 Back
BackBiochemistry Exam II & III Study Guidance
Study Guide - Smart Notes

Q1. The reaction below shows a general method used by proteases to catalyze the cleavage of a peptide bond. In this figure, 1 and 2 are generic proton donors and acceptors that are part of the enzyme active site. The rest of the polypeptide chain, which surrounds the substrate in the active site and sequesters it from bulk solvent, is not shown for clarity.
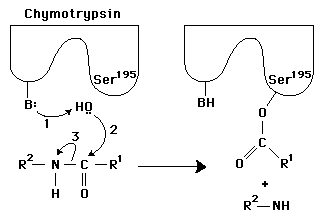
Background
Topic: Enzyme Catalysis & Reaction Mechanisms
This question tests your understanding of how enzymes (specifically proteases like chymotrypsin) catalyze peptide bond hydrolysis, the energy profile of catalyzed vs. uncatalyzed reactions, and the roles of amino acid side chains in catalysis.
Key Terms and Formulas
Reaction coordinate diagram: A plot of energy vs. reaction progress, showing reactants, transition state, and products.
Transition state: The highest energy point along the reaction pathway.
Enzyme catalysis: Enzymes lower the activation energy () of a reaction, increasing the rate.
Mechanisms of catalysis: General acid-base catalysis, covalent catalysis, and proximity/orientation effects.
Step-by-Step Guidance
Draw two reaction coordinate diagrams (energy vs. reaction progress): one for the uncatalyzed reaction and one for the enzyme-catalyzed reaction. Label the energy of reactants, transition state, and products for each.
On the enzyme-catalyzed diagram, show that the activation energy () is lower than in the uncatalyzed reaction, but the overall free energy change () is the same for both.
Explain that enzymes speed up reactions by stabilizing the transition state, thus lowering the activation energy required.
Identify the catalytic mechanisms used by the enzyme in the figure (e.g., general acid-base catalysis, covalent catalysis) based on the roles of the groups labeled 1 and 2.
Propose specific amino acids that could serve as proton donors/acceptors (e.g., histidine, serine, aspartate) and justify your choices based on their chemical properties.
Try solving on your own before revealing the answer!
Q2. Lactate dehydrogenase (LDH) catalyzes the reduction of pyruvate with NADH to form lactate. A mutant form of LDH (Arg109→Gln) shows only 25% of the pyruvate binding and 0.07% of the activity of wild-type enzyme.
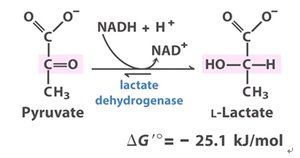
Background
Topic: Enzyme Structure-Function Relationships & Mutagenesis
This question examines how specific amino acid changes affect substrate binding and catalytic activity in enzymes, using LDH as an example.
Key Terms and Concepts
Active site: The region of an enzyme where substrate binding and catalysis occur.
Binding vs. catalysis: Binding refers to substrate association with the enzyme; catalysis refers to the chemical transformation.
Mutagenesis: Changing specific amino acids to study their roles.
Step-by-Step Guidance
Consider why pyruvate can still bind to the mutant enzyme (Arg109→Gln) at a significant level, even though the residue is changed.
Think about the chemical properties of arginine vs. glutamine and how these might affect substrate binding and/or catalysis.
Explain why the catalytic activity is much lower in the mutant, despite some binding still occurring.
For the Ile250→Gln mutant, consider how changing a hydrophobic residue to a polar one might affect NADH binding.
Try solving on your own before revealing the answer!
Q3. From the table above, which enzyme/substrate pair could be classified as the “most efficient?” Include one sentence that justifies your answer.
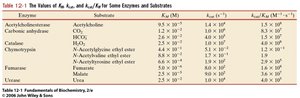
Background
Topic: Enzyme Kinetics (kcat, KM, and Catalytic Efficiency)
This question tests your ability to interpret kinetic parameters and identify the most efficient enzyme-substrate pair using .
Key Terms and Formulas
(turnover number): Number of substrate molecules converted to product per enzyme per second.
(Michaelis constant): Substrate concentration at which the reaction rate is half-maximal.
Catalytic efficiency: (units: Ms$^{-1}$)
Step-by-Step Guidance
Examine the table and identify the enzyme-substrate pair with the highest value.
Justify your choice in one sentence, explaining why a high indicates high efficiency.
Try solving on your own before revealing the answer!
Q4. Dr. Burke is developing inhibitors for alcohol dehydrogenase. Kinetic data for the enzyme in the absence and presence of two inhibitors (B and C) are given as linear equations. Label the axes, determine and for each trial, and identify the type of inhibition.
Background
Topic: Enzyme Inhibition & Lineweaver-Burk Plots
This question tests your ability to interpret kinetic data, calculate and from Lineweaver-Burk plots, and classify types of enzyme inhibition.
Key Terms and Formulas
Lineweaver-Burk plot: Double reciprocal plot of vs. .
Equation:
Inhibition types: Competitive, noncompetitive, uncompetitive (distinguished by changes in and ).
Step-by-Step Guidance
Label the axes: -axis is , -axis is .
For each trial, identify the slope and y-intercept from the equation and relate them to and .
Calculate and for each trial using the slope and intercept.
Compare the values to determine the type of inhibition (e.g., does or change?).
Predict which inhibitor is a better candidate based on the data and your reasoning.
Try solving on your own before revealing the answer!
Q5. Raffinose is a trisaccharide present in beans, broccoli, cabbage, etc. Humans do not possess an α-galactosidase enzyme necessary to break down raffinose. Which of the bonds indicated by the letters is hydrolyzed by the α-galactosidase in Beano®? Explain.
Background
Topic: Carbohydrate Structure & Enzyme Specificity
This question tests your understanding of glycosidic bonds and the specificity of carbohydrate-degrading enzymes.
Key Terms and Concepts
Trisaccharide: A carbohydrate composed of three monosaccharides.
α-galactosidase: An enzyme that hydrolyzes α-galactosidic bonds.
Step-by-Step Guidance
Identify the glycosidic bonds present in raffinose and determine which one is an α-galactosidic bond.
Explain why humans cannot digest this bond and how Beano® helps.
Try solving on your own before revealing the answer!
Q6. Two polysaccharides used for storing sugars include amylopectin and glycogen. These polysaccharides differ in that:
Background
Topic: Polysaccharide Structure & Function
This question tests your knowledge of the structural differences between amylopectin and glycogen.
Key Terms and Concepts
Amylopectin: A branched polysaccharide found in plants.
Glycogen: A highly branched polysaccharide found in animals.
Step-by-Step Guidance
Recall the branching frequency and linkage types in amylopectin and glycogen.
Compare their structures and relate these differences to their biological roles.
Try solving on your own before revealing the answer!
Q7. Snake venom contains the enzyme phospholipase A2. The action of phospholipase A2 cleaves the fatty acid tail from position 2 of a phospholipid. Illustrate the action and discuss the structural ramifications on the plasma membrane. Could this be reversed? Explain.
Background
Topic: Lipid Structure & Enzyme Action
This question tests your understanding of phospholipid structure, enzyme specificity, and the consequences of lipid hydrolysis on membrane integrity.
Key Terms and Concepts
Phospholipase A2: An enzyme that hydrolyzes the ester bond at the sn-2 position of phospholipids.
Plasma membrane: A lipid bilayer that forms the boundary of cells.
Step-by-Step Guidance
Draw or visualize a generic phospholipid and indicate the sn-2 position.
Describe what happens structurally when the fatty acid at position 2 is removed.
Discuss the effects on membrane properties and whether the reaction is reversible.
Try solving on your own before revealing the answer!
Q8. Rank the melting points of the following fatty acids from highest to lowest and justify your answer: (1) cis-Palmitoleic (16:1), (2) trans-oleic (18:1), (3) cis-linolenic (18:3), (4) arachidic (20:0), (5) palmitic (16:0)
Background
Topic: Fatty Acid Structure & Physical Properties
This question tests your understanding of how chain length and degree of unsaturation affect melting points of fatty acids.
Key Terms and Concepts
Saturated fatty acids: No double bonds; higher melting points.
Unsaturated fatty acids: One or more double bonds; lower melting points (cis < trans).
Step-by-Step Guidance
List the fatty acids by chain length and degree/type of unsaturation.
Recall that more double bonds (especially cis) lower melting point, while longer chains increase it.
Arrange the fatty acids from highest to lowest melting point based on these principles.
Try solving on your own before revealing the answer!
Q9. Examine the following nucleotide sequence from the sense strand of DNA. a) What is the amino acid sequence of the encoded protein? b) Would you predict this peptide to reside in the cytosol or plasma membrane? Why? c) Suggest a specific amino acid mutation that might influence the structure and function of this peptide and explain your reasoning. d) For the above suggested mutation to be inheritable, in which biopolymer would the mutation need to occur and why?
Background
Topic: Genetic Code, Protein Structure, and Mutations
This question tests your ability to translate DNA to protein, predict localization, and understand the molecular basis of inheritance.
Key Terms and Concepts
Genetic code: The set of rules by which nucleotide sequences are translated into amino acids.
Mutation: A change in the DNA sequence that can affect protein structure and function.
Step-by-Step Guidance
Translate the DNA sequence into the corresponding amino acid sequence using a codon table.
Analyze the sequence for hydrophobic/hydrophilic regions to predict localization.
Suggest a mutation (e.g., hydrophobic to hydrophilic) and explain its potential effect.
Identify which biopolymer must be mutated for the change to be inheritable and justify your answer.
Try solving on your own before revealing the answer!
Q10. Actinomycin D binds DNA and prevents transcription. a) Define intercalating agent. b) Indicate in the figure where this molecule intercalates. c) What effect does this interaction have with the melting temperature of DNA? Justify your answer.
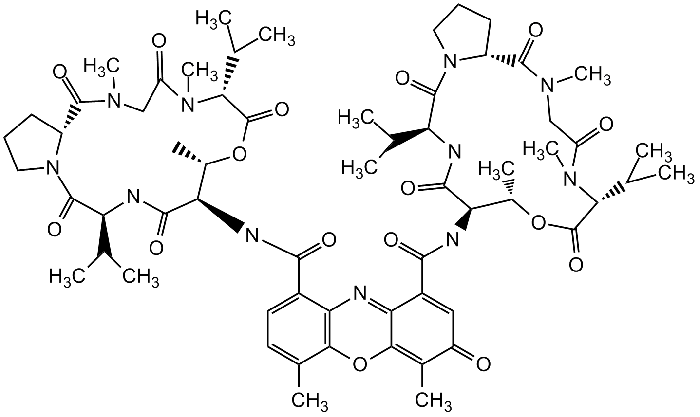
Background
Topic: DNA Structure, Intercalating Agents, and Transcription Inhibition
This question tests your understanding of how small molecules interact with DNA and affect its physical properties and biological function.
Key Terms and Concepts
Intercalating agent: A molecule that inserts between DNA base pairs, distorting the double helix.
Melting temperature (): The temperature at which half of the DNA helix denatures.
Step-by-Step Guidance
Define what an intercalating agent is and how it interacts with DNA.
Identify the planar aromatic region of actinomycin D that intercalates between base pairs.
Explain how intercalation affects the stability and melting temperature of DNA.
Try solving on your own before revealing the answer!
