 Back
BackViruses, Viroids, and Prions: Structure, Replication, and Pathogenicity
Study Guide - Smart Notes

Viruses: Structure, Classification, and Replication
Comparing Cells and Viruses
Viruses are fundamentally different from cellular life forms. They are acellular, obligate intracellular parasites that require a host cell for replication. In contrast, cells are self-replicating, metabolically active, and can grow and divide independently.
Viruses: Acellular, contain either DNA or RNA (never both), do not metabolize or grow outside a host, replicate using host machinery.
Cells: Cellular, contain both DNA and RNA, metabolize and grow independently, replicate by asexual and/or sexual means.

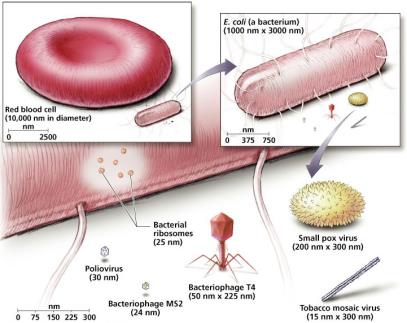
Relative Sizes of Viruses
Viruses are much smaller than most cells and even many organelles. Their size typically ranges from 10 nm to over 500 nm, making them ultramicroscopic.
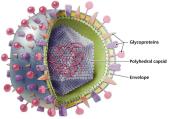
Structure of Viruses
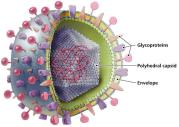
Viruses exist in two states: extracellular (virion) and intracellular. The virion consists of a nucleic acid genome surrounded by a protein capsid, and in some cases, an envelope derived from the host cell membrane.
Capsid: Protein shell that protects the viral genome and aids in attachment to host cells.
Envelope: Lipid bilayer with embedded proteins (often glycoprotein spikes) acquired from the host cell during viral replication; present in some animal viruses.
Genome: DNA or RNA, which may be single- or double-stranded, linear or circular, segmented or non-segmented.


Viral Envelopes
Enveloped viruses are more fragile than non-enveloped viruses due to their lipid bilayer, which is sensitive to environmental factors. The envelope contains viral glycoproteins essential for host cell recognition and entry.
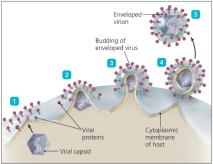
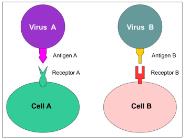
Host Range and Specificity
Viruses exhibit specificity for their host due to interactions between viral surface proteins and host cell receptors. Most viruses have a narrow host range, but some are generalists.
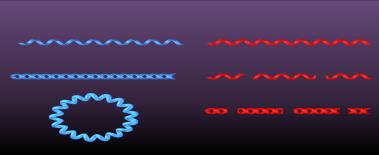
Genetic Material of Viruses
Viral genomes are highly diverse and are a primary basis for virus classification. They may be DNA or RNA, single- or double-stranded, linear, circular, or segmented.
dsDNA: Double-stranded DNA
ssDNA: Single-stranded DNA
dsRNA: Double-stranded RNA
ssRNA: Single-stranded RNA (positive-sense or negative-sense)

Classification of Viruses
Viruses are classified based on:
Type and structure of nucleic acid
Host range
Size and shape (helical, polyhedral, complex)
Capsid structure
Presence or absence of envelope
Viral Replication Mechanisms
Viruses rely on host cell machinery for replication. There are three main replication strategies:
Lytic replication: Results in host cell lysis and release of new virions.
Lysogenic replication: Viral genome integrates into host DNA and can remain dormant (prophage/provirus).
Latent replication: Dormancy in animal cells, with or without integration into host genome.

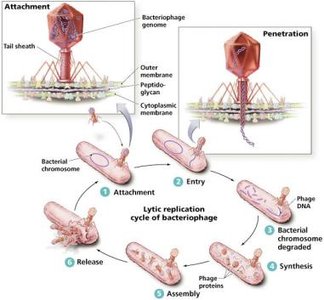
Lytic Replication Cycle
The lytic cycle consists of five steps:
Attachment
Entry
Synthesis
Assembly
Release
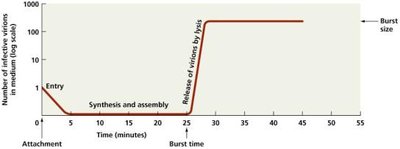
Burst time is the period required to complete the lytic cycle, and burst size is the number of virions released per lysed cell.
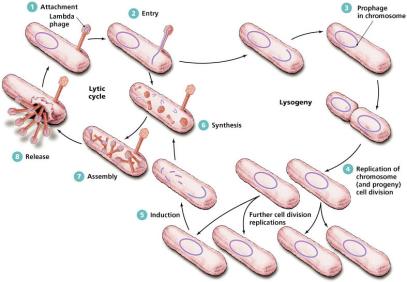
Lysogenic and Latent Replication
In lysogeny, the viral genome integrates into the host chromosome and is replicated along with it. Induction can trigger entry into the lytic cycle. Latency in animal viruses can be temporary or permanent, with the viral genome sometimes becoming a permanent part of the host DNA.
Replication of Animal Viruses
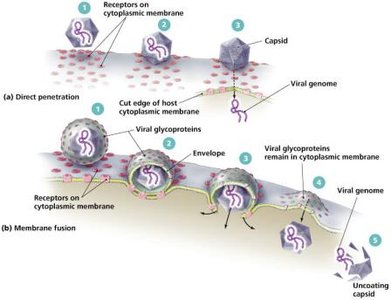
Animal viruses may enter cells by direct penetration, membrane fusion, or phagocytosis. Uncoating of the capsid is required for genome release. DNA viruses often replicate in the nucleus, while RNA viruses replicate in the cytoplasm.
Synthesis Strategies of Animal Viruses
The synthesis of viral components depends on the type of nucleic acid:
dsDNA viruses: Replicate genome in nucleus, proteins in cytoplasm.
ssDNA viruses: Form dsDNA intermediate for replication.
+ssRNA viruses: Genome acts as mRNA.
-ssRNA viruses: Require RNA-dependent RNA polymerase to synthesize mRNA.
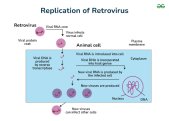
Retroviruses: Use reverse transcriptase to make DNA from RNA, which integrates into host genome.
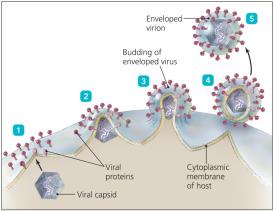
Assembly and Release of Animal Viruses
Most DNA viruses assemble in the nucleus; most RNA viruses assemble in the cytoplasm. Enveloped viruses are released by budding, causing persistent infections, while non-enveloped viruses are released by lysis or exocytosis.
Bacteriophage vs. Animal Virus Replication
Step | Bacteriophage | Animal Virus |
|---|---|---|
Attachment | Proteins on tails attach to cell wall | Spikes, capsids, or envelope proteins attach to cell membrane |
Penetration | Genome injected or diffuses into cell | Capsid enters by direct penetration, fusion, or endocytosis |
Uncoating | None | Capsid removed by cell enzymes |
Site of Synthesis | Cytoplasm | RNA viruses: cytoplasm; DNA viruses: nucleus |
Site of Assembly | Cytoplasm | RNA viruses: cytoplasm; DNA viruses: nucleus |
Release | Lysis | Naked: exocytosis/lysis; Enveloped: budding |
Chronic Infection | Lysogeny (may leave chromosome) | Latency (with/without integration; permanent if integrated) |
Examples of Important Viruses
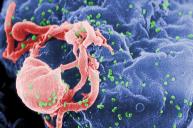
Human Immunodeficiency Virus (HIV)
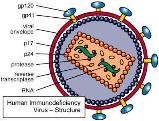
HIV is a retrovirus with a double-stranded RNA genome. It carries reverse transcriptase, integrase, and protease enzymes. HIV infects CD4 T-cells and macrophages, leading to immune deficiency.
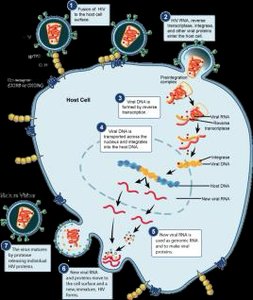
Attachment: gp120 binds CD4
Entry: Reverse transcription of RNA to DNA
Integration: Viral DNA integrates into host genome
Synthesis: Production of viral RNA and proteins
Assembly and Release: New virions assembled and released
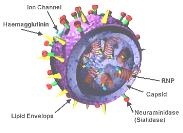
Influenza Virus
Influenza is an enveloped RNA virus with a segmented genome. It uses hemagglutinin (H) for attachment and neuraminidase (N) for release. Antigenic drift (minor changes) and shift (major changes) in H and N proteins can lead to new strains and pandemics.
Attachment: H binds sialic acid on host cell
Entry: Endocytosis and nuclear entry of vRNA
Synthesis: RNA-dependent RNA polymerase synthesizes RNA
Assembly and Release: N cleaves sialic acid for virion release
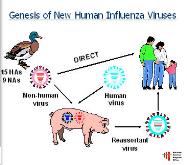
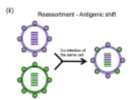
Antigenic Drift and Shift in Influenza
Antigenic drift involves gradual mutations in H and N glycoproteins, while antigenic shift involves gene reassortment, leading to major changes and potential pandemics.
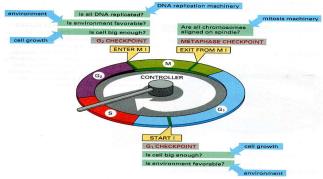
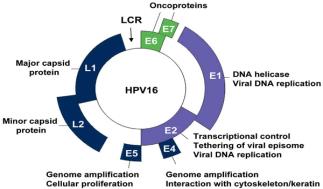
Viruses and Cancer
Oncogenic Viruses
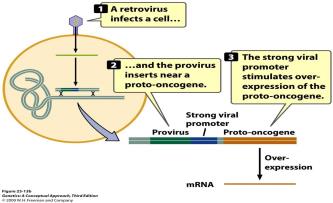
Some viruses can cause cancer by disrupting normal cell cycle regulation. They may insert into tumor suppressor genes, increase oncogene expression, or carry viral oncogenes.
Examples: Epstein-Barr virus, Human papillomavirus (HPV), Hepatitis B & C, HIV, Human herpesvirus 8, HTLV-1
Other Parasitic Particles: Viroids and Prions
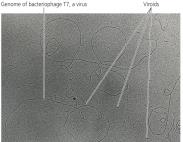
Viroids
Viroids are small, circular ssRNA molecules that infect plants. They lack a protein capsid and do not code for proteins, but can cause significant plant diseases by interfering with host RNA.

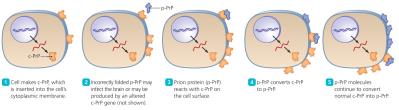
Prions
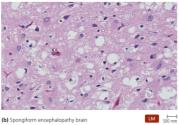
Prions are infectious proteins that cause neurodegenerative diseases. The disease-causing form (Prion PrP) has a β-pleated sheet structure, which can induce normal cellular PrP (with α-helices) to misfold. Prion diseases are fatal and untreatable, causing spongiform encephalopathies in the brain.
Transmission: Ingestion, transplantation, or contact with infected tissues
Diseases: BSE (cows), Scrapie (sheep), CWD (deer/elk), Kuru, vCJD (humans)
Comparison Table: Bacteria, Viruses, Viroids, and Prions
Bacteria | Viruses | Viroids | Prions | |
|---|---|---|---|---|
Width | 200–2000 nm | 10–400 nm | 2 nm | 5 nm |
Length | 200–550,000 nm | 20–800 nm | 40–130 nm | 5 nm |
Nucleic Acid | DNA & RNA | DNA or RNA | RNA only | None |
Protein | Present | Present | Absent | Present (PrP) |
Cellular | Yes | No | No | No |
Cytoplasmic Membrane | Present | Absent (some have envelope) | Absent | Absent |
Functional Ribosomes | Present | Absent | Absent | Absent |
Growth | Present | Absent | Absent | Absent |
Self-Replicating | Yes | No | No | No; transforms PrP in cell |
Responsiveness | Present | Some bacteriophages respond to host | Absent | Absent |
Metabolism | Present | Absent | Absent | Absent |
