 Back
BackViruses and Eukaryotes: Structure, Classification, and Replication
Study Guide - Smart Notes

General Properties of Viruses
Definition and Forms
Viruses are genetic elements that cannot replicate independently of a living host cell. The infectious form of a virus is called a virion, which consists of the nucleic acid genome surrounded by a protein coat and, in some cases, additional layers such as an envelope.
Virus particle: Extracellular form that facilitates transmission between host cells.
Virion: Infectious virus particle; nucleic acid genome surrounded by protein coat.
Viral Genomes
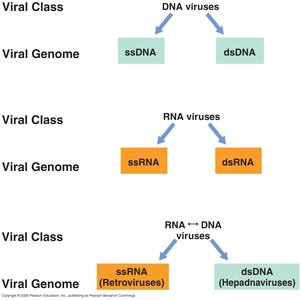
Viral genomes can be composed of either DNA or RNA, and may be linear or circular. Most viral genomes are smaller than those of biological cells.
DNA viruses: Single-stranded (ssDNA) or double-stranded (dsDNA).
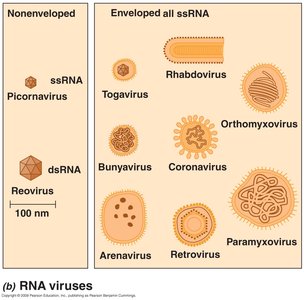
RNA viruses: Single-stranded (ssRNA) or double-stranded (dsRNA).
Retroviruses: ssRNA viruses that replicate via a DNA intermediate.
Viral Structure
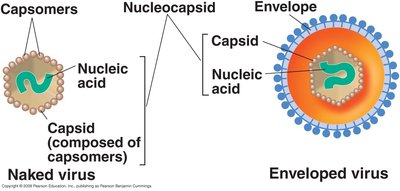
Viruses are classified based on their structure, which includes the capsid, capsomers, nucleocapsid, and envelope.
Capsid: Protein shell surrounding the genome.
Capsomer: Subunit of the capsid.
Nucleocapsid: Complete complex of nucleic acid and protein.
Enveloped virus: Virus with additional lipid bilayer surrounding the nucleocapsid.
Nature of the Virion
Symmetry Types
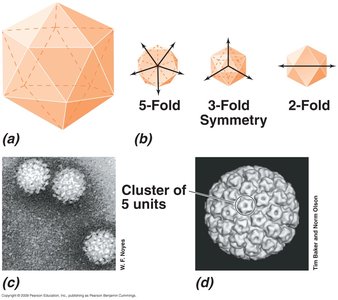
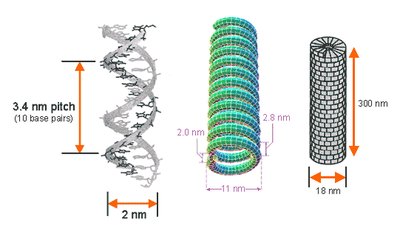
Virions are constructed in highly symmetric ways, most commonly helical or icosahedral symmetry.
Helical symmetry: Rod-shaped viruses; length determined by nucleic acid, width by protein subunits.
Icosahedral symmetry: Spherical viruses; most efficient arrangement of subunits in a closed shell.
Enveloped and Complex Viruses
Enveloped viruses: Have a membrane surrounding the nucleocapsid, derived from the host cell's lipid bilayer.
Complex viruses: Virions composed of several parts, often found in bacterial viruses with icosahedral heads and helical tails.
Viral Hosts and Taxonomy
Viruses are grouped based on the hosts they infect and their genomic characteristics. Most viruses are smaller than prokaryotic cells, ranging from 0.02 to 0.3 µm.
Quantification of Viruses
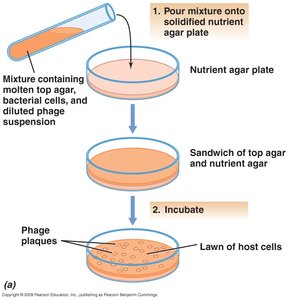
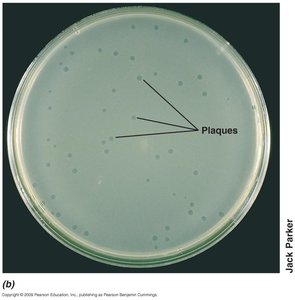
Plaque Assay
The plaque assay is one of the most accurate ways to measure virus infectivity. Plaques are clear zones that develop on lawns of host cells, each resulting from infection by a single virus particle.
Titer: Number of infectious units per volume of fluid.
Plaque assay: Analogous to bacterial colony counting.
General Features of Virus Replication
Phases of Viral Replication
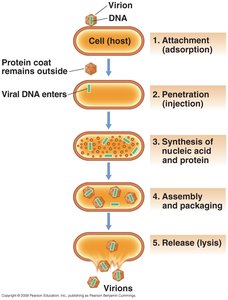
Virus replication typically follows a one-step growth curve, characterized by distinct phases:
Attachment (adsorption): Virus binds to a susceptible host cell.
Entry (penetration): Virion or its nucleic acid enters the host cell.
Synthesis: Virus nucleic acid and protein are synthesized using host cell metabolism.
Assembly: Capsids are assembled and viral genomes are packaged into new virions.
Release: New virions are released, often by cell lysis.
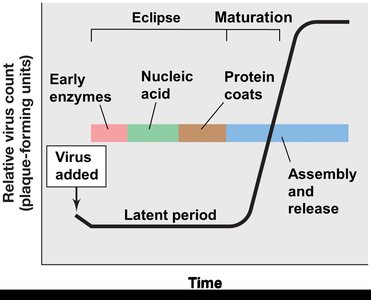
One-Step Growth Curve
Eclipse: Genome replication and protein translation.
Maturation: Packaging of nucleic acids in capsids.
Latent period: Eclipse + maturation; virions undetectable until release.
Burst size: Number of virions released per cell.
Viral Attachment and Penetration
Attachment of the virion to the host cell is highly specific, requiring complementary receptors on the host cell surface. Bacteriophage T4, a virus of E. coli, demonstrates a complex penetration mechanism involving tail fibers, lysozyme-like enzymes, and contraction of the tail sheath. 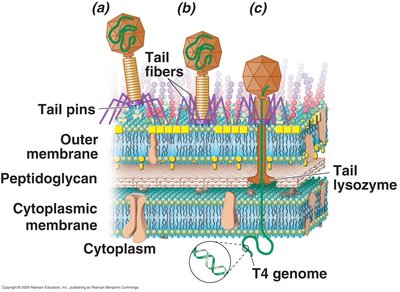
Bacteriophage T4 Replication
Genome Replication and Packaging
Bacteriophage T4 uses a complex replication strategy, including circular permutation and terminal redundancy. The genome is first replicated as a unit, then forms a concatemer (several genomic units recombined).
Circular permutation: Same genes arranged in different orders.
Terminal redundancy: DNA sequences duplicated at both ends.
Concatemer: Long DNA molecule with repeated genome units.
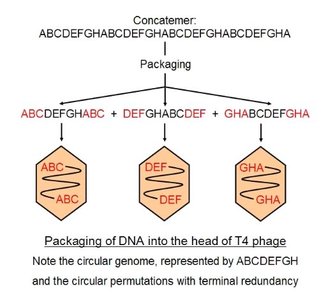
Transcription and Translation
T4 genome is divided into early, middle, and late proteins:
Early proteins: Enzymes for DNA replication and transcription.
Middle and late proteins: Head and tail proteins, enzymes for virion liberation.
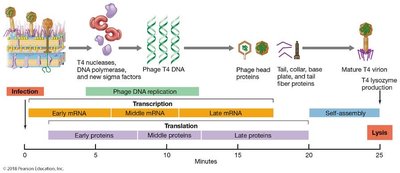
Virion Assembly and Release
Genome is pumped into the head under pressure using ATP. Packaging occurs in three stages: prohead assembly, packaging motor assembly, and DNA pumping. After the head is filled, tail and other components are self-assembled, and late enzymes break the membrane and peptidoglycan, resulting in lysis and virion release. 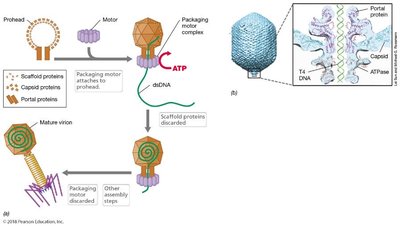
Viral Life Cycles
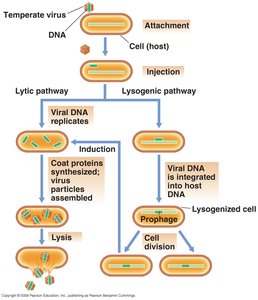
Virulent and Temperate Modes
Virulent mode: Viruses lyse host cells after infection.
Temperate mode: Viruses replicate their genomes in tandem with the host genome without killing the host.
Lysogeny: State where most virus genes are not expressed; virus genome (prophage) is replicated with host chromosome.
Lysogen: Bacterium containing a prophage.
Overview of Animal Viruses
Animal viruses exhibit all known modes of viral genome replication. Many are enveloped, and as they leave the host cell, they may acquire part of the host cell's lipid bilayer for their envelope. 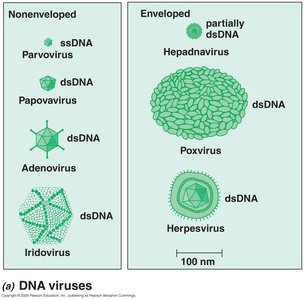
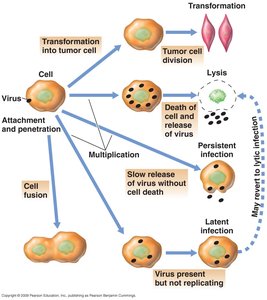
Consequences of Virus Infection in Animal Cells
Lysis: Destruction of host cell.
Persistent infection: Release of virions without cell lysis; cell remains alive.
Latent infection: Delay between infection and lytic events.
Transformation: Conversion of normal cell into tumor cell.
Retroviruses
Retroviruses are RNA viruses that replicate via a DNA intermediate. They are enveloped and contain reverse transcriptase, integrase, and protease.
Entrance into the cell and removal of envelope.
Reverse transcription of one RNA genome.
Integration of retroviral DNA into host genome (integrase).
Transcription of virus mRNA from viral DNA.
Assembly and packaging of genomic RNA.
Budding and release of enveloped virions.
Prions
Prions are infectious proteins known to cause disease in animals. Host cells contain the gene (PrnP) encoding the native prion protein. Misfolding results in neurological symptoms, resistance to proteases, insolubility, and aggregation.
Infectious prion disease: Pathogenic form transmitted between individuals.
Sporadic prion disease: Random misfolding in uninfected individuals.
Inherited prion disease: Mutation in prion gene increases misfolding.
Key Terms and Concepts
Positive-strand RNA virus: ssRNA genome with same orientation as mRNA.
Negative-strand RNA virus: ssRNA genome with complementary orientation to mRNA.
Concatemer: Long DNA molecule with repeated genome units.
Terminal redundancy: DNA sequences duplicated at both ends.
Circular permutation: Same genes arranged in different orders.
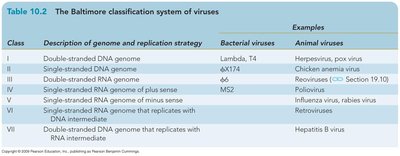
Baltimore Classification System of Viruses
The Baltimore system classifies viruses based on genome type and replication strategy.
Class | Description of genome and replication strategy | Bacterial viruses | Animal viruses |
|---|---|---|---|
I | Double-stranded DNA genome | Lambda, T4 | Herpesvirus, pox virus |
II | Single-stranded DNA genome | φX174 | Chicken anemia |
III | Double-stranded RNA genome | φ6 | Rotavirus |
IV | Single-stranded RNA genome of plus sense | MS2 | Poliovirus |
V | Single-stranded RNA genome of minus sense | Influenza virus, rabies virus | |
VI | Single-stranded RNA genome that replicates with DNA intermediate | Retrovirus | |
VII | Double-stranded DNA genome that replicates with RNA intermediate | Hepatitis B virus |

