 Back
BackMolecular Regulation in Microbiology: Mechanisms and Applications
Study Guide - Smart Notes

Molecular Regulation
Overview of Molecular Regulation
Molecular regulation refers to the complex systems cells use to control their internal processes, ensuring homeostasis and adaptability to environmental changes. This regulation involves the coordinated activity of genes, proteins, and other molecules to ensure that essential biological functions occur at the correct time, location, and quantity.
Central Dogma & Gene Regulation
Transcription Factors: Activators and Repressors
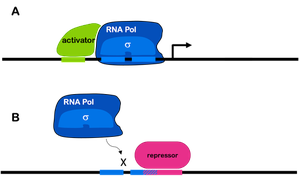
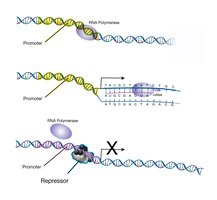
Transcription factors are proteins that act as molecular switches, binding to specific DNA sequences to regulate gene expression by either promoting or inhibiting the initiation of transcription by RNA polymerase.
Activators: Increase gene transcription by recruiting RNA polymerase to the promoter or by remodeling chromatin to make DNA more accessible.
Repressors: Decrease or block gene transcription by preventing RNA polymerase binding, quenching activators, or recruiting enzymes that condense chromatin.
Gene regulation is often a balance between multiple activators and repressors, functioning more like a dimmer switch (rheostat) than a simple on/off mechanism.
Chromatin Remodeling
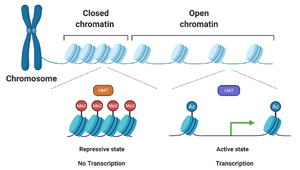
Chromatin structure plays a critical role in gene regulation, especially in eukaryotes. Chromatin can exist in a closed (repressive) or open (active) state, influencing the accessibility of DNA to transcription machinery.
Histone modifications: Enzymes such as histone acetyltransferases (HATs) and histone methyltransferases (HMTs) add or remove chemical groups, altering chromatin structure.
Open chromatin: Associated with active transcription.
Closed chromatin: Associated with gene silencing.

The "Rheostat" Effect in Gene Regulation
Gene expression is often regulated in a graded manner, with the level of mRNA produced depending on the combined influence of multiple activators and repressors. This allows cells to fine-tune gene expression in response to environmental and internal signals.

Prokaryotic Gene Regulation
Alternative Sigma Factors
In bacteria, sigma (σ) factors are essential for directing RNA polymerase to specific promoter sequences. Alternative sigma factors allow bacteria to rapidly switch gene expression programs in response to environmental changes.
Housekeeping sigma factor (σ70): Controls genes for routine growth.
Alternative sigma factors: Activate stress response genes (e.g., σ32 for heat shock, σ38 for starvation).
Regulons: Groups of genes controlled by a specific sigma factor.
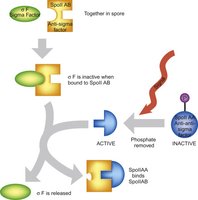
Anti-Sigma and Anti-Anti-Sigma Factors
Anti-sigma factors are proteins that bind and inhibit sigma factors, preventing them from associating with RNA polymerase. Anti-anti-sigma factors can bind to anti-sigma factors, releasing the sigma factor to activate gene expression in response to specific signals.

RNA-Based Regulation
Riboswitches
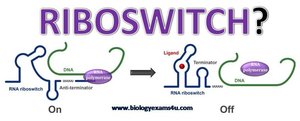
Riboswitches are regulatory segments of mRNA that bind small molecules, causing structural changes that influence gene expression. They can terminate transcription or block translation in response to metabolite concentrations.
Location: Typically found in the 5′ untranslated region (UTR) of mRNA.
Function: Act as built-in sensors for cellular metabolites.
RNA Interference (RNAi)
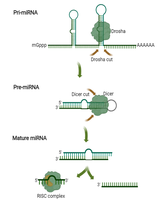
In eukaryotes, small non-coding RNAs such as microRNAs (miRNAs) and small interfering RNAs (siRNAs) regulate gene expression by targeting mRNAs for degradation or translational repression.
miRNA: Endogenously encoded, imperfect base pairing blocks translation or leads to mRNA degradation.
siRNA: Often exogenous, perfect base pairing leads to mRNA cleavage and degradation.
Long Non-coding RNAs (lncRNAs)
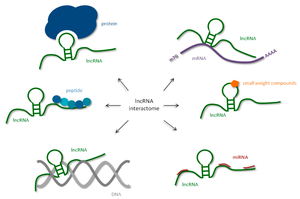
lncRNAs are RNA molecules longer than 200 nucleotides that do not code for proteins but regulate gene expression through chromatin remodeling, acting as decoys, or scaffolding molecular complexes.
Chromatin remodeling: Recruit proteins to modify chromatin structure (e.g., Xist in X-inactivation).
Decoys: Bind and sequester miRNAs or proteins, preventing their regulatory action.
Antisense RNA
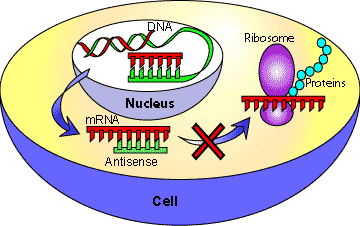
Antisense RNAs are complementary to specific mRNAs and can bind to them, forming double-stranded regions that block translation or trigger mRNA degradation, effectively silencing gene expression.
CRISPR-Cas Systems
In bacteria and archaea, CRISPR-Cas systems provide adaptive immunity against viruses. Small CRISPR RNAs (crRNAs) guide Cas enzymes to recognize and cleave foreign DNA, preventing infection and providing a molecular memory of past invaders.
Second Messengers in Signal Transduction
Role of Second Messengers
Second messengers are small intracellular molecules that relay signals from cell-surface receptors to internal targets, amplifying and distributing the signal to coordinate cellular responses.
Signal amplification: One receptor can generate many second messenger molecules, producing a large response.
Signal distribution: Second messengers activate multiple targets, coordinating complex cellular processes.
Spatiotemporal control: Cells tightly regulate the timing and location of second messenger activity.
Specialized Regulatory Mechanisms
Clocks: Timing and Periodicity
Molecular clocks are internal timing systems that regulate periodic biological processes, such as circadian rhythms, through feedback loops involving gene expression and protein degradation.
Feedback loop: Proteins inhibit their own gene expression, creating oscillatory cycles.
Function: Synchronize cellular activities with environmental cycles (e.g., day-night).
Thermometers: Temperature Sensing
Molecular thermometers are RNA or protein structures that change conformation in response to temperature, regulating gene expression accordingly.
RNA thermometers: Unfold at higher temperatures to allow translation of heat-shock proteins.
Protein thermometers: Change DNA-binding ability with temperature shifts.
Switches: Binary Decision Making
Molecular switches enable cells to toggle between two stable states, such as active/inactive or dividing/non-dividing, often through positive feedback or phosphorylation events.
Bistability: Positive feedback maintains the "on" state even after the initial signal is gone.
Phospho-switches: Protein activity is regulated by phosphorylation (kinases add, phosphatases remove phosphate groups).
G-proteins: Act as molecular switches, active when bound to GTP and inactive with GDP.
Summary Table: Key Mechanisms of Molecular Regulation
Mechanism | Main Function | Example |
|---|---|---|
Transcription Factors | Activate or repress gene transcription | Lac repressor, CAP activator |
Chromatin Remodeling | Control DNA accessibility | Histone acetylation |
Sigma Factors | Direct RNA polymerase to promoters | σ70, σ32 |
Riboswitches | Regulate gene expression via mRNA structure | Thiamine pyrophosphate riboswitch |
RNAi (miRNA/siRNA) | mRNA degradation or translation inhibition | miR-21, viral siRNAs |
lncRNA | Chromatin remodeling, decoy, scaffold | Xist |
Antisense RNA | Block translation, degrade mRNA | MicF RNA in E. coli |
CRISPR-Cas | Adaptive immunity | Cas9 |
Second Messengers | Signal transduction | cAMP, Ca2+ |
