 Back
BackTranslation of mRNA: The Genetic Code and Protein Synthesis
Study Guide - Smart Notes

Translation of mRNA
Overview of Translation
Translation is the process by which the genetic information encoded in messenger RNA (mRNA) is used to synthesize proteins. This process is fundamental to gene expression and involves decoding the nucleotide sequence of mRNA into a specific sequence of amino acids, forming a polypeptide chain.
Codons: Triplets of nucleotides in mRNA that specify amino acids or stop signals.
Ribosomes: Molecular machines that facilitate the translation process by bringing together mRNA and transfer RNA (tRNA).
tRNA: Adapter molecules that bring amino acids to the ribosome, matching codons in mRNA with the correct amino acid.
The Central Dogma and the Genetic Code
Relationship between the Genetic Code and Protein Synthesis
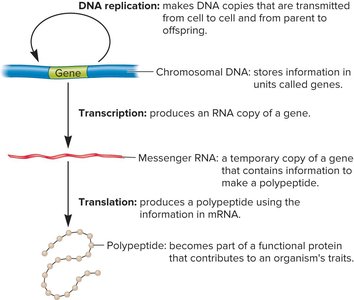
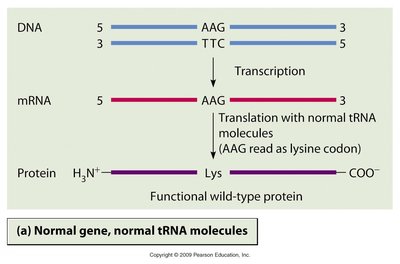
The central dogma of molecular genetics describes the flow of genetic information: DNA is transcribed into RNA, which is then translated into protein. The genetic code is the set of rules by which the nucleotide sequence of mRNA is translated into the amino acid sequence of proteins.
DNA replication: Copies DNA for inheritance.
Transcription: Synthesis of RNA from a DNA template.
Translation: Synthesis of proteins from mRNA templates.
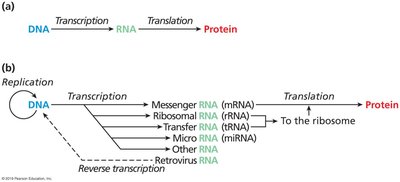
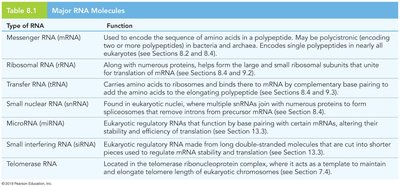
Types of RNA Involved in Translation
Several types of RNA play roles in translation, each with specific functions:
Type of RNA | Function |
|---|---|
Messenger RNA (mRNA) | Encodes the sequence of amino acids in a polypeptide. |
Ribosomal RNA (rRNA) | Forms the core of the ribosome's structure and catalyzes protein synthesis. |
Transfer RNA (tRNA) | Brings amino acids to the ribosome and matches them to the mRNA codons. |

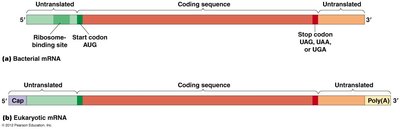
Structure of Genes and mRNA
Gene and mRNA Organization
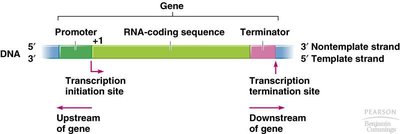
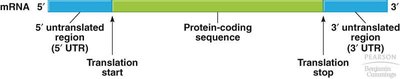
Genes contain coding and non-coding regions. The coding region is transcribed into mRNA, which is then translated into protein. mRNA contains untranslated regions (UTRs) at both ends, which play regulatory roles but are not translated into protein.
Promoter: Site where transcription begins.
RNA-coding sequence: Region that is transcribed and translated.
Terminator: Signals the end of transcription.
The Genetic Code
Codons: Sense and Nonsense
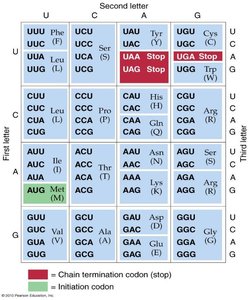
The genetic code consists of 64 codons, each made up of three nucleotides. Codons can be classified as sense (coding for amino acids) or nonsense (stop codons).
Start codon (AUG): Specifies the start of translation and codes for methionine (Met) in eukaryotes or N-formylmethionine (fMet) in bacteria.
Stop codons (UAA, UAG, UGA): Signal the termination of translation.
Characteristics of the Genetic Code
The genetic code has several important properties:
Triplet code: Each codon consists of three nucleotides.
Degeneracy: Most amino acids are encoded by more than one codon.
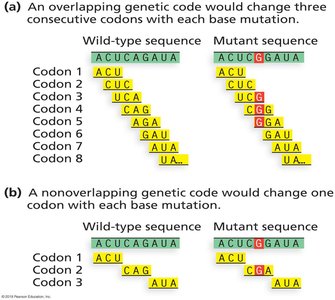
Non-overlapping: Codons are read one after another without overlap.
Universal (with exceptions): The code is nearly universal among organisms, with some exceptions in mitochondria and certain protozoa.
Unambiguous: Each codon specifies only one amino acid.
Continuous: Coding regions are read in a continuous sequence of codons (open reading frame, ORF).
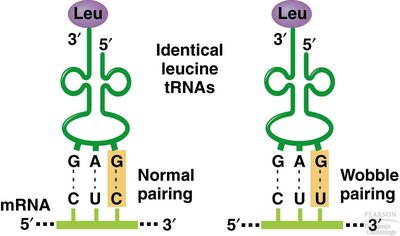
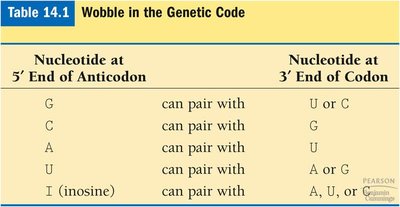
Wobble: Flexibility in base pairing at the third position of the codon allows some tRNAs to pair with more than one codon.
Nucleotide at 5' End of Anticodon | Nucleotide at 3' End of Codon |
|---|---|
G | U or C |
C | G |
A | U |
U | A or G |
I (inosine) | A, U, or C |
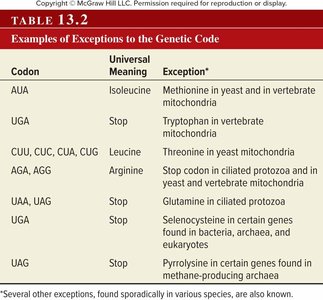
Exceptions to the Genetic Code
Although the genetic code is nearly universal, there are notable exceptions, especially in mitochondrial genomes and some protozoa.
Codon | Universal Meaning | Exception |
|---|---|---|
AUA | Isoleucine | Methionine in yeast and in vertebrate mitochondria |
UGA | Stop | Tryptophan in vertebrate mitochondria |
CUU, CUC, CUA, CUG | Leucine | Threonine in yeast mitochondria |
AGA, AGG | Arginine | Stop codon in ciliated protozoa and in yeast and vertebrate mitochondria |
UAA, UAG | Stop | Glutamine in ciliated protozoa |
UGA | Stop | Selenocysteine in certain genes found in bacteria, archaea, and eukaryotes |
UAG | Stop | Pyrrolysine in certain genes found in methane-producing archaea |
Amino Acids, tRNA, and Ribosome Structure
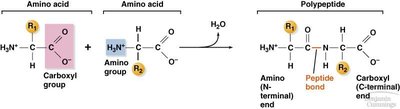
Amino Acids and Peptide Bond Formation
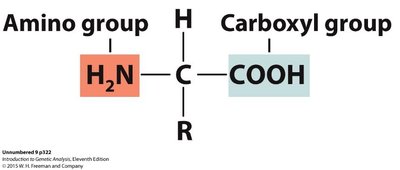
Proteins are polymers of amino acids linked by peptide bonds. Each amino acid has a central carbon atom bonded to an amino group, a carboxyl group, a hydrogen atom, and a variable R group.
Peptide bond: Formed between the carboxyl group of one amino acid and the amino group of the next.
Polypeptide: A chain of amino acids that folds into a functional protein.
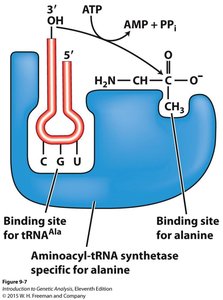
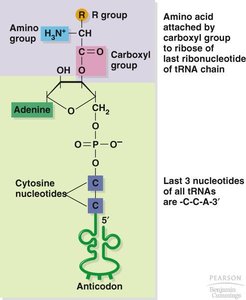
tRNA Structure and Function
tRNAs are adaptor molecules that match amino acids to codons in mRNA. Each tRNA has an anticodon that pairs with a codon and an acceptor stem for amino acid attachment. Aminoacyl-tRNA synthetases are enzymes that attach the correct amino acid to its corresponding tRNA.
Aminoacyl-tRNA synthetase: Enzyme that charges tRNA with its specific amino acid.
Anticodon: Three-nucleotide sequence in tRNA complementary to the mRNA codon.
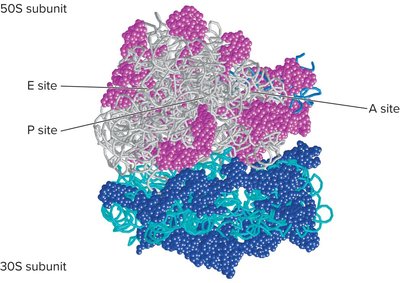
Ribosome Structure and Assembly
Ribosomes are composed of rRNA and proteins, forming two subunits (large and small). The ribosome has three sites for tRNA binding: A (aminoacyl), P (peptidyl), and E (exit).
A site: Binds incoming aminoacyl-tRNA.
P site: Holds the tRNA with the growing polypeptide chain.
E site: Site where uncharged tRNA exits the ribosome.
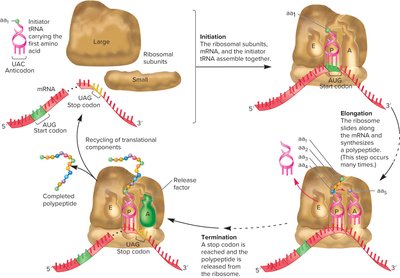
Stages of Translation
Initiation in Bacteria
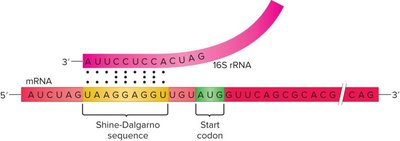
Translation initiation in bacteria involves the assembly of the ribosome on the mRNA at the start codon, guided by the Shine-Dalgarno sequence.
30S subunit binds to mRNA at the Shine-Dalgarno sequence.
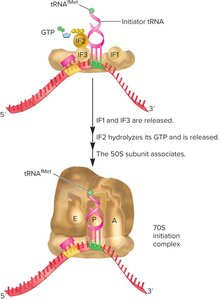
Initiator tRNAfMet binds to the start codon (AUG).
50S subunit joins to form the complete ribosome.
Elongation in Bacteria
During elongation, amino acids are added one by one to the growing polypeptide chain.
Incoming aminoacyl-tRNA binds to the A site.
Peptidyl transferase forms a peptide bond between the polypeptide and the new amino acid.
Ribosome translocates, moving the tRNA and mRNA through the sites.
Termination in Bacteria
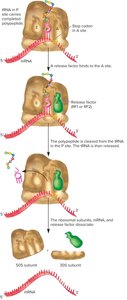
Termination occurs when a stop codon is reached. Release factors bind to the A site, causing the release of the polypeptide and dissociation of the translation complex.
Stop codon enters the A site.
Release factor binds, triggering release of the polypeptide.
Ribosomal subunits dissociate.
Translation in Eukaryotes
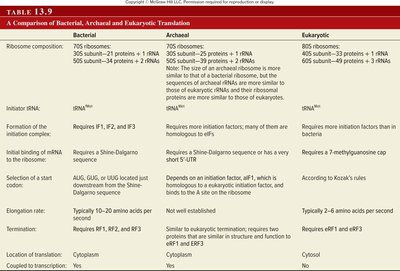
Translation in eukaryotes is similar to that in bacteria but has key differences, such as the use of a 5' cap for ribosome binding and the scanning mechanism for start codon recognition (Kozak's rules).
40S subunit binds to initiator tRNAMet and the 5' cap of mRNA.
Scans for AUG start codon in a favorable context (Kozak's rules).
60S subunit joins to form the complete ribosome.
Nonsense Suppressor Mutations and Nonsense-Mediated Decay
Nonsense Suppressor Mutations
Nonsense suppressor mutations allow the insertion of an amino acid at a stop codon, enabling translation to continue and potentially restoring protein function.
Suppressor tRNAs recognize stop codons and insert an amino acid.
Can partially compensate for nonsense mutations in the coding sequence.
Nonsense-Mediated Decay (NMD)
NMD is a cellular quality control mechanism that degrades mRNAs containing premature termination codons (PTCs), preventing the production of truncated, potentially harmful proteins.
Exon junction complexes (EJCs) mark exon-exon boundaries after splicing.
Normally, ribosomes remove EJCs during translation.
PTCs prevent removal of downstream EJCs, triggering mRNA degradation.
Diseases such as cystic fibrosis and Duchenne muscular dystrophy can result from PTCs. Therapeutic strategies aim to promote readthrough of PTCs to restore full-length protein production.
Posttranslational Modifications
Types of Posttranslational Modifications
After translation, proteins often undergo modifications that are essential for their function, stability, and localization.
Protein folding: Assisted by chaperones to achieve correct 3D structure.
Terminal modifications: Removal of formyl group from fMet in bacteria, acetylation of N-terminus in eukaryotes.
Modification of amino acids: Phosphorylation, acetylation, methylation, carboxylation, glycosylation.
Proteolytic processing: Cleavage of precursor proteins into active forms.
Addition of prosthetic groups: Attachment of vitamins or metals.
Protein trafficking: Directs proteins to correct cellular compartments.
Ubiquitination: Marks proteins for degradation by the proteasome.
