 Back
BackTranslation and the Genetic Code: From DNA to Protein
Study Guide - Smart Notes

Translation and the Genetic Code
Overview of Gene Expression
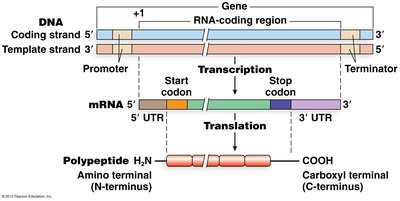
Gene expression is the process by which information encoded in DNA is used to direct the synthesis of proteins, the functional molecules of the cell. This process involves two main steps: transcription (DNA to RNA) and translation (RNA to protein).
Transcription: The DNA sequence of a gene is transcribed to produce a complementary messenger RNA (mRNA) molecule.
Translation: The mRNA sequence is decoded by the ribosome to synthesize a specific polypeptide (protein).
The Structure and Properties of Amino Acids
The 20 Common Amino Acids
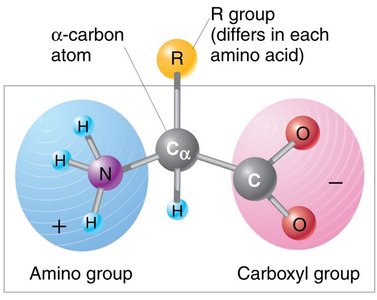
Proteins are polymers of amino acids. Each amino acid has a central carbon (α-carbon) bonded to an amino group, a carboxyl group, a hydrogen atom, and a variable R group (side chain) that determines its properties.
Amino group (–NH2): Basic group.
Carboxyl group (–COOH): Acidic group.
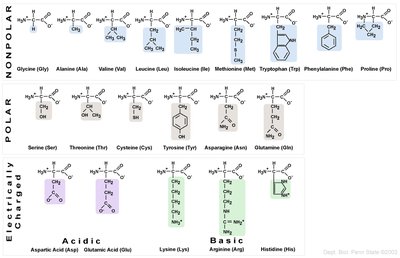
R group: Unique for each amino acid; determines chemical nature (nonpolar, polar, acidic, or basic).
Formation of Polypeptides
Peptide Bond Formation
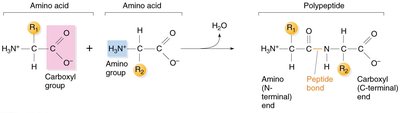
Amino acids are joined by peptide bonds, which are covalent bonds formed between the carboxyl group of one amino acid and the amino group of the next. This reaction releases water (a condensation reaction).
Polypeptide chain: Has an amino (N) terminus and a carboxyl (C) terminus.
Directionality: Proteins are synthesized from the N-terminus to the C-terminus.
The Flow of Genetic Information: The Central Dogma
DNA to RNA to Protein
The central dogma of molecular biology describes the flow of genetic information: DNA is transcribed into RNA, which is then translated into protein. The sequence of bases in DNA determines the sequence of amino acids in a protein.
Codon: A sequence of three nucleotides in mRNA that specifies a particular amino acid.
Reading frame: The way nucleotides are grouped into codons for translation.

The Genetic Code
Properties of the Genetic Code
The genetic code is the set of rules by which the sequence of bases in mRNA is translated into the sequence of amino acids in a protein.
Triplet code: Each codon consists of three nucleotides.
Non-overlapping: Codons are read one after another, without overlap.
Continuous: The code is read without punctuation from a fixed starting point.
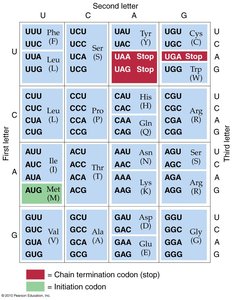
Start and stop codons: AUG (methionine) is the start codon; UAA, UAG, and UGA are stop codons.
Degenerate: Most amino acids are encoded by more than one codon.
Nearly universal: The code is conserved across almost all organisms.
Experimental Evidence for the Triplet Code
Crick and Brenner’s Experiment
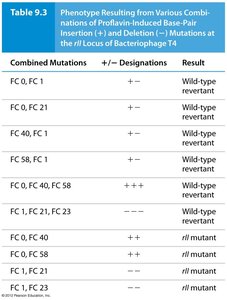
Francis Crick and Sydney Brenner used mutations in bacteriophage T4 to demonstrate that the genetic code is read in triplets. They showed that adding or deleting one or two bases disrupted the code, but adding or deleting three bases restored function, indicating a triplet code.
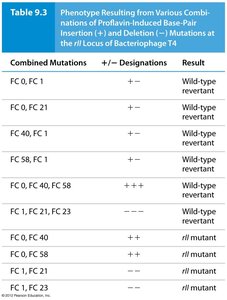
Proflavin-induced mutations: Caused insertions (+) or deletions (–) of single base pairs.
Revertants: Combinations of three insertions or deletions restored the reading frame and wild-type function.
Combined Mutations | + / – Designations | Result |
|---|---|---|
FC 0, FC 21 | + – | Wild-type revertant |
FC 40, FC 1 | + – | Wild-type revertant |
FC 58, FC 23 | + – | Wild-type revertant |
FC 0, FC 40, FC 58 | + + + | Wild-type revertant |
FC 1, FC 21, FC 23 | – – – | Wild-type revertant |
FC 0, FC 40 | + + | rII mutant |
FC 1, FC 21 | – – | rII mutant |
Deciphering the Genetic Code
Nirenberg and Khorana’s Experiments
Marshall Nirenberg and Har Gobind Khorana deciphered which codons specify which amino acids using synthetic RNAs and cell-free translation systems. Their work established the genetic code table used today.
Poly-U experiment: Poly-U RNA directed the synthesis of polyphenylalanine, showing UUU codes for phenylalanine.
Mixed copolymers: Used to determine codons for other amino acids.

Translation: From mRNA to Protein
The Role of tRNA
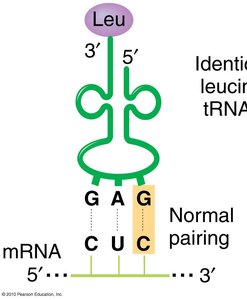
Transfer RNAs (tRNAs) are adapter molecules that match codons in mRNA with the correct amino acids during translation. Each tRNA has an anticodon that base-pairs with a codon in mRNA and an attached amino acid corresponding to that codon.
Anticodon: Three-nucleotide sequence in tRNA that pairs with the mRNA codon.
Charging: Aminoacyl-tRNA synthetases attach the correct amino acid to each tRNA.
Wobble position: The third base of the codon allows for some flexibility in pairing, enabling one tRNA to recognize multiple codons.
Translation Machinery and Process
The Ribosome and Translation Stages
The ribosome is the molecular machine that synthesizes proteins by reading the mRNA sequence and catalyzing peptide bond formation. Translation occurs in three stages: initiation, elongation, and termination.
Initiation: Ribosome assembles at the start codon (AUG) on the mRNA.
Elongation: Amino acids are added one by one to the growing polypeptide chain.
Termination: Translation ends when a stop codon is reached, and the completed protein is released.
Key sites on the ribosome: A (aminoacyl), P (peptidyl), and E (exit) sites for tRNA binding and movement.
Summary Table: Properties of the Genetic Code
Property | Description |
|---|---|
Triplet | Each codon is three nucleotides long |
Non-overlapping | Codons are read sequentially, without overlap |
Continuous | No punctuation; read 5' to 3' without gaps |
Start/Stop Codons | One start (AUG), three stop (UAA, UAG, UGA) |
Degenerate | Most amino acids have multiple codons |
Nearly Universal | Shared by almost all organisms |
Example: The codon CUC in mRNA encodes leucine. A charged Leu-tRNA with the GAG anticodon binds to CUC by normal base pairing.

Additional info: The genetic code's universality and degeneracy are crucial for the robustness and evolution of life. Mutations in the DNA sequence can lead to changes in protein sequence, which underlies genetic variation and evolution.
