the carboxyl group of another through a dehydration reaction. This linkage creates the backbone of polypeptide chains.
Dipeptide: Two amino acids joined by a peptide bond.
Tripeptide: Three amino acids joined together, and so on.
Levels of Protein Structure
Proteins have four levels of structural organization, each critical for their function:
Primary Structure: Linear sequence of amino acids.
Secondary Structure: Local folding into α-helices and β-pleated sheets, stabilized by hydrogen bonds.
Tertiary Structure: Overall three-dimensional shape of a single polypeptide chain.
Quaternary Structure: Assembly of multiple polypeptide chains into a functional protein complex.
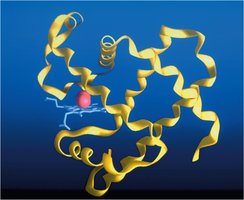
Protein Folding and Misfolding
Protein folding is a highly regulated process, often assisted by chaperone proteins. Misfolded proteins are tagged by ubiquitins and degraded by proteasomes. Improper folding can lead to diseases such as prion diseases (e.g., mad cow disease, Creutzfeldt–Jakob disease) and neurodegenerative disorders (e.g., Huntington's, Alzheimer's, Parkinson's).
Chaperones: Proteins that assist in proper folding.
Ubiquitin-Proteasome System: Degrades misfolded proteins to prevent cellular damage.
Disease Link: Protein misfolding and aggregation are associated with several human diseases.
Functional Diversity of Proteins
Major Protein Functions
Proteins are the most abundant macromolecules in cells and serve a wide variety of functions:
Respiratory Pigments: Hemoglobin and myoglobin transport oxygen.
Structural Proteins: Collagen and keratin provide support in tissues.
Contractile Proteins: Actin and myosin enable muscle contraction.
Microtubule Proteins: Tubulin forms the basis of spindle fibers in cell division.
Immunoglobulins: Function in the immune response.
Transport Proteins: Facilitate movement of molecules across membranes.
Hormones and Receptors: Regulate physiological processes.
Histones: Bind DNA and regulate gene expression in eukaryotes.
Transcription Factors: Control gene expression.
Enzymes
Enzymes are specialized proteins that catalyze biochemical reactions, increasing reaction rates and determining the metabolic capabilities of cells.
Catalytic Function: Lower activation energy for reactions.
Specificity: Each enzyme catalyzes a specific reaction or set of reactions.
Protein Domains
Protein domains are distinct functional and structural units within a protein, typically consisting of 50–300 amino acids. Each domain can impart a specific function, such as catalytic activity or DNA binding, contributing to the versatility of proteins.
Examples: Catalytic domains, DNA-binding domains.
Significance: Domains allow proteins to evolve new functions and interact with different molecules.
