 Back
BackClassification and Identification of Microorganisms: Taxonomy, Phylogeny, and Laboratory Methods
Study Guide - Smart Notes

Classification of Microorganisms
Introduction to Taxonomy and Systematics
Taxonomy is the science of classifying organisms, providing a universal framework for naming and grouping organisms based on shared characteristics. Systematics, or phylogeny, is the study of the evolutionary history and relationships among organisms, often visualized as phylogenetic trees.
Taxonomy: Involves identification, classification, and nomenclature of organisms.
Systematics/Phylogeny: Focuses on evolutionary relationships, often using genetic data.
The Three-Domain System
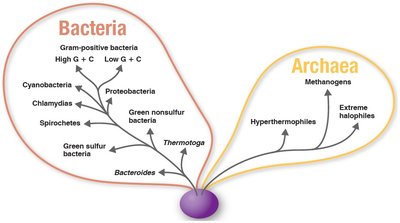
The three-domain system, developed by Carl Woese in 1978, classifies all life into three domains based on rRNA nucleotide sequences: Bacteria, Archaea, and Eukarya. This system reflects fundamental differences in cell structure and genetics.
Bacteria: Prokaryotic, includes common bacteria.
Archaea: Prokaryotic, includes extremophiles such as methanogens, extreme halophiles, and hyperthermophiles.
Eukarya: Eukaryotic, includes animals, plants, fungi, and protists.
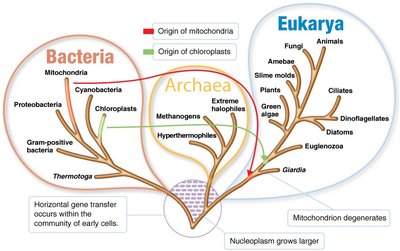
Key Concepts: All organisms evolved from a common ancestor. DNA sequences are conserved and used to determine evolutionary relationships. Eukaryotes originated from infoldings of prokaryotic membranes and endosymbiotic events.
Comparative Characteristics of Domains
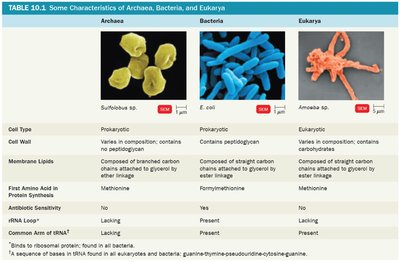
Archaea, Bacteria, and Eukarya differ in cell structure, membrane composition, and genetic machinery. These differences are summarized in the following table:
Characteristic | Archaea | Bacteria | Eukarya |
|---|---|---|---|
Cell Type | Prokaryotic | Prokaryotic | Eukaryotic |
Cell Wall | Varies; no peptidoglycan | Contains peptidoglycan | Varies; no peptidoglycan |
Membrane Lipids | Branched carbon chains attached to glycerol by ether linkage | Straight carbon chains attached to glycerol by ester linkage | Straight carbon chains attached to glycerol by ester linkage |
First Amino Acid in Protein Synthesis | Methionine | Formylmethionine | Methionine |
Antibiotic Sensitivity | No | Yes | No |
rRNA Loop | Lacking | Present | Lacking |
Common Arm of tRNA | Lacking | Present | Present |
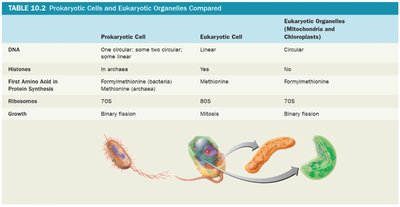
Prokaryotic Cells and Eukaryotic Organelles Compared
Prokaryotic cells, eukaryotic cells, and eukaryotic organelles (mitochondria and chloroplasts) share some similarities, supporting the endosymbiotic theory of organelle origin.
Feature | Prokaryotic Cell | Eukaryotic Cell | Eukaryotic Organelles |
|---|---|---|---|
DNA | One circular; some two circular; some linear | Linear | Circular |
Histones | In archaea | Yes | No |
First Amino Acid in Protein Synthesis | Formylmethionine (bacteria), Methionine (archaea) | Methionine | Formylmethionine |
Ribosomes | 70S | 80S | 70S |
Growth | Binary fission | Mitosis | Binary fission |
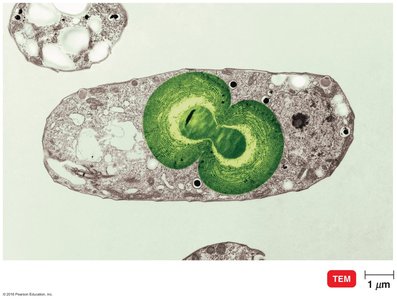
Endosymbiotic Theory and Origin of Eukaryotes
The endosymbiotic theory proposes that eukaryotic organelles such as mitochondria and chloroplasts originated from free-living bacteria that were engulfed by ancestral eukaryotic cells. This is supported by similarities in DNA, ribosomes, and reproduction between these organelles and prokaryotes.
Phylogenetic Relationships and Evolution
Phylogenetic Trees and Fossil Evidence
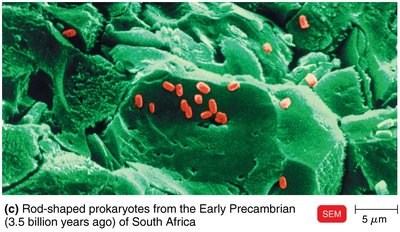
Phylogenetic trees group organisms based on common properties, such as fossil records and genetic sequences. Mutations in genomes serve as molecular clocks to estimate evolutionary divergence.
Fossilized prokaryotes provide evidence for early life forms and evolutionary history.
Each species retains some ancestral characteristics.


Scientific Nomenclature and Taxonomic Hierarchy
Binomial Nomenclature
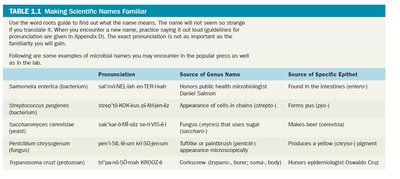
Scientific names use binomial nomenclature, consisting of a genus and specific epithet (species). This system ensures consistency and accuracy in naming organisms worldwide.
Genus: Capitalized, italicized or underlined.
Species: Lowercase, italicized or underlined.
Example: Escherichia coli
Microbe | Pronunciation | Source of Genus Name | Source of Specific Epithet |
|---|---|---|---|
Salmonella enterica | sal-mo-NEL-la en-ter-IK-a | Honors Daniel Salmon | Found in the intestines (enteron) |
Streptococcus pyogenes | strep-to-KOK-us pi-OJ-e-neez | Appearance of cells in chains (strepto-) | Forms pus (pyo-) |
Saccharomyces cerevisiae | sac-kar-o-MY-seez ser-e-VIS-e-a | Fungus (myces) that uses sugar (saccharo-) | Makes beer (cerevisia) |
Pseudomonas aeruginosa | su-do-MO-nas a-roo-jin-O-sa | False unit (pseudo-) | Produces a blue-green pigment |
Trypanosoma cruzi | tri-pan-o-SO-ma KROO-zi | Corkscrew (trypano-) | Honors Oswaldo Cruz |
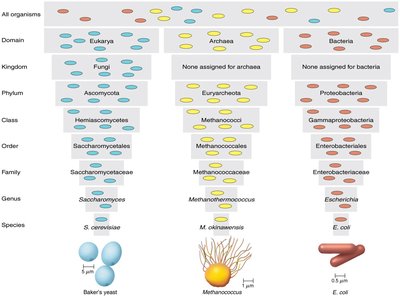
Taxonomic Hierarchy
The taxonomic hierarchy is a series of ranked groups developed by Carolus Linnaeus. It organizes organisms from broad to specific categories: Domain, Kingdom, Phylum, Class, Order, Family, Genus, Species.
Eukaryotic species: Group of closely related organisms that breed among themselves.
Prokaryotic species: Population of cells with similar characteristics; includes concepts of culture, clone, and strain.
Classification of Prokaryotes, Eukaryotes, and Viruses
Prokaryotic Classification
Prokaryotic species are defined by genetic and phenotypic similarity. Strains within a species may differ genetically. Phylogenetic relationships are often depicted in trees based on genetic data.
Eukaryotic Classification
Eukaryotes are classified into kingdoms based on cell structure, nutrition, and genetic data:
Protista: Diverse, grouped by rRNA clades; autotrophic and heterotrophic.
Fungi: Chemoheterotrophic, chitin cell walls, develop from spores or hyphae.
Plantae: Multicellular, cellulose cell walls, photosynthetic.
Animalia: Multicellular, no cell walls, chemoheterotrophic.
Classification of Viruses
Viruses are not classified within the three domains because they are acellular and require host cells for replication. Viral species are defined by shared characteristics and ecological niche.
Methods of Classifying and Identifying Microorganisms
Overview of Classification and Identification
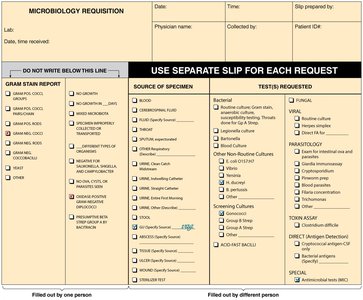
Classification involves grouping organisms based on similarities, while identification matches unknown organisms to known groups. Bergey's Manual and Approved Lists of Bacterial Names are key references. Clinical labs use requisition forms and transport media for specimen handling.
Phenotypic Methods
Morphological characteristics: Useful for eukaryotes, limited for prokaryotes.
Differential staining: Gram and acid-fast stains distinguish major groups but are not useful for wall-less bacteria.
Biochemical tests: Detect presence of specific enzymes or metabolic pathways.
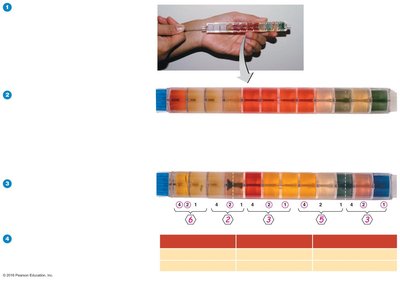
Rapid Identification Methods
Rapid identification systems perform multiple biochemical tests simultaneously, generating a code number for database comparison. Automated systems use mass spectrophotometry to analyze cellular proteins.
Serological Methods
Serology studies immune responses in serum. Microorganisms are antigenic and stimulate antibody production. Serological tests include slide agglutination, ELISA, and Western blotting.
Slide agglutination: Detects antigen-antibody reactions.
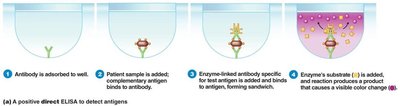
ELISA: Uses enzyme-linked antibodies to detect specific antigens or antibodies.
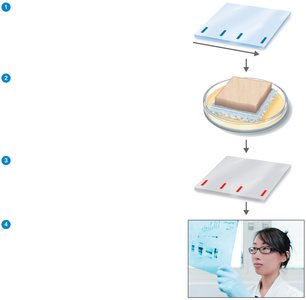
Western blotting: Identifies specific proteins or antibodies, used for confirming infections like HIV and Lyme disease.
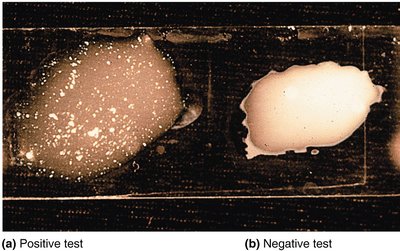

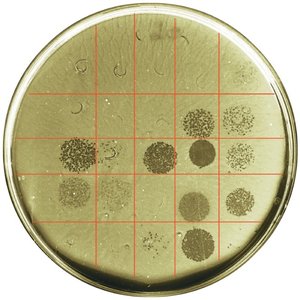
Phage Typing
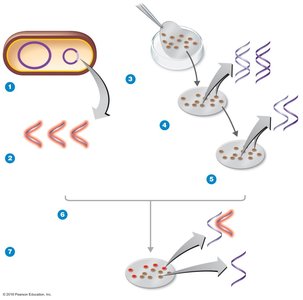
Phage typing determines bacterial susceptibility to specific bacteriophages. Plaques (clearings) on a bacterial lawn indicate lysis by phages, helping differentiate bacterial strains.
Molecular Methods
DNA base composition: Compares G+C content; closely related organisms have similar base ratios.
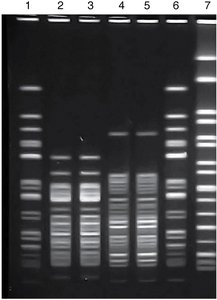
DNA fingerprinting: Uses restriction enzyme digestion and electrophoresis to compare genetic patterns.
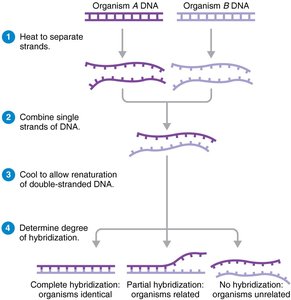
Nucleic acid hybridization: Measures the ability of DNA from different organisms to hybridize; >70% hybridization indicates same species.
Nucleic Acid Amplification Tests (NAATs): Use PCR to amplify DNA from unculturable organisms.
Southern blotting: Uses DNA probes to identify specific sequences in unknown samples.
DNA chips (microarrays): Detect pathogens by hybridization with fluorescently labeled probes.
Ribotyping: Uses rRNA sequencing for identification.
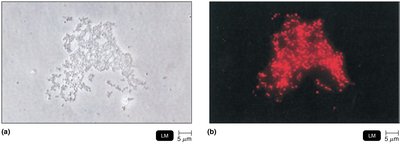
Fluorescent in situ hybridization (FISH): Uses fluorescent probes to stain and identify microorganisms in situ.

Integrating Classification Methods
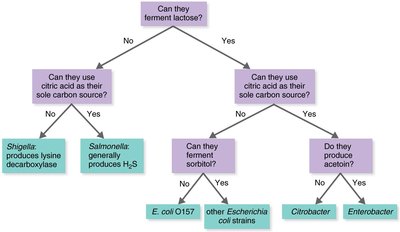
Dichotomous keys use a series of questions to identify organisms. Cladograms are diagrams that show evolutionary relationships, often based on rRNA sequences.
Additional info: This guide integrates foundational concepts in microbial taxonomy, phylogeny, and laboratory identification, providing a comprehensive overview for microbiology students. It includes definitions, examples, and visual aids to reinforce learning.
