Back
BackBiochemistry Study Guide: Protein Structure, Amino Acids, Buffers, and Protein Techniques
Study Guide - Smart Notes

Hydrophobic Interactions
The Hydrophobic Effect
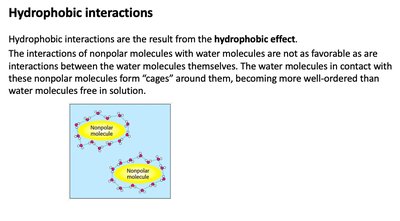
Hydrophobic interactions are a fundamental non-covalent force in biochemistry, arising from the tendency of nonpolar molecules to minimize contact with water. This effect is crucial for protein folding, membrane formation, and molecular assembly.
Hydrophobic effect: Water molecules form 'cages' around nonpolar molecules, increasing order and decreasing entropy locally.
Biological relevance: Drives the aggregation of hydrophobic amino acids in protein interiors and the formation of lipid bilayers.
Example: Protein folding relies on hydrophobic residues clustering away from water.

Acid-Base Equilibria and Buffers
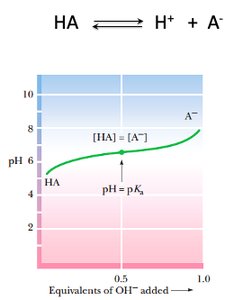
Monoprotic Acid Titration Curve
Acid-base equilibria are central to biochemistry, affecting enzyme activity, protein structure, and metabolic pathways. The titration curve of a monoprotic acid illustrates the relationship between pH and the degree of dissociation.
Key equation:
Henderson-Hasselbalch equation:
Buffer region: pH changes slowly near pKa, where [HA] = [A^-].
Polyprotic Acid Titration Curve
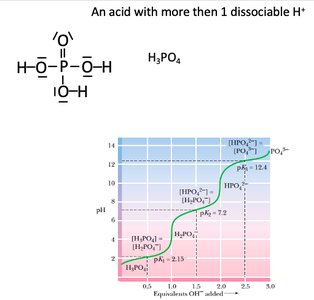
Some acids, such as phosphoric acid (H3PO4), have multiple dissociable protons, resulting in multiple buffer regions and pKa values.
Multiple pKa values: Each dissociation step has its own pKa and buffer region.
Biological relevance: Polyprotic acids are important in cellular buffering systems (e.g., phosphate buffer).
Amino Acids
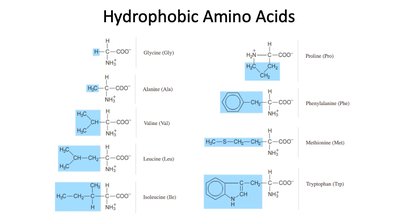
Hydrophobic Amino Acids
Hydrophobic amino acids possess nonpolar side chains that do not interact favorably with water. They are typically found in the interior of proteins, stabilizing the folded structure through hydrophobic interactions.
Examples: Glycine, Alanine, Valine, Leucine, Isoleucine, Proline, Phenylalanine, Methionine, Tryptophan.
Structural features: Side chains highlighted in blue are nonpolar and hydrophobic.
Role in proteins: Promote compact folding and stability.
Protein Structure
Polypeptide Directionality
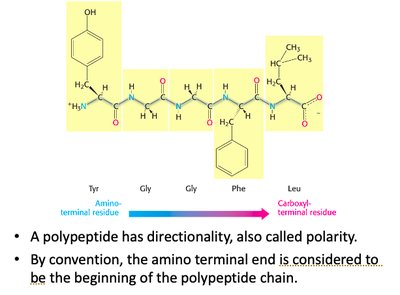
Proteins are linear polymers of amino acids, exhibiting directionality from the amino (N-) terminus to the carboxyl (C-) terminus. This polarity is essential for understanding protein synthesis and function.
Convention: The N-terminus is considered the start of the polypeptide chain.
Polarity: Directionality affects protein folding and biological activity.
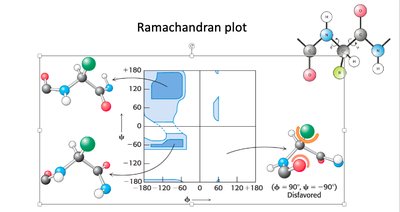
Ramachandran Plot
The Ramachandran plot is a graphical representation of the allowed dihedral angles (phi and psi) in polypeptide backbones. It is used to predict secondary structure and assess protein models.
Allowed regions: Indicate conformations with minimal steric hindrance.
Secondary structure: Alpha helices and beta sheets occupy distinct regions.
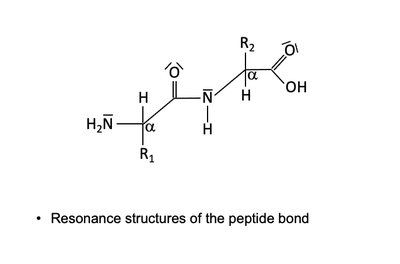
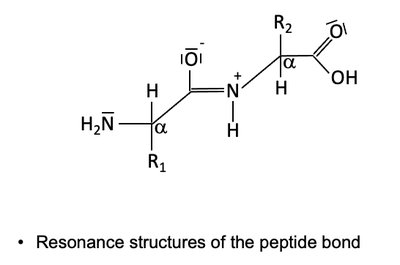
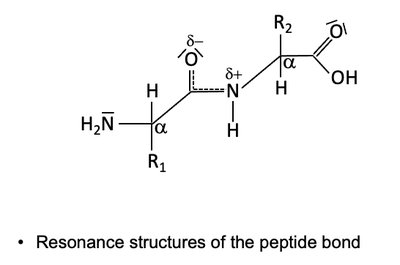
Peptide Bond Resonance
The peptide bond exhibits resonance, resulting in partial double-bond character and planarity. This restricts rotation and contributes to protein structure stability.
Resonance forms: Peptide bond can be represented by multiple resonance structures.
Implication: Planarity and rigidity of the peptide bond affect protein folding.
Hemoglobin and Myoglobin
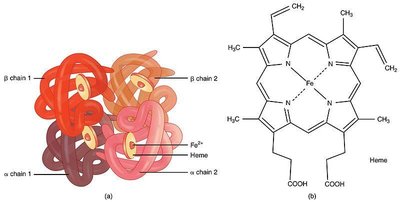
Hemoglobin Structure
Hemoglobin is a tetrameric protein composed of two alpha and two beta chains, each containing a heme group. It is responsible for oxygen transport in blood.
Heme group: Contains Fe2+ at its center, essential for oxygen binding.
Quaternary structure: Four polypeptide chains assemble into a functional complex.
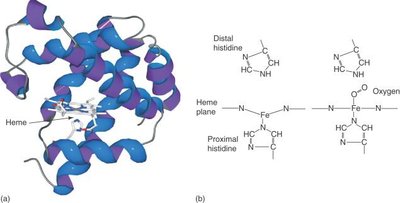
Myoglobin Structure
Myoglobin is a monomeric protein found in muscle tissue, specialized for oxygen storage. It contains a single heme group and exhibits high oxygen affinity.
Structure: Composed mainly of alpha helices.
Heme coordination: Proximal and distal histidine residues stabilize oxygen binding.
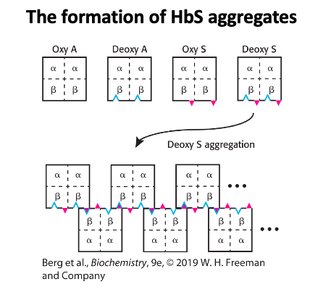
Sickle Cell Aggregation
Sickle cell disease arises from a mutation in hemoglobin, causing aggregation of deoxy-HbS molecules and distortion of red blood cells.
Mutation: Glutamic acid replaced by valine in beta chain.
Aggregation: Deoxy-HbS forms rigid fibers, leading to sickle-shaped cells.
Protein Secondary Structure
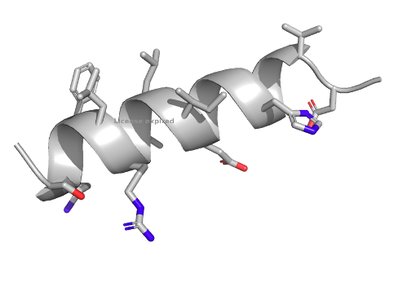
Alpha Helix
The alpha helix is a common secondary structure stabilized by hydrogen bonds between backbone carbonyl and amine groups.
Hydrogen bonding: Each carbonyl oxygen forms a hydrogen bond with the amine four residues ahead.
Structure: Right-handed, 3.6 residues per turn.
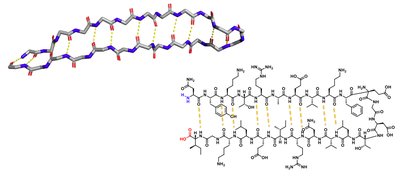
Beta Sheet
Beta sheets are formed by hydrogen bonding between adjacent beta strands, which can be parallel or antiparallel.
Hydrogen bonding: Occurs across the sheet.
Structure: Can be flat or twisted; forms barrels in some proteins.
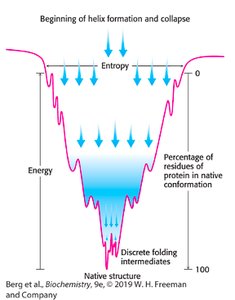
Protein Folding Funnel
Protein folding is often visualized as a funnel, representing the decrease in entropy and energy as the protein adopts its native conformation.
Folding intermediates: Proteins pass through discrete intermediates before reaching native structure.
Thermodynamics: Native structure is the lowest energy, most stable form.
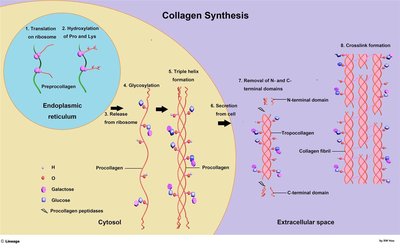
Collagen Structure and Synthesis
Collagen Synthesis
Collagen is a structural protein with a unique triple helix structure, synthesized through a multi-step process involving post-translational modifications.
Steps: Translation, hydroxylation, glycosylation, triple helix formation, secretion, cleavage, crosslinking.
Enzymes: Prolyl and lysyl hydroxylases require vitamin C.
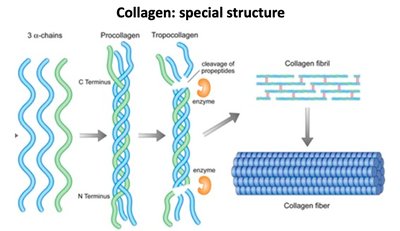
Collagen Special Structure
Collagen's triple helix is formed by three polypeptide chains, which are processed into tropocollagen and assembled into fibrils and fibers.
Triple helix: Provides tensile strength and stability.
Fibril formation: Essential for connective tissue integrity.
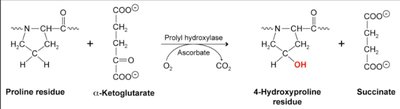
Posttranslational Hydroxylation
Hydroxylation of proline and lysine residues is a critical post-translational modification in collagen, stabilizing the triple helix and enabling crosslinking.
Hydroxyproline: Forms interchain hydrogen bonds.
Hydroxylysine: Allows interchain covalent bonds.
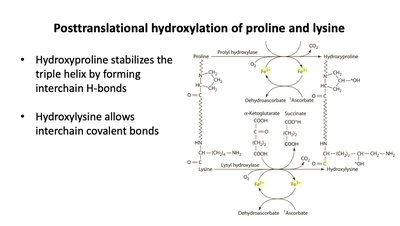
Vitamin C and Vitamin K
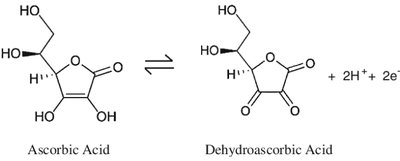
Vitamin C (Ascorbic Acid)
Vitamin C acts as a reducing agent, essential for the activity of prolyl and lysyl hydroxylases in collagen synthesis.
Redox reaction: Converts Fe3+ to Fe2+ in enzyme active sites.
Deficiency: Leads to impaired collagen synthesis (scurvy).
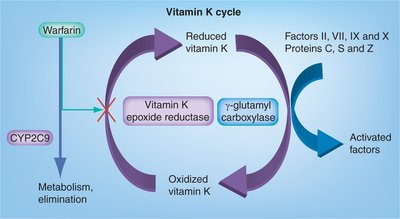
Vitamin K Cycle
Vitamin K is required for the gamma-carboxylation of glutamate residues in clotting factors, enabling calcium binding and proper coagulation.
Cycle: Vitamin K is recycled via epoxide reductase, which is inhibited by warfarin.
Clinical relevance: Warfarin is an anticoagulant that blocks vitamin K recycling.
Protein Techniques
Gel Filtration Chromatography
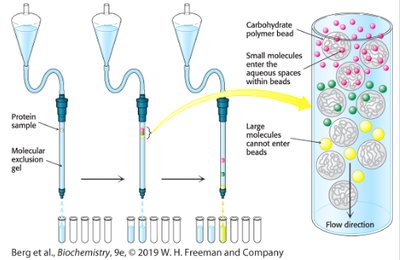
Gel filtration (size-exclusion) chromatography separates proteins based on size. Large molecules elute first, while small molecules enter beads and elute later.
Principle: Separation by molecular size.
Application: Purification of proteins and determination of molecular weight.
Ion Exchange Chromatography
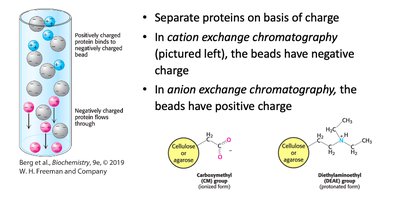
Ion exchange chromatography separates proteins based on charge. Cation exchange uses negatively charged beads; anion exchange uses positively charged beads.
Principle: Separation by net charge.
Application: Protein purification and analysis.
Affinity Chromatography
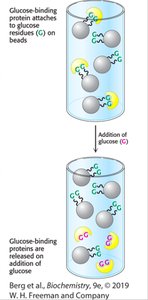
Affinity chromatography exploits specific binding interactions between a protein and a ligand attached to the column. Only proteins with affinity for the ligand are retained.
Principle: Separation by specific binding.
Application: Purification of tagged proteins (e.g., His-tag).
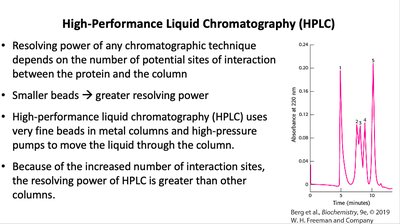
High-Performance Liquid Chromatography (HPLC)
HPLC uses fine beads and high-pressure pumps for high-resolution separation of proteins and other biomolecules.
Principle: Increased interaction sites yield greater resolving power.
Application: Analytical and preparative separation of biomolecules.
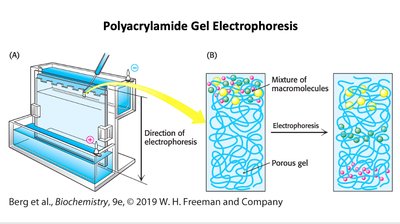
Polyacrylamide Gel Electrophoresis (PAGE)
PAGE separates proteins based on size, charge, and shape. SDS-PAGE standardizes charge and shape, allowing separation solely by molecular weight.
Principle: Migration through gel under electric field.
Application: Protein purity, size determination, and subunit analysis.
Enzyme Structure
Active Sites of Enzymes
Enzyme active sites are three-dimensional pockets where substrates bind and reactions occur. The specificity and catalytic efficiency of enzymes depend on the precise arrangement of amino acids in the active site.
Binding site: Positions substrate correctly.
Catalytic site: Facilitates chemical transformation.