 Back
BackClassification of Microorganisms: Taxonomy, Phylogeny, and Identification Methods
Study Guide - Smart Notes

Classification of Microorganisms
Introduction to Taxonomy and Phylogeny
Taxonomy is the science of classifying organisms, providing a universal framework for naming and grouping organisms based on similarities. Systematics, or phylogeny, studies the evolutionary history and relationships among organisms, often visualized as phylogenetic trees. These disciplines are foundational for understanding microbial diversity and evolution.
Taxonomy: Involves classification, nomenclature, and identification of organisms.
Systematics (Phylogeny): Focuses on evolutionary relationships, often using genetic data.
Historical Milestones: Linnaeus (1735) established kingdoms Plantae and Animalia; later, bacteria and fungi were classified, and the five-kingdom system was proposed by Whittaker (1969).
The Three-Domain System
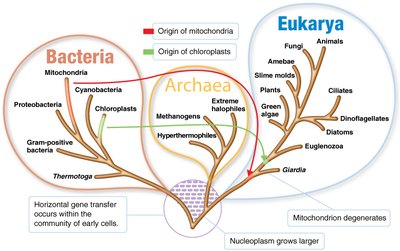
The three-domain system, developed by Carl Woese in 1978, is based on differences in ribosomal RNA (rRNA) sequences. It divides all life into three domains: Bacteria, Archaea, and Eukarya.
Bacteria: True bacteria, including most known prokaryotes.
Archaea: Prokaryotes distinct from bacteria, often found in extreme environments (e.g., methanogens, extreme halophiles, hyperthermophiles).
Eukarya: Organisms with eukaryotic cells, including animals, plants, fungi, and protists.
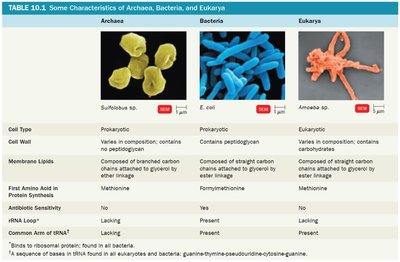
Characteristics of Archaea, Bacteria, and Eukarya
Each domain has unique cellular and molecular features. The table below summarizes key differences among Archaea, Bacteria, and Eukarya.
Feature | Archaea | Bacteria | Eukarya |
|---|---|---|---|
Cell Type | Prokaryotic | Prokaryotic | Eukaryotic |
Cell Wall | Varies; no peptidoglycan | Peptidoglycan | Varies; contains carbohydrates |
Membrane Lipids | Branched carbon chains | Straight carbon chains | Straight carbon chains |
First Amino Acid in Protein Synthesis | Methionine | Formylmethionine | Methionine |
Antibiotic Sensitivity | No | Yes | No |
rRNA Loop | Lacking | Present | Lacking |
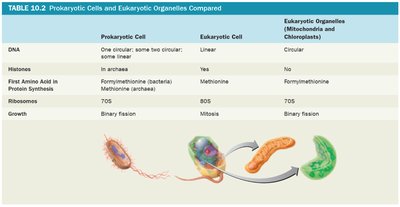
Prokaryotic Cells and Eukaryotic Organelles Compared
Prokaryotic cells differ from eukaryotic cells in several fundamental ways, including DNA structure, presence of histones, and ribosome type. Some eukaryotic organelles (mitochondria and chloroplasts) share similarities with prokaryotes, supporting the endosymbiotic theory.
Feature | Prokaryotic Cell | Eukaryotic Cell | Eukaryotic Organelles |
|---|---|---|---|
DNA | Circular | Linear | Circular |
Histones | No (except in Archaea) | Yes | No |
Ribosomes | 70S | 80S | 70S |
Growth | Binary fission | Mitosis | Binary fission |
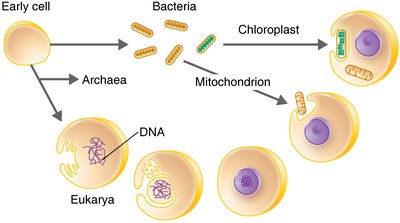
Endosymbiotic Theory
The endosymbiotic theory explains the origin of eukaryotic organelles such as mitochondria and chloroplasts. According to this theory, these organelles originated from free-living bacteria that were engulfed by ancestral eukaryotic cells.
Mitochondria and chloroplasts contain their own DNA and ribosomes, similar to bacteria.
They replicate independently within the cell by binary fission.
Taxonomic Hierarchy and Nomenclature
Scientific Nomenclature
Binomial nomenclature is the standardized system for naming organisms, consisting of a genus and a specific epithet (species). This system ensures consistency and avoids confusion caused by common names.
Genus: Capitalized and italicized (e.g., Escherichia).
Species: Lowercase and italicized (e.g., coli).
Names often reflect characteristics, discoverers, or habitats.
Taxonomic Hierarchy
The taxonomic hierarchy is a series of ranked categories used to classify organisms. The main ranks are: Domain, Kingdom, Phylum, Class, Order, Family, Genus, and Species (mnemonic: "Dear King Philip Came Over For Good Soup").
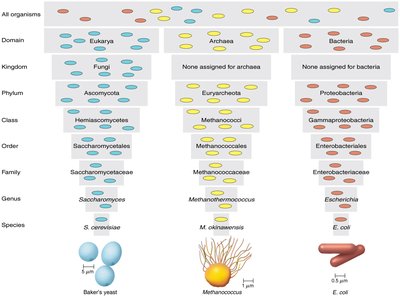
Classification of Prokaryotes and Eukaryotes
Prokaryotic species are defined as populations of cells with similar characteristics. Eukaryotic species are groups of organisms that can breed among themselves. Strains are genetically distinct subgroups within a species.
Culture: Bacteria grown in laboratory media.
Clone: Population derived from a single parent cell.
Strain: Genetically different cells within a clone.
Classification of Viruses
Viruses are not classified within the three domains because they are acellular and require host cells for replication. Viral species are defined as populations of viruses with similar characteristics occupying a particular ecological niche.
Methods of Classifying and Identifying Microorganisms
Overview of Classification and Identification
Classification involves grouping organisms based on shared characteristics, while identification matches unknown organisms to known groups. Bergey’s Manual is a key reference for bacterial classification.
Laboratory Identification Methods
Morphological characteristics: Useful for eukaryotes but limited for bacteria.
Differential staining: Gram and acid-fast stains distinguish major groups of bacteria.
Biochemical tests: Detect metabolic capabilities, such as enzyme presence.
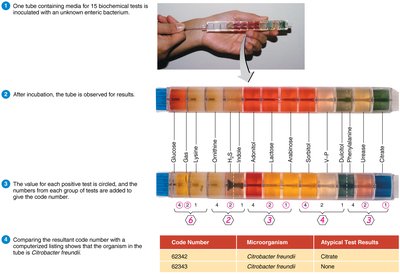
Serological Methods
Serology studies immune responses in serum. Microorganisms are antigenic and can be identified by their reaction with specific antibodies.
Slide agglutination test: Bacteria clump when mixed with specific antibodies.
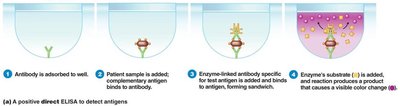
ELISA (Enzyme-Linked Immunosorbent Assay): Detects antigens or antibodies using enzyme-linked reactions.
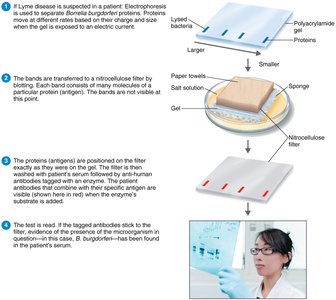
Western blotting: Identifies specific proteins or antibodies in a sample.
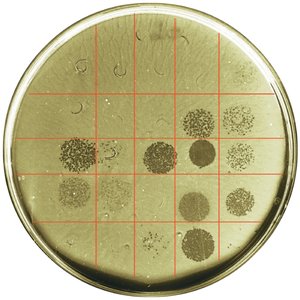
Phage Typing
Phage typing determines bacterial susceptibility to specific bacteriophages. Clear zones (plaques) on a bacterial lawn indicate lysis by phages.
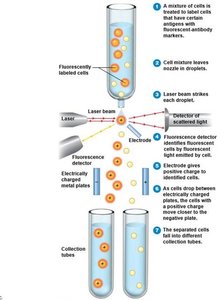
Flow Cytometry
Flow cytometry distinguishes cells based on electrical conductivity or fluorescence. It is useful for rapid identification and sorting of microorganisms.
Genetic and Molecular Methods
DNA base composition: The percentage of guanine and cytosine (G+C) in DNA is characteristic for each species.
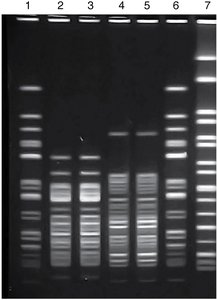
DNA fingerprinting: Electrophoresis of restriction enzyme digests reveals genetic similarities and differences.
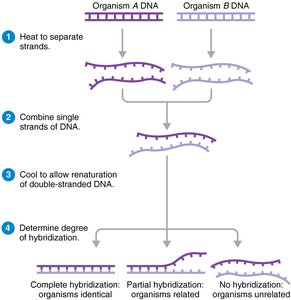
Nucleic acid hybridization: Measures the ability of DNA from different organisms to hybridize, indicating relatedness. Hybridization above 70% suggests the same species.
NAATs (Nucleic Acid Amplification Tests): Use PCR to amplify DNA for identification.
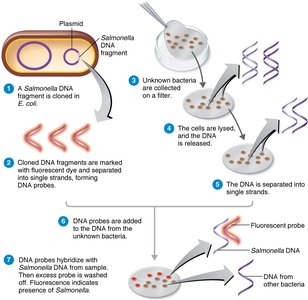
Southern blotting: Uses DNA probes to detect specific sequences in unknown samples.
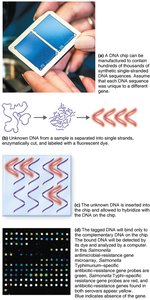
DNA chips (microarrays): Detect pathogens by hybridization with DNA probes on a chip.
Ribotyping: Uses rRNA sequencing for identification.
FISH (Fluorescent in situ hybridization): Uses fluorescent probes to detect and localize specific DNA or RNA sequences in cells.

Tools for Classification: Dichotomous Keys and Cladograms
Dichotomous Keys
Dichotomous keys are tools for identifying organisms based on a series of choices that lead to the correct name. Each step presents two alternatives, guiding the user to the next step or the final identification.
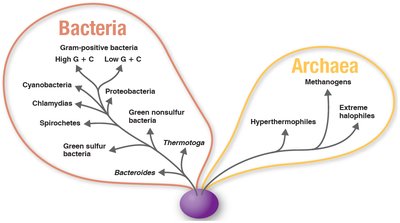
Cladograms
Cladograms are branching diagrams that depict evolutionary relationships among organisms, often based on genetic data such as rRNA sequences. They help visualize the evolutionary pathways and relatedness of different groups.
Summary Table: Key Methods for Microbial Classification and Identification
Method | Principle | Application |
|---|---|---|
Morphological | Shape, structure | Eukaryotes, some bacteria |
Differential Staining | Cell wall properties | Gram, acid-fast bacteria |
Biochemical Tests | Metabolic reactions | Bacterial identification |
Serology | Antigen-antibody reactions | Species/strain identification |
Phage Typing | Phage susceptibility | Bacterial strain typing |
Genetic Methods | DNA/RNA analysis | Phylogeny, species ID |
